Construction of CRISPR Plasmids and Detection of Knockout Efficiency in Mammalian Cells through a Dual Luciferase Reporter System
Summary
Here, we present a protocol describing a streamlined method for the efficient generation of plasmids expressing both the CRISPR enzyme and associated single guide RNA (sgRNAs). Co-transfection of mammalian cells with this sgRNA/CRISPR vector and a dual luciferase reporter vector that examines double-strand break repair allows evaluation of knockout efficiency.
Abstract
Although highly efficient, modification of a genomic site by the CRISPR enzyme requires the generation of a sgRNA unique to the target site(s) beforehand. This work describes the key steps leading to the construction of efficient sgRNA vectors using a strategy that allows the efficient detection of the positive colonies by PCR prior to DNA sequencing. Since efficient genome editing using the CRISPR system requires a highly efficient sgRNA, a preselection of candidate sgRNA targets is necessary to save time and effort. A dual luciferase reporter system has been developed to evaluate knockout efficiency by examining double-strand break repair via single strand annealing. Here, we use this reporter system to pick up the preferred xCas9/sgRNA target from candidate sgRNA vectors for specific gene editing. The protocol outlined will provide a preferred sgRNA/CRISPR enzyme vector in 10 days (starting with appropriately designed oligonucleotides).
Introduction
The CRISPR sgRNAs comprise a 20-nucleotide sequence (the protospacer), which is complementary to the genomic target sequence1,2. Although highly efficient, the ability of the CRISPR/Cas system to modify a given genomic site requires the generation of a vector carrying an efficient sgRNA unique to the target site(s)2. This paper describes the key steps in the generation of that sgRNA vector.
For successful genome editing using the CRISPR/Cas system, the use of highly efficient sgRNAs is a crucial prerequisite3,4,5. Since engineered nucleases used in genome editing manifest diverse efficiencies at different targeted loci1, a pre-selection of candidate sgRNA targets is necessary in order to save time and effort6,7,8,9. A dual luciferase reporter system has been developed to evaluate knockout efficiency by examining double-strand break repair via single strand annealing3,10. Here we use this reporter system to choose a preferred CRISPR sgRNA target from different candidate sgRNA vectors designed for specific gene editing. The protocol stated here has been implemented in our group and collaborating laboratories for the last few years to generate and evaluate CRISPR sgRNAs.
The following protocol sums up how to design suitable sgRNA through network software. Once the suitable sgRNAs are selected, we describe the different steps to obtain the required oligonucleotides as well as the approach for inserting the paired oligonucleotides into the pX330-xCas9 expression vector. We also present a method for assembling sgRNA-expressing and dual luciferase reporter vectors based on the ligation of these sequences into a predigested expression vector (steps 2-10, Figure 1A). Finally we describe how to analyze the the DNA cutting efficiency for each of the sgRNAs (steps 11-12).
Protocol
1. sgRNA oligonucleotide design
- Design sgRNAs using online tools such as the Cas-Designer online tool (http://www.rgenome.net/cas-designer/). The PAM sequence is important based on the Cas9 being used. For xCas9, the relevant PAM sequences are NG and the former referred Cas-Designer online tool can generate xCas9 relevant sgRNAs.
- Use sgRNA design tools that encompass algorithms for on- and off-target prediction (http://www.broadinstitute.org/rnai/public/analysis-tools/sgrna-design)11. A score of 0.2 or greater is preferred.
- Select up to 3 gene editing targets for an optimum screening (e.g., T1, T2 and T3 were designed for sheep DKK2 exon 1 gene targeting [Table 1]).
2. Oligonucleotide modification
- To modify the sgRNA oligonucleotide, delete the 3’-NG protospacer adjacent motif (PAM), keeping the protospacer sequence (e.g., starting sequence for T1: TGCCTGCTCCTACTGGCCGC [20 nt]).
- Add the pentanucleotide CACCG to the 5’-end of the oligo.
NOTE: Upon ligation to the pX330-xCas9 skeleton, this sequence will contain the 3’-end of the U6 promoter motivating sgRNA transcription. The array “CACC” guarantees that the oligo is matching with the overhangs of the BbsI-digested pX330-xCas9 plasmid. The base “G” is a prerequisite of RNA Polymerase III promoters and guarantees the effective startup of sgRNA transcription (e.g., appending the 5’-CACCG array to the protospacer of T1, achieving T1-F: CACCGTGCCTGCTCCTACTGGCCGC [25 nt]). The cleavage efficiency of 21 nt gRNA is significantly different from that of 20 nt gRNA1,12,13,14. It is generally advised to use 20 nt with 5'-G by generating a shorter protospacer, if this is not possible then 21 nt with extra 5'-G can be used. - Create a reverse complement (rc) of the protospacer sequence.
NOTE: For example, the rc of the T1 protospacer is GCGGCCAGTAGGAGCAGGCA (20 nt). - Append AAAC to the 5’-end of the rc protospacer sequence. Append an additional C to the 3’-end of the rc protospacer.
NOTE: The “AAAC” sequence ensures that the oligonucleotide is suitable for cloning into the BbsI-digested pX330-xCas9 plasmid. The additional“C”on the 3’-end is essential for annealing with the initiating “G” for the sgRNA transcription described above (e.g., AAACGCGGCCAGTAGGAGCAGGCAC [25 nt] is the final oligonucleotide sequence for T1 rc protospacer). - Order the oligonucleotides.
3. Oligonucleotide annealing
- Dilute lyophilized oligonucleotides to a final concentration of 10 μM in double distilled water (ddH2O).
- Mix forward and reverse oligonucleotides in a thin wall PCR tube maintaning a 1:1 ratio (e.g., 20 μL each) without adding any extra buffer.
- Incubate the mixture at 95 °C for 5 min and then ramp down the temperature to 72 °C for 10 min. Follow with a cooling period at room temperature (RT) consisting of simply removing the sample from the PCR machine and placing it at RT.
NOTE: It is not necessary to phosphorylate the oligonucleotide mixtures to facilitate the ligation.
4. sgRNA/CRISPR vector digestion
- Digest 1 μg of the selected pX330-xCas9 vector with BbsI (10 units of enzyme per 1 μg of plasmid) for 2 h at 37 °C (Figure 2). Conduct the digestion of the pX330-xCas9 sgRNA expression vector in a total volume of 50 μL, containing 5 μL of 10x digestion buffer and distilled water to achieve the final volume.
- Purify the digested vector by band extraction from a 2% agarose gel under 10 V/cm and subsequently purify using a silica column using a commercial gel extraction kit.
5. Ligation of the annealed sgRNA oligonucleotides to the expression vector
- Mix the annealed sgRNA oligonucleotides (5 μL of the mixture from step 4) with the BbsI-digested pX330-xCas9 vector (purified, 100 ng).
- Add 1 μL of ligase and 1 μL of 10x ligase buffer.
- Add distilled water with appropriate volume, up to 10 μL.
- Incubate overnight at 4 °C.
6. Competent cell transformation
- Take out E. coli DH5α competent cells from storage at -80 °C and thaw it on ice.
- Add 5 μL of the ligation mix to 50 μL of competent E. coli DH5α and keep the mixture on ice for 30 min.
- Heat shock the mixture at 42 °C for 90 s.
- Rest it on ice for 2 min.
- Recover the culture on a rotary shaker in 500 μL of LB media for 1 h at 37 °C.
- Plate 200 μL of the culture on an ampicillin resistance LB agarose plate and incubate it overnight at 37 °C.
7. Identification of the correct recombinant plasmids by PCR
- Choose 5 to 10 bacterial colonies from the LB plate and use each of them to inoculate one 1.5 mL tube containing 1 mL of LB media with 60 mg/mL ampicillin.
- Incubate the tubes on a rotary shaker for 2-3 h.
- Carry out detection of correct recombinant plasmids by using specific primer pairs for the sgRNA oligonucleotides [e.g., forward primer for T1 sgRNA expression vector construction (T1-F): CACCGTGCCTGCTCCTACTGGCCGC, reverse primer (BbsI-R): AAAGTCCCTATTGGCGTTAC, producing a 287 bp amplicon (Figure 3)].
- Prepare the PCR mixture (Table 2).
- Use the following PCR cycling conditions: 95 °C for 5 min for pre-denaturation; 30 cycles of 95 °C for 30 s for denaturation, 60 °C for 30 s for annealing, and 72 °C for 30 s for extension. After the 30 cycles are completed, perform a final extension step by heating for 5 min at 72 °C.
- Run the PCR product on a 2% agarose gel under 10 V/cm. A band at the right size [e.g., 287 bp] is considered as positive.
8. Validate the sequence of sgRNA expression plasmid
- Verify the sequence of PCR positive colonies by Sanger sequencing15 using the reverse primer BbsI-R (see step 7.3). This primer anneals at the site downstream of the sgRNA oligo insert. The pX330-xCas9-T1 expressing sgRNA sequence containing protospacer TGCCTGCTCCTACTGGCCGCGG and xCas9 was constructed.
- Use the forward primer T1-F to sequence the positive colonies. The forward one could not give complete sequence information for the site surrounding the insertion site of the sgRNA oligo, because 30-50 bp fragment following the sequencing primer could not be exactly read out.
9. Construction of dual luciferase reporter vector
- Synthesize 300-500 bp DNA fragments containing the sgRNA targets and subclone it into a dual luciferase reporter vector through double digestion [e.g., subclone 440 bp sheep DKK2 exon1 fragment into pSSA-Dual plasmid16,17 by using double digestion with AscI and SalI, resulting pSSA-Dual-DKK2].
- Synthesize 300-500 bp DNA fragments containing the sgRNA targets. Note that the sequences of the DNA fragments must be parts of the genomic sequences, which could be obtained from NCBI website (https://www.ncbi.nlm.nih.gov/) or related references.
- Select a suitable dual luciferase reporter vector such as pSSA-Dual16,17 (or two vectors expressing firefly luciferase and renilla luciferase respectively) and then digest this vector and the DNA fragments stated above with two endonucleases such as AscI and SalI.
- Finally ligate these two fragments with T4 DNA Ligase, resulting into pSSA-Dual-Target. Details for double digestion and ligation are displayed in Table 3 and Table 4, respectively. It is worth mentioning that the reporter vector should be amplified in stable bacteria strains (such as Top10), which are recommended to be cultured with lower rotational speed (typically no more than 200 rpm), for the purpose of avoiding DNA recombination.
10. Cell transfection
- Extract endotoxin-free plasmids for vectors stated above. Evaluate the purity and concentration of the plasmids by using suitable instruments. A final concentration of no less than 500 ng/μL and a purity ratio of 1.7-1.9 at absorbance 260/280 nm (A260/A280) are recommended for these plasmids.
- Co-transfect an appropriate cell line such as PIEC18 with the plasmids in equal proportion (e.g., pX330-xCas9-T1: pSSA-Dual-DKK2=1:1, 0.5 μg plasmid for each duplication when using 24 well plate). Use an empty vector such as pX330-xCas9 as a negative control.
- One day before the transfection, plate cells in an 24-well culture plate at a density of 2 x 105 cells/well. Cells will be ready for transfection when they achieve a confluence of 60-80%.
- Before the transfection time point, remove the supernatant as much as possible and gently add 0.5 mL of fresh media for each well.
- Dilute transfection reagent in DMEM media at the ratio of 1:25 and mix well.
- Prepare master mix of DNA by diluting 0.5 μg of DNA in 25 μL of DMEM media, and then add 1 μL of P3000 reagent.
- Add diluted DNA to each tube of diluted transfection reagent (1:1 ratio).
- Incubate for 10–15 min at room temperature.
- Add 50 μL of DNA-lipid complex to cells.
- Incubate cells for 24 h at 37 °C. Then, analyze transfected cells as in Step 11.
11. Dual-luciferase detection
- Prepare a sufficient quantity of the 1x passive lysis buffer (PLB) by adding 1 volume of 5x PLB to 4 volumes of distilled water and mix well.
- Passive lysis of cells cultured in 24-well culture plates.
- Remove the growth medium from the cultured cells, and gently apply a sufficient volume of phosphate buffered saline (PBS) to wash the surface of the culture vessel. Swirl the vessel briefly to remove detached cells and residual growth medium. Completely remove the rinse solution before applying PLB reagent.
- Dispense 100 μL of 1x PLB into each culture well to completely cover the cell monolayer. Let the culture plates stand for 20 min.
- Transfer the lysate to a tube or vial for further handling or storage.
- Prepare luciferase assay reagent (LAR) by resuspending the provided lyophilized luciferase assay substrate in 10 mL of the supplied luciferase assay buffer.
- Prepare an adequate volume to perform the desired number of dual-luciferase reporter assays (100 µL reagent per assay). Add 1 volume of 50x stop substrate to 50 volumes of stop buffer in a glass or siliconized polypropylene tube.
- Dual-luciferase reporter (DLR) assay
- Program luminometers to provide a 2-second pre-read delay, followed by a 10-second measurement period.
- Predispense 100 µL of luciferase assay reagent into the appropriate number of luminometer tubes to complete the desired number of DLR Assays.
- Carefully transfer up to 20 µL of cell lysate into the luminometer tube containing LAR; mix by pipetting 2 or 3 times. Place the tube in the luminometer and initiate reading.
- Remove the sample tube from the luminometer, add 100 µL of stop reagent and vortex briefly to mix. Replace the sample in the luminometer, and initiate reading.
- Record the renilla luciferase activity normalized to the firefly luciferase activity, namely the reciprocal of the ratio displayed on screen.
- Discard the reaction tube, and proceed to the next assay.
Representative Results
The methods outlined in this protocol are for the construction of sgRNA and xCas9 expression vectors and then for the optimization screening of sgRNA oligos with relatively higher gene targeting efficiencies. Here we display a representative example of 3 sgRNA targets to sheep DKK2 exon 1. SgRNA and xCas9 expressing vectors can be built by predigesting the vector backbone (Figure 2) followed by ligating it in a series of short double-strand DNA fragments through annealing oligo pairs. The positive colonies could be detected through specific primer pairs guided PCR (Figure 3). An 440 bp DNA fragment from sheep DKK2 exon 1 was subcloned into pSSA-Dual plasmid15 by using double digestion with AscI and SalI, resulting in pSSA-Dual-DKK2. The gene targeting capacities of pX330-xCas9-T1, pX330-xCas9-T2 and pX330-xCas9-T3 were simutanusly detected (Figure 5), and then the last sgRNA vector was identified as the relatively better one, which we pick up for sheep gene editing research at the next step.
This detection method combines a single strand annealing (SSA) mechanism (Figure 1B and Figure 4) with a luciferase report gene in order to monitor DNA cutting efficiency. As illustrated in Figure 4, SSA is a process initiated when a double strand break is made between two repeated sequences oriented in the same direction. Single strand regions are created adjacent to the breaks that extend to the repeated sequences so the complementary strands can anneal to each other. This annealing intermediate can be processed by digesting away the single stranded tails and filling in the gaps. The Dual-Luciferase Reporter gene mainly includes the luciferase genes from the firefly Photinus pyralis and from Renilla reniformis (also known as sea pansy). The activities of firefly and renilla luciferases are measured sequentially from a single sample. When the recognition area that has the termination codon is not cut in its middle, the gene in SSA system is blocked and cannot be translated into a functional protein. When the processing takes place, the SSA system will merge the homologous sequence automatically, and overlapping sequences becomes a single sequence, gene gets recombination repaired and can then be read throughout producing a functional protein.
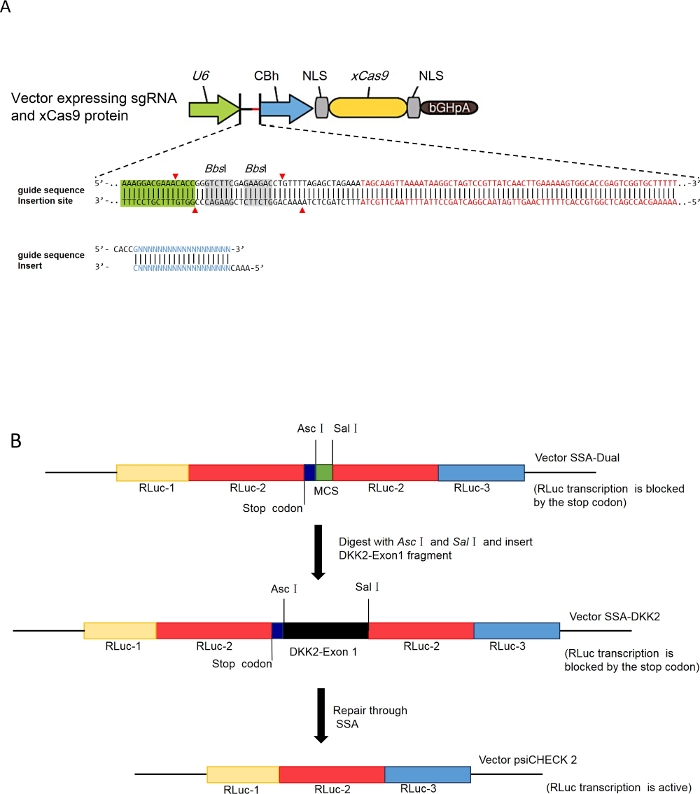
Figure 1: Schematic representation of the different steps of the cloning process.
(A) Schematic of sgRNA vector construction, adapted from the protocols of Feng Zhang Lab. (B) Schematic of dual luciferase reporter vector building (select vector pSSA-DKK2 as an example), DNA fragments Rluc-, Rluc-2 and Rluc-3 represent three parts of full-length coding sequence of Renilla luciferase. MCS represents “multiple cloning site”. Please click here to view a larger version of this figure.
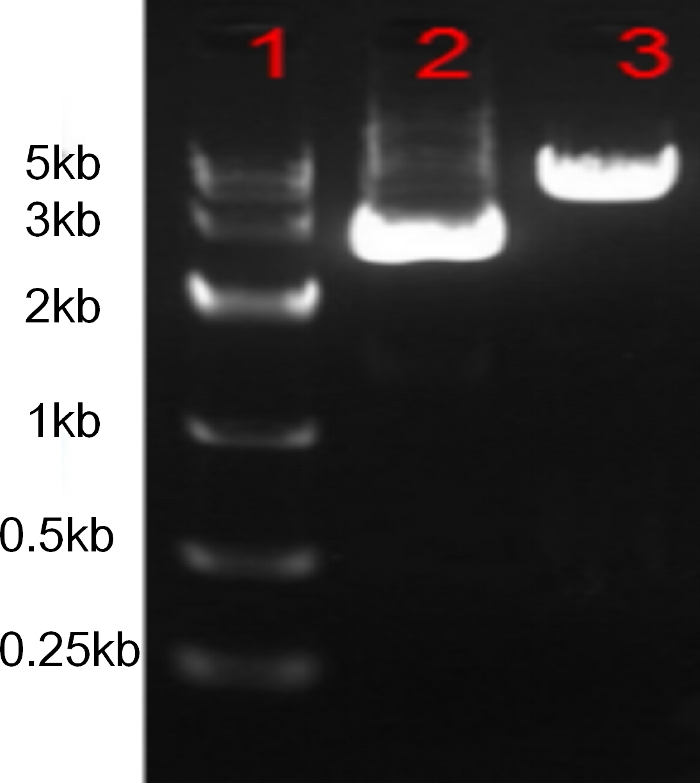
Figure 2: Predigesting the vector backbone pX330-xCas9 with BbsI.
1: DNA Marker, 2: pX330-xCas9 plasmid, 3: pX330-xCas9 plasmid digested with BbsI. Please click here to view a larger version of this figure.
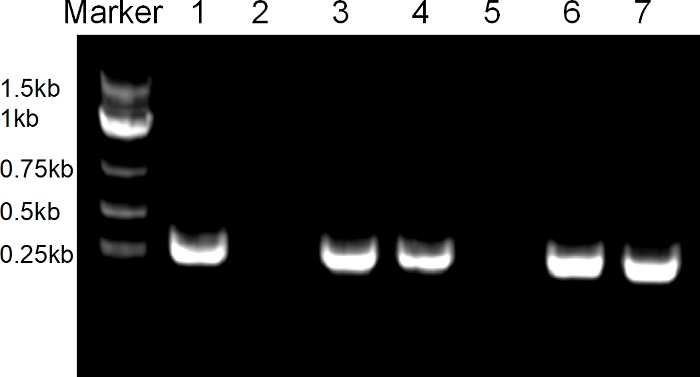
Figure 3: Specific primer pairs guided PCR for DKK2-T1 sgRNA vector detection.
Lanes 1-7 indicate PCR bands for 7 different bacteria colonies separately. Please click here to view a larger version of this figure.
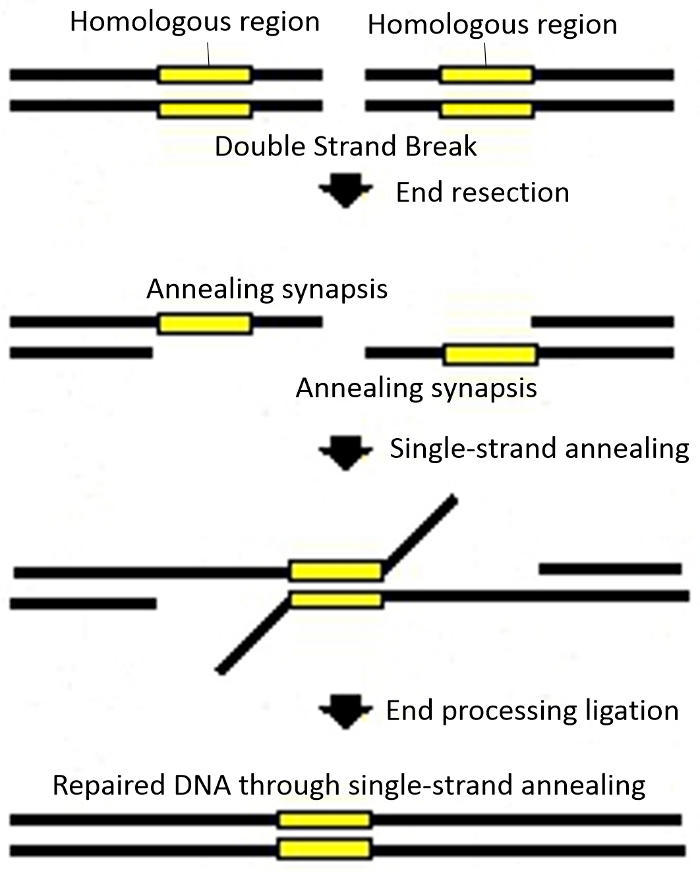
Figure 4: Schematics for single strand annealing (SSA). Please click here to view a larger version of this figure.
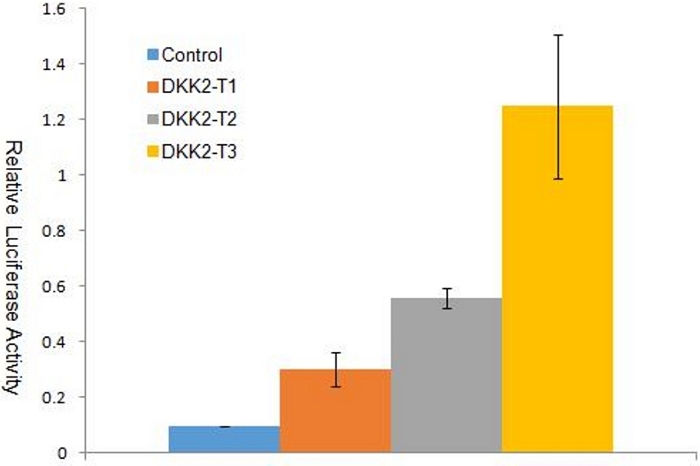
Figure 5: Dual luciferase assay with reporter vector pSSA-DKK2 in cell line PIEC.
The Ranilla luciferase luciferase activities are significantly induced (P<0.01, Student's t test) in PIEC by overexpresing of pX330-xCas9-T1, pX330-xCas9-T2 or pX330-xCas9-T3 with the fold change of 3.15, 5.84 or 13.10, respectively. The error bars indicate standard deviations (SD)(n=3) for each group. Please click here to view a larger version of this figure.
| Name | Genomic DNA Targets (5’-3’) | sgRNA Oligos (5’-3’) | ||
| T1 | TGCCTGCTCCTACTGGCCGCGG | T1-F: CACCGTGCCTGCTCCTACTGGCCGC | ||
| T1-R: AAACGCGGCCAGTAGGAGCAGGCAC | ||||
| T2 | ATCAAGTCCTCTCTGGGCGGGG | T2-F: CACCGATCAAGTCCTCTCTGGGCGG | ||
| T2-R: AAACCCGCCCAGAGAGGACTTGATC | ||||
| T3 | GCCCGCGAGCTGCCGAACTGTG | T3-F: CACCGCCCGCGAGCTGCCGAACTG | ||
| T3-R: AAACCAGTTCGGCAGCTCGCGGGC | ||||
Table 1: sgRNA targets and oligos designed for gene editing of sheep DKK2.
| PCR Component | 25 µL reaction |
| 10 µM Forward Primer (e.g. T1-F) | 1 µL |
| 10 µM Reverse Primer BbsI-R | 1 µL |
| 10x PCR Buffer | 2.5 µL |
| 2.5 mM dNTPs | 1 µL |
| bacterial fluid | 1 µL |
| DNA Taq Polymerase | 2.5 units |
| Nuclease-free water | Up to 25 µL |
Table 2: PCR mixture for positive bacterial colonies detection.
| Reagent | Volume |
| pSSA-Dual vector (synthesized DNA fragment or PCR product) | 1-2 μg |
| CutSmart Buffer | 5 μL |
| AscI | 1 μL |
| SalI-HF | 1 μL |
| Distilled water | up to 50 μL |
| Total volume | 50 μL |
Table 3: Double digestion of pSSA-Dual vector and synthesized DNA fragment (or PCR product).
| Reagent | Volume |
| pSSA-Dual vector(predigested) | 0.5 μg |
| DNA fragments containing the sgRNA targets (predigested) | 0.2 μg |
| T4 DNA Ligase Buffer (10x) | 1 μL |
| T4 DNA Ligase | 1 μL |
| Distilled water | up to 10 μL |
| Total volume | 10 μL |
Table 4: A ligation reaction of DNA fragments containing the sgRNA targets into a predigested pSSA-Dual vector.
| Step 1: reagent preparation | |
| Transfection reagent | 2 μL |
Table 5: Details of cell transfection in a 24 well plate.
Discussion
The sgRNA vector cloning procedures we have described here facilitates efficient production of sgRNAs, with most of the costs derived from the oligonucleotide ordering and vector sequencing. While the outlined method is designed to allow users to generate sgRNAs for use with CRISPR/Cas9, the protocol can easily be adapted for use with Cas9 orthologues or other RNA-guided endonucleases such as Cpf1, introducing minor modifications to the vector backbone and the oligonucleotide overhanging sequences.
The protocol outlined above will provide a preferred sgRNA target in 10 days when starting with appropriately designed oligonucleotides. This includes the sgRNA design (1 h, steps 1 – 2), dilution, aliquot and annealing of the oligonucleotides (30 min, step 3), digestion and purification of the sgRNA expression vector (3 h, step 4), the cloning of the sgRNA oligonucleotides into a predigested empty vector (overnight, steps 5 – 6), the PCR detection of the colonies (4-5 h, step 7) and the validation of the sequence of the sgRNA expression plasmid (24 h, step 8). Meanwhile, the construction and sequence validation of dual luciferase reporter vector takes 5 – 7 d (step 9). The preparation of the cell line, plasmid transfection and dual luciferase detection take about 48 h (steps 10 – 11). The failure rates for generating each plasmid are nearly 0.
The most critical step of the protocol is the restrictive enzyme digestion of sgRNA vector by enzymes such as BbsI. If the digestion is not sufficient, then the positive rate of bactera colonies might be very low. While the PCR-based detection approach could sensitively monitor the digestion efficiency and the positive rate. The forward primers, formerly used as forward oligos of the inserted sgRNA fragment, ensured highly effective distinction of right insertions and empty vectors. One of the advantages of this strategy is precisely the efficient detection of the positive colonies by PCR before DNA sequencing. As stated above, sequence verification is still relatively more expensive compared to PCR, especially when incomplete digestion of BbsI occurs. In terms of resource saving, we paired the forward oligos for sgRNA fragment annealing (such as T1-F, T2-F or T3-F) with the sequencing primer BbsI-R for PCR amplification. More than 500 sgRNA vectors have been successfully constructed in the lab through this strategy for colony identification.
Another merit of the protocol is the effective pre-selection of candidate sgRNA targets using dual luciferase reporter system. Huge variance of cutting efficiency indeed exists at different sgRNA targets4. To identify a highly efficient sgRNA target by T7E1 assay or Surveyor nuclease assay1 in a difficult-to-transfect cell line is time and labor consuming. In addition, for PCR amplification in the T7E1 assay or Surveyor nuclease assay, it is sometimes difficult to specifically amplify the target sequence in the genome DNA because of the lack of suitable primer pairs1. In contrast, the dual luciferase reporter assay is independent of genome amplification procedures. Previously we have used this method to get highly active sgRNAs16,19.
Despite the clear advantages of the method described in this work, there are also some limitations that must be pointed out. Although it ensures a high rate of success, the luciferase-based method for sgRNA selection proposed could be more expensive than other approaches (surveyor/high-resolution melt analysis, etc.) due to the need to synthesize the target sequence. In addition, this approach is not faster than either of these alternative methods due to the need to clone the reporter plasmid if an easy-to-transfect cell line will be conducted for gene targeting. It is also necessary to clarify that the method is suitable for making knockouts but not to edit specific nucleotides. Additionally, only double-stranded breaks and not single-stranded breaks can be evaluated.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This project was funded by First Class Grassland Science Discipline Program of Shandong Province (China), National Natural Science Foundation of China (31301936, 31572383), the Special Fund for Agro-scientific Research in the Public Interest (201403071), National risk assessment major special project of milk product quality and safety (GJFP201800804) and Projects of Qingdao People's Livelihood Science and Technology (19-6-1-68-nsh, 14-2-3-45-nsh, 13-1-3-88-nsh).
Materials
| A new generation of full touch screen gradient PCR instrument | LongGene | A200 | Target gene amplification |
| AscI restriction enzymes | New England Biolabs | R0558V | Cutting target vectors |
| BbsI restriction enzyme | New England Biolabs | R0539S | Cutting target vectors |
| Clean workbench | AIRTECH | SW-CJ-2FD/VS-1300L-U | A partial purification device in the form of a vertical laminar flow, which creates a local high clean air environment |
| DH5α Competent Cells | TaKaRa | K613 | Plasmid vector transformation |
| Dual-Luciferas Reporter Assay System | Promega | E1910 | Dual-luciferas reporter assay |
| Electric thermostatic water bath | Sanfa Scientific Instruments | DK-S24 | Heating reagent by constant temperature in water bath |
| Electrophoresis | Beijing Liuyi Biotechnology Co., Ltd. | DYY-6C | Control voltage, current, etc. |
| Eppendorf Reference 2 | Eppendorf China Ltd. | Reference 2 | Accurately draw and transfer traces of liquid |
| Gel imaging analyzer | Beijing Liuyi Biotechnology Co., Ltd. | WD-9413B | For the analysis of electrophoresis gel images |
| GloMax 20/20 Luminometer | Promega | E5311 | Detect dual luciferase activity |
| High speed refrigerated centrifuge | BMH | sigma 3K15 | Nucleic acid extraction and purification |
| Intelligent biochemical incubator | Sanfa Scientific Instruments | SHP-160 | Provide a suitable temperature environment for the enzyme digestion experiment |
| LB Broth Agar | Sangon Biotech | A507003-0250 | For the cultivation of E.coli |
| Lipofectamine 3000 Transfection Reagent Kit | Thermo Fisher | L3000015 | DNA Transfection |
| SalI restriction enzymes | New England Biolabs | R3138V | Cutting target vectors |
| SanPrep Column DNA Gel Extraction Kit | Sangon Biotech | B518131-0050 | Recycling DNA fragments |
| SanPrep Column Plasmid Mini-Preps Kit | Sangon Biotech | B518191-0100 | Extraction of plasmid DNA |
| T4 DNA Ligase | New England Biolabs | M0202V | Link DNA fragment |
| TaKaRa MiniBEST DNA Fragment Purification Kit Ver.4.0 | TaKaRa | 9761 | DNA purification |
| Vertical pressure steam sterilizer | JIBIMED | LS-50LD | High temperature and autoclave to kill bacteria, fungi and other microorganisms in laboratory equipment |
| Water bath thermostat | Changzhou Guoyu Instrument Manufacturing Co., Ltd. | SHZ-82 | Let the bacteria keep shaking, which is good for contact with air. |
Riferimenti
- Zhang, H., et al. A surrogate reporter system for multiplexable evaluation of CRISPR/Cas9 in targeted mutagenesis. Scientific Reports. 8 (1), 1042 (2018).
- Nageshwaran, S., et al. CRISPR Guide RNA Cloning for Mammalian Systems. Journal of Visualized Experiments. (140), (2018).
- Dang, Y., et al. Optimizing sgRNA structure to improve CRISPR-Cas9 knockout efficiency. Genome Biology. 16, 280 (2015).
- Doench, J. G., et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nature Biotechnology. 34 (2), 184-191 (2016).
- Moreno-Mateos, M. A., et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nature Methods. 12 (10), 982-988 (2015).
- Joung, J., et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nature Protocols. 12 (4), 828-863 (2017).
- Yang, L., Yang, J. L., Byrne, S., Pan, J., Church, G. M. CRISPR/Cas9-Directed Genome Editing of Cultured Cells. Current Protocols in Molecular Biology. 107, 1-17 (2014).
- Yang, L., Mali, P., Kim-Kiselak, C., Church, G. CRISPR-Cas-mediated targeted genome editing in human cells. Methods in Molecular Biology. 1114, 245-267 (2014).
- Vidigal, J. A., Ventura, A. Rapid and efficient one-step generation of paired gRNA CRISPR-Cas9 libraries. Nature Communications. 6, 8083 (2015).
- Mazon, G., Mimitou, E. P., Symington, L. S. SnapShot: Homologous recombination in DNA double-strand break repair. Cell. 142 (4), 646 (2010).
- Doench, J. G., et al. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nature Biotechnology. 32 (12), 1262-1267 (2014).
- Lee, J. K., et al. Directed evolution of CRISPR-Cas9 to increase its specificity. Nature Communications. 9 (1), 3048 (2018).
- Sung, Y. H., et al. Highly efficient gene knockout in mice and zebrafish with RNA-guided endonucleases. Genome Research. 24 (1), 125-131 (2014).
- Ran, F. A., et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell. 154 (6), 1380-1389 (2013).
- Sanger, F., Nicklen, S., Coulson, A. R. DNA sequencing with chain-terminating inhibitors. Proceedings of the National Academy of Sciences of the United States of America. 74 (12), 5463-5467 (1977).
- Ruan, J., et al. Highly efficient CRISPR/Cas9-mediated transgene knockin at the H11 locus in pigs. Scientific Reports. 5, 14253 (2015).
- Li, K., et al. An plasmid with double fluorescent groups and its application as standard substance. China patent. , (2012).
- Li, H., et al. Characterization of the porcine p65 subunit of NF-kappaB and its association with virus antibody levels. Molecular Immunology. 48 (6-7), 914-923 (2011).
- Li, H., et al. A pair of sgRNAs targeting porcine RELA gene. China patent. , (2015).

