Engineering and Characterization of an Optogenetic Model of the Human Neuromuscular Junction
Summary
We describe a reproducible, automated, and unbiased imaging system for characterizing neuromuscular junction function using human engineered skeletal muscle tissue and optogenetic motoneurons. This system allows for the functional quantification of neuromuscular connectivity over time and detects diminished neuromuscular function caused by neurotoxins and myasthenia gravis patient serum.
Abstract
Many neuromuscular diseases, such as myasthenia gravis (MG), are associated with dysfunction of the neuromuscular junction (NMJ), which is difficult to characterize in animal models due to physiological differences between animals and humans. Tissue engineering offers opportunities to provide in vitro models of functional human NMJs that can be used to diagnose and investigate NMJ pathologies and test potential therapeutics. By incorporating optogenetic proteins into induced pluripotent stem cells (iPSCs), we generated neurons that can be stimulated with specific wavelengths of light. If the NMJ is healthy and functional, a neurochemical signal from the motoneuron results in muscle contraction. Through the integration of optogenetics and microfabrication with tissue engineering, we established an unbiased and automated methodology for characterizing NMJ function using video analysis. A standardized protocol was developed for NMJ formation, optical stimulation with simultaneous video recording, and video analysis of tissue contractility. Stimulation of optogenetic motoneurons by light to induce skeletal muscle contractions recapitulates human NMJ physiology and allows for repeated functional measurements of NMJ over time and in response to various inputs. We demonstrate this platform's ability to show functional improvements in neuromuscular connectivity over time and characterize the damaging effects of patient MG antibodies or neurotoxins on NMJ function.
Introduction
The neuromuscular junction (NMJ) is the chemical synapse between motoneurons (MNs) and skeletal muscle cells (SkM) that allows for muscle contraction. Toxins, such as neurotoxin α-bungarotoxin (BTX), or neuromuscular diseases (NMD) like myasthenia gravis (MG) can lead to degeneration of the NMJ and reductions in muscle control1. Bioengineered human tissue models better recapitulate the functional and physiological mechanisms of human NMJs and offer greater translational potential than animal models.
While animal models have advanced the understanding of the formation and function of the NMJ, there are significant differences between human and animal synapses that limit the translation of results to humans and make in vivo characterization of the NMJ challenging2,3,4. Studies have shown distinct physiological differences between mouse and human NMJs. Mice have larger NMJs and smaller active zone densities when compared to human NMJs4. Additionally, drug studies conducted in animal models do not always reflect the effects found in human clinical trials. Engineered human tissue models provide the opportunity to study the healthy development of the NMJ and the pathology of neuromuscular diseases and allow for drug screenings. Human induced pluripotent stem cells (hiPSCs)5 can be differentiated into a variety of cell types, including skeletal muscle cells6,7 and motoneurons8,9. hiPSCs can be generated easily from patient cells, allowing for better disease modeling10 and drug screening11,12 through patient-specific tissue models.
Two-dimensional (2D) monolayer co-cultures of SkMs and MNs lack the morphology, phenotype, organization, and functional behavior of physiological NMJs. NMJs randomly form in 2D culture, which inhibits the isolation of motor units for analysis, limits accurate functional measurements, and prevents their use for repeated, systematic experiments13. Three-dimensional (3D) tissue models of NMJs overcome many of these limitations, recapitulating the morphological and functional characteristics of physiological NMJs7,14,15,16,17. Using this model, the two tissue types are developed separately and then integrated by directing axonal growth, allowing for more organized NMJs to develop compared to 2D culture systems.
Our previous study demonstrated that combining optogenetics with tissue engineering can allow for accurate non-invasive stimulation and evaluation of NMJ function18,19. Through genetic engineering, light-sensitive proteins can be integrated into the genome of hiPSCs. Integrating channelrhodopsin-2 (ChR2), an ion channel that opens in response to blue light, into the membrane of excitable cells such as neurons allows for non-contact spatiotemporal control over cell activation20,21,22. hiPSCs carrying ChR2 can be differentiated into optogenetic motoneurons sensitive to blue light, removing the need for typical invasive electrodes that stimulate neurons and avoiding unwanted stimulation of the muscle cells by electrodes23. This system uses optogenetic motoneurons to stimulate contractions in non-optogenetic skeletal muscle cells. Combining video acquisition and controlled blue light illumination allows for the co-cultured tissues to be simultaneously stimulated and recorded for NMJ function.
MG is caused by autoantibodies targeting nicotinic acetylcholine receptors (AChR), which results in decreased NMJ function and muscle weakness24. It is diagnosed based on presented symptoms, electrodiagnosis, and detection of autoantibodies via serological blood tests. However, not all autoantibodies involved in MG have been identified, and some seronegative patients are diagnosed with MG but with no recognized antibodies25,26. Our system allows for repeated functional assessment of the NMJ before and after the addition of serum from MG patients, providing invaluable insight into the functional and biochemical changes caused by the MG antibodies18. Our protocol illustrates how to produce 3D in vitro models of functional human NMJ that can be used to diagnose and investigate NMJ pathologies and test potential therapeutics. We demonstrate the versatility of the system in two platforms, a microfluidic device, and a larger open-well bioreactor platform.
Protocol
All cell lines for this work were created and used in compliance with the institutional guidelines of Columbia University, NY, USA.
1. Bioreactor preparation
- Make bioreactor molds
- Download a bioreactor CAD file from the Supplementary CAD File or create a custom own design.
- Generate a CNC toolpath from the 3D model using CAM software.
- Machine acetal molds using a CNC milling machine.
- Fabrication of bioreactors
- Mix a 10:1 base to curing agent mixture of polydimethylsiloxane (PDMS, 77 g of mixture per 4 platforms/molds).
- Place the mixture into a vacuum chamber, close all the valves, turn on the vacuum, and de-gas the mixture for at least 30 min until no air bubbles remain. Pour the mixture into molds and de-gas the molds in the vacuum chamber for 1 h.
- Close the molds with the top half of the mold in the correct orientation. Place a steel hexagonal rod over the center and clamp on both sides.
- Refill the top with PDMS.
- Cure the molds in a 65 °C oven for at least 4 h.
- Remove the platforms from the molds after cooling down to room temperature.
- Clean the devices in an ultrasonic bath using 1 h cycles of dish soap, 400 mL of 100% isopropanol, and distilled water.
- Dry overnight at 65 °C.
- Soak glass coverslips in 1% nonionic surfactant polyol for 30 min, and ensure that the coverslips do not stack so that they are all properly coated.
- Rinse well with distilled water and dry overnight at 65 °C.
- Use a plasma cleaner on high with 6 L/min of oxygen for 1-2 min to the treat glass coverslips and PDMS platform. Once treated, bond the two together by pressing the PDMS down onto the coverslip for at least 30 s.
- Autoclave the bonded devices before adding cells.
2. Building an optical stimulation setup
- Using a 30 mm cage cube system, attach a 573 nm dichroic mirror in the center. Couple a red 627 nm LED with a 594 nm long-pass excitation filter and attach it to the top side of the cube, with the LED facing the mirror. Then attach a blue 488 nm LED with a 546 nm short-pass excitation filter to the adjacent side of the cube, with the LED facing the mirror as shown in the schematic in Figure 3A.
- Attach a ring-actuated iris diaphragm to the bottom of the cage cube to control the size of the illuminated area.
- Power each LED by a T-Cube LED driver and control via an Arduino Uno Rev3 board. Connect the side input of the LED driver to a power source plug, connect the middle output to the Arduino, and connect the side output directly to the LEDs.
- Connect the Arduino Uno to a PC by a USB cable.
- Download files from GitHub link (https://github.com/ofvila/NMJ-function-analysis).
- Open the "Optical_Stimulation_Ramp_Arduino.ino" file from GitHub folder.
- Set starting parameters: Steps = 30; startFrequency = 0.5; endFrequency = 3; pulseLength = 100.
- Ensure that pin output 4 is assigned to the Blue LED driver and pin output 2 is assigned to the Red LED driver. Check the middle wires of the LED driver are connected to the ground and to the respective pin output.
- Compile and load the "Optical_Stimulation_Ramp_Arduino.ino" program to Arduino.
- Connect each LED driver to its corresponding channel.
3. Cell culture setup (day -21-0)
- Primary skeletal muscle cells
- Thaw and expand primary skeletal myoblasts (obtained from Cook Myosite) for a maximum of six passages using Myotonic growth medium (+ supplement). Maintain cells in an incubator set to 37 °C and 5% CO2.
- Change the media every 2 days.
- Once cells are about 60% confluent, passage them using 0.05% Trypsin-EDTA (1x, for 5 min at 37 °C). Collect the dissociated cells and add fresh media to neutralize the trypsin.
- Spin down the cells at 300 x g and aspirate the supernatant above the cell pellet.
- Resuspend the cells with MGM and seed 1:3 with 60 mL per triple-layer flask.
- ChR2-hiPSC
NOTE: All cell lines for this work were created and used in compliance with the institutional guidelines of Columbia University, NY, USA. In this protocol, ChR2-expressing hiPSCs were generated via CRISPR-Cas9 genome editing using previously described methods18, but any stable optogenetic cell lines (lentiviral, piggyBac, etc.) can be used in the same way. Constitutive promoters expressed in both iPSCs and motoneurons were chosen (CAG). The cells were found to have healthy karyotype, as noted in previous publications18, 19.- Coat 6-well plates with 1 mL/well of solubilized basement membrane matrix diluted in DMEM/F12 (1:80) and incubate the plates at room temperature for 1 h.
- Seed iPSCs on coated 6-well plates with 2 mL of feeder-free cell culture medium (iPSC media), exchanging 2 mL of media every other day and passaging every 5-7 days. Maintain cells in an incubator set to 37°C and 5% CO2.
- Passaging
- Dissociate the stem cells by incubating with 1 mL of enzyme-free stem cell releasing reagent for 4 min and mechanically shearing with a wide tip P1000 pipette.
- Seed cells at a ratio of 1:24 or 1:48 in iPSC media with 2 μM of Y-27632 dihydrochloride in 2 mL/well onto coated 6-well plates.
4. Skeletal muscle tissue seeding (day -3)
- Isolate and count 30 x 106 myoblasts using an automated cell counter.
- Resuspend the myoblasts in a 4:1 mix of 3 mg/mL collagen I and solubilized basement membrane matrix (800 μL of collagen mixture + 200 μL of basement membrane matrix to reach a final volume of 1 mL). For the collagen component, add the accompanying neutralizing agent (9:1 mix of collagen to neutralizing agent) and dilute the mixture with 1x PBS to achieve a volume of 800 μL.
- Add 15 μL of cell-collagen suspension to each muscle chamber of the bioreactor, being sure to spread the suspension across both pillars using the pipette tip (Figure 2).
- Let the cell-gel mixture polymerize at 37 °C for 30 min and then fill each bioreactor well with 450 μL of Myotonic growth media.
- After 3 days (once the gel has compacted), begin myotube differentiation.
5. Myotube differentiation (days 0-14)
- Begin myotube infusion by switching tissues to 450 μL of fusion inducing media (FS, Table 1) for 7 days, changing the media every other day.
NOTE: Try to begin myotube differentiation on the same day as motoneuron differentiation from hiPSCs in order to seed motoneurons at the end of both cell types' differentiation schedules. - On day 7 change the media to 450 μL of maturation media Ia (MMIa, Table 1).
- On day 9, change to 450 μL of maturation media Ib (MMIb, Table 1).
- On day 11, change the media to 450 μL of NbActiv4. Keep changing the media every 2 days until the motoneurons are seeded (Step 7).
6. Motoneuron differentiation (days 0-14)
NOTE: Our motoneuron differentiation protocol was adapted from Maury et al8.
- On day 0, transfer 4 x 106 ChR2- hiPSCs to an ultra-low attachment Petri dish with 15 mL of motoneuron suspension culture medium (MSCM, Table 1). Supplement MSCM with 3 μM CHIR99021, 0.2 μM LDN193189, 40 μM SB431542 hydrate and 5 μM Y-27632 dihydrochloride.
- On day 2, isolate the neurospheres (NS) with a 37 μM reversible strainer and replate in 15 mL of MSCM with 3 μM CHIR99021, 0.2 μM LDN193189, 40 μM SB431542 hydrate, and 0.1 μM retinoic acid. NS should be visible in the Petri dish without using a microscope after day 2.
- On day 4, transfer the cells and media to a 50 mL tube and allow the neurospheres to settle to the bottom (5 min). Aspirate the supernatant and resuspend the cells in 15 mL of MSCM supplemented with 0.5 μM SAG, 0.2 μM LDN193189, 40 μM SB431541, and 0.1 μM retinoic acid.
- On day 7, repeat Step 6.3. but resuspend the cells in 15 mL of MSCM supplemented with 0.5 μM SAG and 0.1μM retinoic acid.
- On day 9, repeat Step 6.3. but resuspend cells in 15 mL of MSCM supplemented with 10 μM DAPT.
- On day 11, repeat Step 6.3. but resuspend the cells in 15 mL of MSCM supplemented with 20 ng/mL BDNF and 10 ng/mL GDNF.
- On day 14, seed the motoneurons into the platform.
7. Seeding motoneurons aggregates in bioreactor (day 14)
- Prepare a 4:1 gel mixture of 2 mg/mL collagen I and Matrigel (800 μL of collagen mixture + 200 μL of Matrigel to reach a final volume of 1 mL). For the collagen component, add the accompanying neutralizing agent (9:1 mix of collagen to neutralizing agent) and dilute the mixture with 1x PBS to achieve a final volume of 800 μL.
- Use a 400 nm cell strainer to select large neurospheres and resuspend them in the gel mixture.
- Aspirate media from the reservoir and carefully from the neurosphere well (Figure 2).
- Add 15 μL of gel mixture into the neurosphere channel.
- Load a 10 μL pipette with 10 μL of gel and then pick one neurosphere.
- Deposit the NS into the neurosphere channel and ensure that the NS is in the chamber. Slowly raise the pipette while releasing the remaining gel once the NS is deposited. If unsure that the NS was correctly deposited, check its location using a microscope.
- Allow the gel to polymerize for 30 min at 37 °C.
- Add 450 μL of NbActiv4 supplemented with 20 ng/mL BDNF and 10 ng/mL GDNF to the reservoirs.
- Change the media every other day to allow for axonal growth from NS to muscle tissue.
8. Simultaneous optical stimulation and video recording of NMJ function (day 24+)
- For imaging, use an inverted fluorescent microscope with a scientific complementary metal-oxide-semiconductor (sCMOS) camera.
- Set the camera software binning to 2×2, exposure to 20 ms, rolling shutter ON, readout rate to 540 MHz, dynamic range: 12-bit & gain 1, and sensor mode: overlap.
- Use 2x objective on the microscope to image the microtissues.
- Attach a live-cell chamber (37 °C, 5% CO2) to the microscope stage.
- Select the region of interest (ROI) that contains the innervated skeletal tissues tissue to minimize the file size and processing time.
- Place a 594 nm long-pass emission filter between the sample and the imaging objective to filter out blue light pulses from the camera.
- Place a rectangular 4-well plate containing 4 bioreactors (24 tissues) into the live-cell chamber.
- Click Live View. Center and focus the image with the desired ROI.
- Upload the custom macro code from the GitHub folder (https://github.com/ofvila/NMJ-function-analysis) to control the stage position, the Arduino board, and video acquisition.
- Set output movie as: day_tissue group_tissue name_experiment.nd2.
- Run the macro code with the desired X,Y coordinates set on the stage and acquire a fast time lapse with 1700 frames at 50 frames/s.
- Replace the media after imaging and return the samples to the incubator. Allow at least 24 h between image acquisition sessions to avoid tissue fatigue.
9. Batch processing and analysis (day 24+)
- Movie processing
- Use the custom MATLAB code to process the videos in batch analysis. The files can be downloaded from the GitHub folder (https://github.com/ofvila/NMJ-function-analysis). The functions are listed and explained in Table 2.
NOTE: The code is compatible with .nd2 and .czi formats. It requires parallel processing to be activated in MATLAB and needs the bioformats package. - Run the recursiveOSAnalysis script to analyze the movies through parallel processing. Have all of the acquired videos in the same folder and select that folder when running the code.
- Adjust the post-analysis parameters as needed.
- Change baselineTime (option 1) if spontaneous contraction is picked up at beginning of recording. This will show an initial reading way above 0 and will need the frame to be shifted to compensate.
- Change peakThreshold (option 2) if the contractions are not being registered. The code will detect contractions that are above 25% of the highest peak by default so that this value can be changed.
- Change minMinProminence and minMinWidth to adjust the sensitivity when detecting the start of each peak.
- Generate video and graph output.
- Run recursiveOSMovie to generate a video file for each tissue with its respective contractility trace.
- Use the custom MATLAB code to process the videos in batch analysis. The files can be downloaded from the GitHub folder (https://github.com/ofvila/NMJ-function-analysis). The functions are listed and explained in Table 2.
10. Perturbation of NMJ function (day 24+)
- Prepare treatment media by adding external effector to NbActiv4 basal media supplemented with 20 ng/mL BDNF and 10 ng/mL GDNF.
- For the MG sera experiment, use 20% MG patient sera in NbActiv4 + BDNF + GDNF.
- For the BTX experiment, use 5 μg/mL BTX in NbActiv4 + BDNF + GDNF.
- Stimulate/image using Step 3.2. to record the baseline.
- Replace the media in tissues with treatment media (450 μL/tissue).
- Incubate for the desired time (48 h for patient sera, 20 min for BTX).
- Stimulate/image again using Step 3.2. to record function after treatment.
- Remove the treatment media and allow the tissues to rest for the desired time (48 h after sera removal)
NOTE: Allow at least 24 h between stimulation and imaging to avoid tissue fatigue
Representative Results
Neuromuscular junctions were generated by co-culturing optogenetic hiPSC-derived motoneurons with non-optogenetic skeletal muscle tissue. Human primary skeletal myoblasts (SkM) were seeded into the platforms and differentiated into multinucleated myotubes using the 2-week protocol. The optogenetic motoneurons were differentiated separately, but in parallel with the myotube differentiation, and then seeded into the platform (Figure 1). The tissues began contracting in response to blue light stimulation 7-12 days after MN seeding, indicating successful development and maturation of the NMJs. The NMJs can be cultured, stimulated, and imaged for up to 40 days after motoneuron seeding, but the optimal time point occurs on day 1619. The morphological and biochemical validation of the differentiations and presence of the optogenetic proteins are shown in Supplementary Figure 1 and have been validated in previous studies18,19.
To demonstrate that the protocol can be easily adapted to different culture systems, the strategy was tested using two different compartmentalized reactors: a microfluidic device and an open-well bioreactor. Both systems have a similar design, with one chamber provided with pillars for the generation of skeletal muscle tissues and a second chamber for the culture of 3D motoneuron aggregates that can extend axons to innervate the skeletal tissues (Figure 2A,B). However, the two systems have different scales, with the microfluidic device yielding 1 mm long skeletal muscle tissues comprising approximately 20,000 myoblasts (Figure 2C) and the open-well bioreactor system resulting in 4 mm long muscle tissues comprising approximately 450,000 myoblasts (Figure 2D).
To allow for controlled stimulation of the optogenetic NMJs, an optical setup was built, comprising of a red 637 nm LED for brightfield illumination, a blue 488 nm LED for activation of ChR2-MNs, and a blue light filter between the tissue sample and objective to prevent blue light from interfering with the image analysis (Figure 3A). The LEDs were powered by LED drivers and controlled precisely via an Arduino microprocessor. NMJ function was evaluated by simultaneously acquiring time-lapse videos while stimulating MNs with blue light using a custom microscope macro-script paired with the Arduino code (Figure 3B). The protocol increased the frequency of stimulation from 0.2 Hz to 2 Hz in 30 steps. Once the movies were acquired, the tissue function was analyzed using a custom MATLAB package (Figure 3C). The analysis measured tissue displacement, generated a contractility trace, and synchronized it with the light stimulation protocol. It then determined the fraction of effective pulses (F) and an expected fraction of effective pulses (E) to account for potential unstimulated contractions. The expected fraction (E) was calculated as the fraction of pulses that would be labeled as effective if the contractions were randomly distributed within the 30 s. The tissue score was then calculated as (F-E)/(1-E), with a value between 0 and 1 (Figure 3C). The code determines if a contraction is triggered based on the time between a light pulse and the start of the contractility peak, as shown in Figure 3D. This automated, unbiased method allows for repeated characterization of NMJ function in large sample sizes and over time. The code includes tunable parameters that can be adjusted after processing to ensure correct scoring (Supplementary Figure 2).
The system's ability to quantitatively characterize NMJ function over time was demonstrated by imaging a group of tissues for 11 days (day 9 to day 20 after motoneuron seeding). The system captured improvement of NMJ function in tissues, showing an increase in triggered responses 1 week after tissue fabrication (Figure 4A), followed by a stable function for an additional week (Figure 4B). To verify that muscle stimulation occurred through the NMJs, another group of tissues was analyzed before and after 20 min of incubation with 5 μg/mL of the neurotoxin α-bungarotoxin (BTX), which binds specifically and irreversibly to acetylcholine receptors. BTX stopped all triggered and spontaneous contractions, resulted in a complete disruption of NMJ function, and demonstrated that light stimulation of the tissues requires a functional NMJ (Figure 4C,D). Serial dilutions of BTX were initially tested to identify the optimal BTX concentration (results not shown).
To evaluate the translational capability of the system, NMJ function was assessed before and after the addition of serum from five myasthenia gravis patients. Analysis of the tissues treated with 20% MG serum for 48 h showed decreased function compared to control tissues treated with serum from healthy donors (Figure 4E,F). The engineered NMJs were also evaluated again 48 h after removal and washout of the serum and showed recovery of function (Figure 4F). Additionally, increasing concentrations of patient serum and normal human serum control (5%, 20%, 40%) were tested to identify the optimal concentration that affected tissue function. The results showed the ability of the system to characterize and quantify NMJ function and to model myasthenia gravis in vitro.
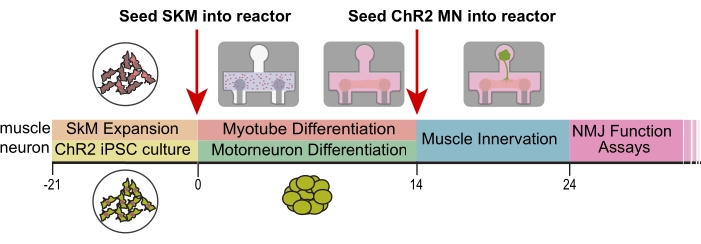
Figure 1. Experimental design and timeline. Timeline (days) of differentiation and seeding into platforms. SkM = skeletal muscle, ChR2 = optogenetic motoneurons, NMJ = neuromuscular junction. This figure has been modified with permission from Vila et al.18. Please click here to view a larger version of this figure.
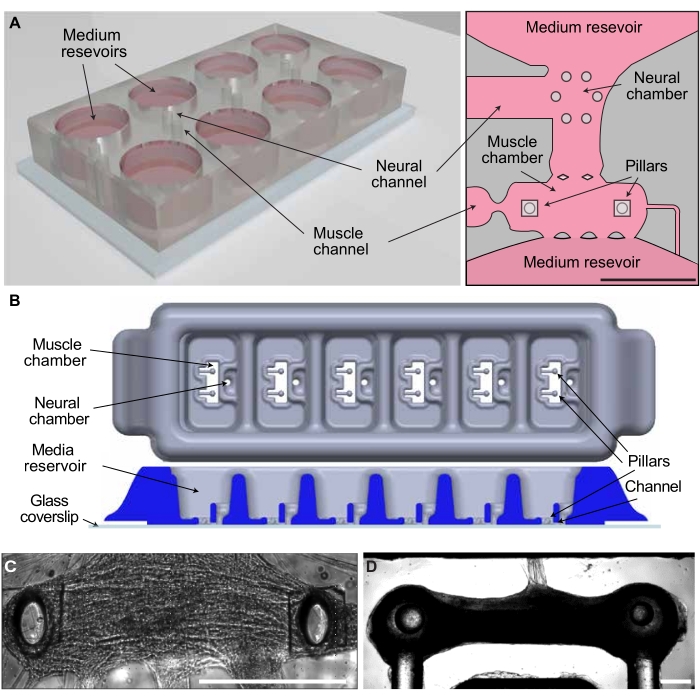
Figure 2. Co-culture systems for the generation of 3D NMJs. (A) Schematic of the microfluidic platform. (B) Schematic of the open-well PDMS bioreactor. (C) Innervated skeletal muscle tissue in the microfluidic platform. (D) Innervated tissue in the open-well bioreactor. Scale bar 0.5 mm. This figure has been modified with permission from Vila et al.18. Please click here to view a larger version of this figure.
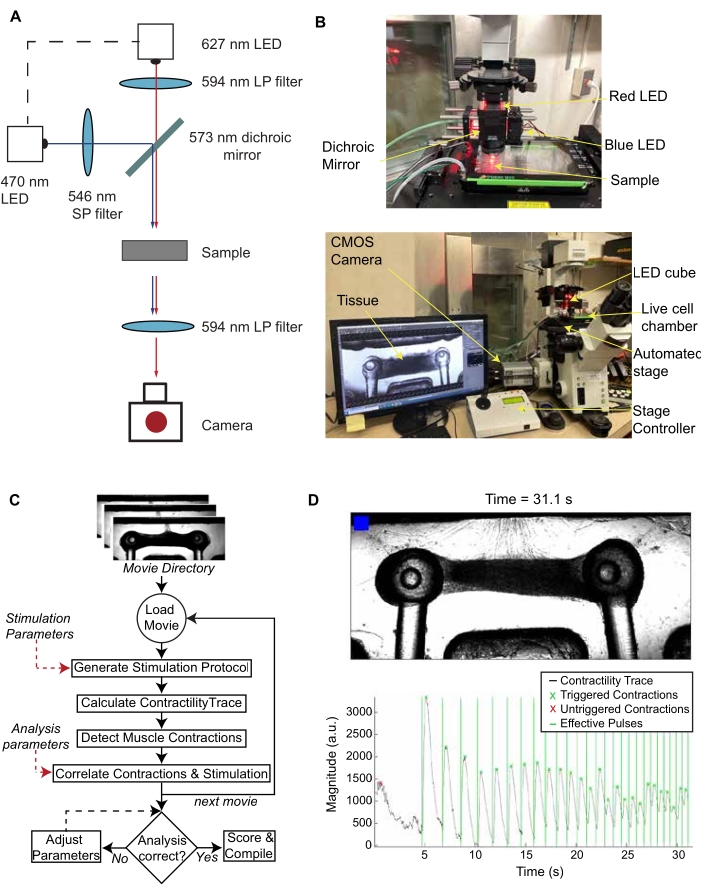
Figure 3. Stimulation, recording, and analysis of NMJ function. (A) Schematic of LED set up and light path. (B) Optical set-up with an inverted microscope, live cell chamber, automated stage, and sCMOS camera. (C) Algorithm for the batch processing of optical stimulation videos of NMJ cultures. (D) Representative frame of the video generated by the MATLAB code and the corresponding contractility trace graph. The blinking blue box in the video and the vertical lines in the graph indicate when the blue light stimulation occurs. This figure has been modified with permission from Vila et al.18. Please click here to view a larger version of this figure.
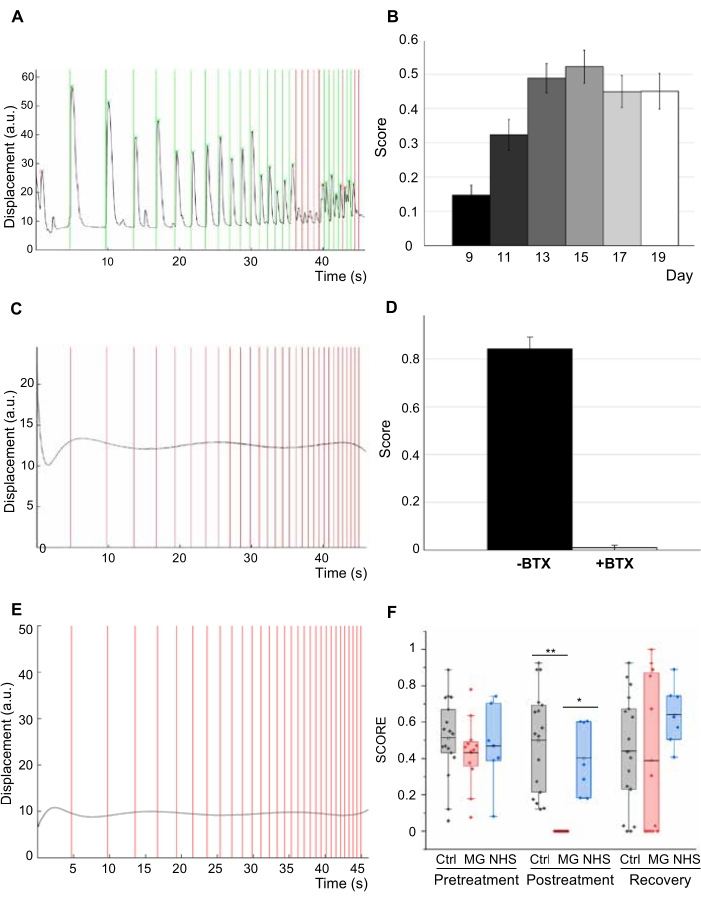
Figure 4. NMJ function. (A) Contractility trace for 3D engineered human NMJs. (B) Score of NMJs in response to light, showing improvement of function over 11 days (n = 47, one-way ANOVA p = 1 x 10-8). Error bars = SEM. (C) Contractility trace after 20 min treatment with 5 mg/mL of the neurotoxin α-bungarotoxin (BTX). (D) Quantification of NMJ function (score) before and after treatment with BTX (n = 6; one-way ANOVA F = 9 x 10-8). (E) Contractility trace of tissues with 20% MG sera after 48 h of treatment. (F) NMJ function assessment for treated and control groups before and after treatment and post-recovery (n = 12; after treatment post-hoc ANOVA F = 0.0002; *indicates p = 0.0015 **indicates p = 3.5 x10−6) (MG = Myasthenia Gravis; NHS=normal human serum). n indicates the number of biological replicates. This figure has been modified with permission from Vila et al.18,19. Please click here to view a larger version of this figure.
Table 1. Myogenic and neural media formulations. Organized list of growth factors and supplements for media. Please click here to download this Table.
Table 2. List of MATLAB functions. Brief explanations of MATLAB functions used for image analysis. This table has been modified with permission from Vila et al.18. Please click here to download this Table.
Supplemental Files
Supplementary Figure 1. Myotube and motoneuron differentiations. (A) Myotube differentiation protocol (MM = maturation media). (B) Immunofluorescence (IF) images showing expression of skeletal muscle markers myosin heavy chain (MHC), α-actinin, and desmin in differentiated myotubes. (C) Motoneuron differentiation protocol. (D) Localization of channelrhodopsin-2 with YFP (ChR2-YFP) to membrane in optogenetic iPSCs and (E) MNs confirming successful infection. (F) Co-expression of choline acetyltransferase (ChAT, red) and ChR2 (green) in optogenetic MNs. Scale bars 100 μm. Reproduced with permission from Vila et al.18. Please click here to download this File.
Supplementary Figure 2. Incorrect processing that requires adjustment of parameters. (A) Incorrect baseline time results in peaks of the wrong shape. (B) A peak threshold that is too high results in peaks being undetected. (C) Too lenient conditions for the detection of minima (i.e., too low values chosen for minMinProminence and/or minMinWidth) result in minima being labeled in the middle or even top of the peaks, therefore being counted as "untriggered" because they start too far from the preceding light pulse. (D) Too stringent conditions for the detection of minima result in the start of the peak being labeled before the light pulse or even missing, as in this example. Reproduced with permission from Vila et al.18. File showing 3D model of bioreactor platform that can be edited and exported to make molds for bioreactor manufacturing using PDMS. Please click here to download this File.
https://github.com/ofvila/NMJ-function-analysis
Supplementary CAD File. File showing 3D model of bioreactor platform that can be edited and exported to make molds for bioreactor manufacturing using PDMS. Please click here to download this File.
Discussion
This system is an engineered 3D human tissue model that combines optogenetics and video processing to enable automated and unbiased evaluation of NMJ function. Using a standardized protocol, we have demonstrated the ability to measure changes in NMJ function during physiological development and characterize the damaging effects of pathologies such as neurotoxin exposure and myasthenia gravis patient sera.
Previous studies have reported the ability to model MG with optogenetic hPSC-derived motoneurons in co-culture with skeletal myotubes using MG patient sera22. However, the system was in 2D and recorded contractions in random locations, thereby limiting reproducibility and decreasing the quantitative ability of the model. One of the greatest advantages of our system, compared to a 2D system, is that it allows us to emulate the 3D environment, which more accurately facilitates cellular and morphological development between MNs and SkMs. There is great evidence that supports that the biomechanical cues required for a tissue to behave like its native physiology are superiorly mimicked in a 3D system13,14,15,16. Additionally, various aspects of the 3D system allow for a more controlled environment: directed formation of synapses, less heterogeneity between samples, increased spatial-temporal control of stimulation via optogenetics, and an automated high-throughput system that decreases the subjectivity of each sample's analysis.
The developed system can be used to model other neuromuscular diseases (NMDs), such as amyotrophic lateral sclerosis (ALS), Duchenne muscular dystrophy (DMD), and others. Although ALS pathophysiology involves the dysfunction of the motor neuron itself rather than the neuromuscular junction17,27, our system has the ability to recapitulate the disease state of ALS by showing diminished muscle contraction after neuron degeneration or damage. DMD is a muscular degenerative disease caused by a lack of dystrophin protein, which leads to muscle weakness and eventual degeneration28. The system could model DMD by incorporating genetically engineered skeletal muscle that carries a dystrophin inhibitor or diseased muscle cells that lack dystrophin. Overall, the system has the flexibility and power to recapitulate various NMDs in an in vitro 3D platform.
For a protocol that requires various sources of cells, each with particular levels of confluency, one of the most critical aspects is timing. During the expansion of primary skeletal muscle cells, it is important to not allow them to become too confluent (65%-70%) due to fusion. At times, even when using trypsin to break up the clumps, it has been observed that this translates to lower efficacious seeding into the bioreactors. Another critical aspect is the open-well bioreactor platform production. In order to increase homogeneity among the PDMS platforms, the amount of time they spend in the oven should be kept uniform across all platforms to avoid any fluctuations in the mechanical properties of the platform pillars. Additionally, during the seeding of the muscle cells into the reactors, in order to maintain consistency of seeding density, it is sensible to vortex the tube gently (1-1.5 s) between every two to three compartments within the bioreactors. Lastly, the success rate for NMJ development was seen to be dependent largely on the success of motoneuron seeding since the addition of neurospheres is so delicate and varies from person to person. With seeding practice, the success rate increases, allowing for about 90% of initiated cultures to be stimulated and imaged.
Currently, some limitations that could be addressed in the near future are the incorporation and validation of optogenetic proteins other than the ChR2 discussed. Specific control over both cell types and optogenetic sensors for voltage/calcium imaging or muscle metabolism could be added to the system for increased readouts and analysis. As blue light induces phototoxicity at high doses, the stimulation regimen was limited to 30 s. This effect could be ameliorated by incorporating a red-shifted channelrhodopsin for longer duration stimulation studies. Additionally, another limitation that could be improved in future iterations of the model is the lack of neuronal supporting cells like Schwann cells. The versatility of this platform and analysis code, however, allows for a variety of optogenetic constructs and cell types to be incorporated. While our ChR2-hiPSC line was created using CRISPR, other methods could be used to implement an optogenetic response into hiPSCs. Furthermore, with the adaptability of hiPSCs, the NMJ model can be developed from a single cell source, allowing for patient-specific models to be used to further study neuromuscular diseases18.
This platform provides repeated, automated, and unbiased analysis of human NMJ function over time and can detect quantifiable changes in function due to external effectors, like neurotoxins or MG patient serum. Overall, our 3D system allows for the investigation of human NMJ function early in disease pathology, provides the opportunity for drug screening, and lays the foundation for a robust system to advance precision medicine.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We gratefully acknowledge funding support by the NIH [grant numbers EB025765 and EB027062], DOD [award number W81XWH-18-1-0095], and the UCSF Health Innovation via Engineering (HIVE Fellowship). We gratefully acknowledge the Columbia University Stem Cell Core for their help and guidance with cell reprogramming.
Materials
| Cells | |||
| SkMDC | Cook Myosite | P01059-14M | |
| Media and Supplements | |||
| Advanced DMEM/F12 | ThermoFisher Scientific | 12634-020 | |
| Bovine Serum Albumin solution | Millipore Sigma | A9576-50ML | |
| G-5 Supplement (100X) | ThermoFisher Scientific | 17503-012 | |
| Geneticin Selective Antibiotic (G418 Sulfate) (50 mg/mL) | ThermoFisher Scientific | 10131-035 | |
| Insulin, Recombinant Human | Millipore Sigma | 91077C-100MG | |
| Matrigel | Corning | 354277 | |
| mTeSR Plus | Stem Cell Technologies | 100-0276 | |
| MyoTonic Growth Media Kit | Cook Myosite | MK-4444 | |
| N-2 Supplement | ThermoFisher Scientific | 17502-048 | |
| NBactiv4 500 mL | BrainBits LLC | Nb4-500 | |
| Neurobasal Medium | ThermoFisher Scientific | 21103-049 | |
| Neurobasal-A Medium | ThermoFisher Scientific | A13710-01 | |
| Pluronic F-127 | Sigma Aldrich | P2443 | |
| ReLeSR | Stem Cell Technologies | 5872 | |
| Plasticware | |||
| 30 mm cage cube system | ThorLabs | CM1-DCH, CP33, ER1-P4 and ER2-P4 | |
| 37 µm Reversible Strainer, large | Stem Cell Technologies | 27250 | |
| 546 nm short-pass excitation filter | Semrock | FF01-546/SP-25 | |
| 573 nm dichroic mirror | Semrock | FF573-Di01–25×36 | |
| 594 nm long- pass emission filter | Semrock | BLP01-594R-25 | |
| 594 nm long-pass excitation filter | Semrock | BLP01-594R-25 | |
| Blue (470nm) Rebel LED on a SinkPAD-II 10mm Square Base – 65 lm @ 700mA | LuxeonStarLEDs | SP-05-B4 | |
| Carclo 29.8° Frosted 10 mm Circular Beam Optic – Integrated Legs | LuxeonStarLEDs | 10413 | |
| Corning 60 mm Ultra-Low Attachment Culture Dish | Corning | 3261 | |
| Heat sink | LuxeonStarLEDs | LPD-19-10B | |
| Optics | |||
| pluriStrainer 400 µm, 25 pack, sterile | PluriSelect | 43-50400-03 | |
| pluriStrainer 500 µm, 25 pack, sterile | PluriSelect | 43-50500-03 | |
| Red (627nm) Rebel LED on a SinkPAD-II 10mm Square Base – 65 lm @ 700mA | LuxeonStarLEDs | SP-05-R5 | |
| ring-actuated iris diaphragm | ThorLabs | SM1D12D | |
| T-Cube LED drivers | ThorLabs | LEDD1B, KPS101 | |
| Molds | |||
| Female Threaded Hex Standoffs, 3 1/2" 10-32, Partially Threaded 1/2" | McMaster | 91920A046 | |
| Low-Profile C-Clamp | McMaster | 1705A12 | |
| Growth Factors | |||
| Adenosine 3′,5′-cyclic monophosphate | Millipore Sigma | A9501-1G | |
| CHIR 99021, 10 mg | Tocris | 4423/10 | |
| DAPT 10 mg | R&D Systems | 2634/10 | |
| Human CNTF, research grade, 5 µg | Miltenyl Biotec | 130-096-336 | |
| Human Vitronectin Protein, CF | R&D Systems | 2349-VN-100 | |
| Human Vitronectin Protein, CF | R&D Systems | 2349-VN-100 | |
| IGF1 Recombinant Human Protein | ThermoFisher Scientific | PHG0078 | |
| Laminin mouse protein, natural | ThermoFisher Scientific | 23017015 | |
| Recombinant Human Agrin Protein | R&D Systems | 6624-AG-050 | |
| Recombinant Human GDNF Protein, CF 50ug | R&D Systems | 212-GD-050/CF | |
| Recombinant Human Neurotrophin 3 100 ug | Cell Sciences | CRN500D | |
| Recombinant Human Neurotrophin-4 | Cell Sciences | CRN501B | |
| Recombinant Human Sonic Hedgehog/Shh (C24II) N-Terminus | R&D Systems | 1845-SH-100 | |
| Recombinant Human/Murine/Rat BDNF 50 ug | Peprotech | 450-02 | |
| Retinoic Acid, 50 mg | Millipore Sigma | R2625-50 | |
| SAG Smoothened Agonist | Millipore Sigma | 566660 | |
| SB431542 10 mg | Stem Cell Technologies | 72234 | |
| StemMACS LDN-193189 | Miltenyl Biotec | 130-103-925 | |
| Vitronectin from human plasma | Millipore Sigma | V8379-50UG | |
| Y-27632 dihydrochloride | Tocris | 1254 | |
| Antibodies | |||
| α-actinin mAb (Mouse IgG1) | Abcam | ab9465 | |
| Choline Acetyltransferase (ChAT) (Goat) | Millipore | AB144P | |
| Desmin mAb (Mouse IgG1) | Dako | M076029-2 | |
| Myosin Heavy Chain (MHC) (Mouse IgG2b) | DSHB | MF20 | |
| Equipment | |||
| Arduino Uno R3 | Arduino | A000066 | |
| Automated stage | Applied scientific instrumentation | MS- 2000 XYZ | |
| Expanded plasma cleaner | Harrick Plasma | PDC-001 (115V) | |
| Invitrogen Countess Automated Cell Counter | Marshal Scientific | I-CACC | |
| IX-81 Inverted fluorescence microscope | Olympus | IX-ILL100LH | |
| Series Stage Top Incubator System | Tokai Hit STX | TOKAI-HIT-STXG | |
| Zyla 4.2 sCOMS Camera | Andor Technology | ZYLA-4.2P-CL10 | |
| Software | |||
| Arduino Software (IDE) | Arduino | IDE 1.8.19 | |
| Mastercam | Mastercam | Mastercam for Solidworks | |
| Matlab | Matlab | R2021b | |
| NIS elements | Nikon | Basic Research | |
| Solidworks 3D CAD | Solidworks | Solidworks Standard |
Riferimenti
- Al-bassam, W., et al. Characteristics, incidence, and outcome of patients admitted to the intensive care unit with myasthenia gravis. Journal of Critical Care. 45, 90-94 (2018).
- Vila, O. F., Qu, Y., Vunjak-Novakovic, G. In vitro models of neuromuscular junctions and their potential for novel drug discovery and development. Expert Opinion on Drug Discovery. 15 (3), 307-317 (2020).
- Webster, R. G. Animal models of the neuromuscular junction, vitally informative for understanding function and the molecular mechanisms of congenital myasthenic syndromes. International Journal of Molecular Sciences. 19 (5), 1326 (2018).
- Jones, R. A., et al. Cellular and molecular anatomy of the human neuromuscular junction. Cell Reports. 21 (9), 2348-2356 (2017).
- Takahashi, K., et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 131 (5), 861-872 (2007).
- Rao, L., Qian, Y., Khodabukus, A., Ribar, T., Bursac, N. Engineering human pluripotent stem cells into a functional skeletal muscle tissue. Nature Communications. 9 (1), 126 (2018).
- Madden, L., Juhas, M., Kraus, W. E., Truskey, G. A., Bursac, N. Bioengineered human myobundles mimic clinical responses of skeletal muscle to drugs. eLife. 4, 04885 (2015).
- Maury, Y., et al. Combinatorial analysis of developmental cues efficiently converts human pluripotent stem cells into multiple neuronal subtypes. Nature Biotechnology. 33 (1), 89-96 (2015).
- Bianchi, F., et al. Rapid and efficient differentiation of functional motor neurons from human iPSC for neural injury modelling. Stem Cell Research. 32, 126-134 (2018).
- Turan, S., Farruggio, A. P., Srifa, W., Day, J. W., Calos, M. P. Precise correction of disease mutations in induced pluripotent stem cells derived from patients with limb girdle muscular dystrophy. Molecular Therapy. 24 (4), 685-696 (2016).
- Ebert, A. D., Liang, P., Wu, J. C. Induced pluripotent stem cells as a disease modeling and drug screening platform. Journal of Cardiovascular Pharmacology. 60 (4), 408-416 (2012).
- Lin, C. -. Y., et al. iPSC-derived functional human neuromuscular junctions model the pathophysiology of neuromuscular diseases. JCI Insight. 4 (18), (2021).
- Centeno, E. G. Z., Cimarosti, H., Bithell, A. 2D versus 3D human induced pluripotent stem cell-derived cultures for neurodegenerative disease modelling. Molecular Neurodegeneration. 13 (1), 27 (2018).
- Okano, T., Matsuda, T. Tissue engineered skeletal muscle: preparation of highly dense, highly oriented hybrid muscular tissues. Cell Transplantation. 7 (1), 71-82 (1998).
- Powell, C. A., Smiley, B. L., Mills, J., Vandenburgh, H. H. Mechanical stimulation improves tissue-engineered human skeletal muscle. American Journal of Physiology-Cell Physiology. 283 (5), 1557-1565 (2002).
- Ronaldson-Bouchard, K., et al. Advanced maturation of human cardiac tissue grown from pluripotent stem cells. Nature. 556 (7700), 239-243 (2018).
- Guo, X., et al. A human-based functional NMJ system for personalized ALS modeling and drug testing. Advanced Therapeutics. 3 (11), 2000133 (2020).
- Vila, O. F., et al. Bioengineered optogenetic model of human neuromuscular junction. Biomaterials. 276, 121033 (2021).
- Vila, O. F., et al. Quantification of human neuromuscular function through optogenetics. Theranostics. 9 (5), 1232-1246 (2019).
- Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G., Deisseroth, K. Millisecond-timescale, genetically targeted optical control of neural activity. Nature Neuroscience. 8 (9), 1263-1268 (2005).
- Nagel, G., et al. Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proceedings of the National Academy of Sciences. 100 (24), 13940-13945 (2003).
- Steinbeck, J. A., et al. Functional connectivity under optogenetic control allows modeling of human neuromuscular disease. Cell Stem Cell. 18 (1), 134-143 (2016).
- Santhanam, N., et al. Stem cell derived phenotypic human neuromuscular junction model for dose response evaluation of therapeutics. Biomaterials. 166, 64-78 (2018).
- Phillips Ii, L. H. The epidemiology of myasthenia gravis. Annals of the New York Academy of Sciences. 998 (1), 407-412 (2003).
- Sanders, D. B., et al. Does change in acetylcholine receptor antibody level correlate with clinical change in myasthenia gravis. Muscle & Nerve. 49 (4), 483-486 (2014).
- Vernino, S. Unraveling the enigma of seronegative myasthenia gravis. JAMA Neurology. 72 (6), 630-631 (2015).
- Osaki, T., Uzel, S. G. M., Kamm, R. D. Microphysiological 3D model of amyotrophic lateral sclerosis (ALS) from human iPS-derived muscle cells and optogenetic motor neurons. Science Advances. , (2018).
- Paredes-Redondo, A., et al. Optogenetic modeling of human neuromuscular circuits in Duchenne muscular dystrophy with CRISPR and pharmacological corrections. Science Advances. 7 (37), (2021).

