Chemo-enzymatic Synthesis of N-glycans for Array Development and HIV Antibody Profiling
Summary
A modular approach to the synthesis of N-glycans for attachment to an aluminum oxide-coated glass slide (ACG slide) as a glycan microarray has been developed and its use for the profiling of an HIV broadly neutralizing antibody has been demonstrated.
Abstract
We present a highly efficient way for the rapid preparation of a wide range of N-linked oligosaccharides (estimated to exceed 20,000 structures) that are commonly found on human glycoproteins. To achieve the desired structural diversity, the strategy began with the chemo-enzymatic synthesis of three kinds of oligosaccharyl fluoride modules, followed by their stepwise α-selective glycosylations at the 3-O and 6-O positions of the mannose residue of the common core trisaccharide having a crucial β-mannoside linkage. We further attached the N-glycans to the surface of an aluminum oxide-coated glass (ACG) slide to create a covalent mixed array for the analysis of hetero-ligand interaction with an HIV antibody. In particular, the binding behavior of a newly isolated HIV-1 broadly neutralizing antibody (bNAb), PG9, to the mixture of closely spaced Man5GlcNAc2 (Man5) and 2,6-di-sialylated bi-antennary complex type N-glycan (SCT) on an ACG array, opens a new avenue to guide the effective immunogen design for HIV vaccine development. In addition, our ACG array embodies a powerful tool to study other HIV antibodies for hetero-ligand binding behavior.
Introduction
N-glycans on glycoproteins are covalently linked to the asparagine (Asn) residue of the consensus Asn-Xxx-Ser/Thr sequon, which affect several biological processes such as protein conformation, antigenicity, solubility, and lectin recognition1,2. The chemical synthesis of N-linked oligosaccharides represents a significant synthetic challenge because of their huge structural micro heterogeneity and highly branched architecture. Careful selection of protecting groups to tune reactivity of building blocks, achieving selectivity at anomeric centers, and proper use of promoter/activator(s) are key elements in synthesis of complex oligosaccharides. To solve this problem of complexity, a great amount of work to advance N-glycan synthesis was reported recently3,4. In spite of these robust approaches, finding an effective method for the preparation of a wide range of N-glycans (~20,000) remains a major challenge.
The rapid mutation rate of HIV-1 to achieve the extensive genetic diversity and its ability to escape from neutralizing antibody response, is among the greatest challenges to develop a safe and prophylactic vaccine against HIV-15,6,7. One effective tactic that HIV uses to avoid the host immune response is the post-translational glycosylation of envelope glycoprotein gp120 with a diverse N-linked glycans derived from the host glycosylation machinery8,9. A recent report regarding the precise analysis of recombinant monomeric HIV-1 gp120 glycosylation from human embryonic kidney (HEK) 293T cells suggests the occurrence of structural microheterogeneity with a characteristic cell-specific pattern10,11,12. Therefore, understanding the glycan specificities of HIV-1 bNAbs requires well characterized gp120 related N-glycan structures in a quantity sufficient for analysis.
The discovery of glycan microarray technology provided high throughput-based exploration of specificities of a diverse range of carbohydrate-binding proteins, viruses/bacterial adhesins, toxins, antibodies, and lectines13,14. The systematic glycans arrangement in an arrayed chip-based format could determine problematic low affinity protein-glycan interactions through multivalent presentation15,16,17,18. This chip-based glycan arrangement conveniently appears to effectively mimic cell-cell interfaces. To enrich the technology and overcome the uneven issue associated with conventional array formats, our group recently developed a glycan array on an aluminum oxide-coated glass (ACG) slide using phosphonic acid-ended glycans to enhance the signal intensity, homogeneity, and sensitivity19,20.
To improve the current understanding about glycan epitopes of newly isolated HIV-1 broadly neutralizing antibodies (bNAbs), we have developed a highly efficient modular strategy for the preparation of a broad array of N-linked glycans21,22 to be printed on an ACG slide (see Figure 1). Specificity profiling studies of HIV-1 bNAbs on an ACG array offered the unusual detection of hetero-glycan binding behavior of highly potent bNAb PG9 that was isolated from HIV infected individuals23,24,25.
Protocol
1. Preparation of D1/D2 Arm Modules22
- Preparation of intermediate 2
- Weigh starting material 1 (shown in Figure 2, p-methoxyphenyl-O-2-acetamido-2-deoxy-β-D-glucopyranosyl-(1→2)-α-D-mannopyranoside (100 mg, 0.204 mmol)) into a 15 mL tube and dissolve in Tris buffer (25 mM, pH 7.5) containing manganese dichloride (MnCl2, 10 mM) to achieve a final glycan concentration of 5 mM.
- Add 2 equivalents of uridine diphosphate galactose (UDP–Gal).
- Add 150 units of enzyme β-1, 4 galactosyl transferases from bovine milk, and incubate the mixture at 37 oC for 15 h.
- After 15 h, perform thin layer chromatography (TLC) to indicate total consumption of starting material by spotting the reaction mixture on a TLC plate, developing with n-butanol/acetic acid/water (H2O) in a 1:1:1 ratio and staining by a solution of 0.25 M cerium ammonium molybdate followed by heating. Then quench the reaction by heating at 90 oC for 5 min.
- Centrifuge the reaction mixture at a speed of 2,737 x g for 3 min, and load the supernatant at the top of polyacrylamide gel column (see Table of Materials). Elute the column using distilled de-ionized water, and collect the product with fractions of 1 – 2 mL.
- Monitor the collected fractions by TLC26, developing with n-butanol: H2O: acetic acid in a 1:1:1 ratio, and staining by a solution of cerium ammonium molybdate followed by heating. Lyophilize27 the product containing fractions to obtain intermediate 2 (115 mg, 86%) as white powder. Characterize the product by NMR28 and mass spectroscopy29 (see Supplementary Data File).
- Preparation of intermediate 3
- Weigh starting material 2 (shown in Figure 2, p-methoxyphenyl-O-β-D-galactopyranosyl-(1→4)-2-acetamido-2-deoxy-β-D-glucopyranosyl-(1→2)-α-D-mannopyranoside (80 mg, 0.122 mmol)) in a 15 mL tube and dissolve it in Tris buffer (25 mM, pH 7.5) containing MnCl2 (10 mM) to prepare a final glycan concentration of 5 mM.
- Add 2 equivalents of guanosine 5'-diphospho-β-L-fucose disodium salt (GDP-Fuc).
- Add 150 units of enzyme α-1, 3 fucosyl transferases from Helicobacter pylori (Hp1-3FTΔ26695), and incubate the mixture at 37 oC for 15 h.
- Follow step 1.1.4.
- Follow step 1.1.5.
- Follow step 1.1.6 to obtain intermediate 3 (82 mg, 84%) as white powder. Characterize the product by NMR and mass spectroscopy (See Supplementary Data File).
- Preparation of Modules 4 and 5
- Weigh compound 2, p-methoxyphenyl-O-β-D-galactopyranosyl-(1→4)-2-acetamido- 2-deoxy-β-D-glucopyranosyl-(1→2)-α-D-mannopyranoside (0.230 g, 0.360 mmol) or compound 3, p-methoxyphenyl-O-β-D-galactopyranosyl-(1→4)-[α-L-fucopyranosyl-(1→3)-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-α-D-mannopyranoside (0.100 g, 0.125 mmol) into a 25 mL single-neck round-bottom flask and dry under vacuum for 30 min.
- Remove the flask from vacuum, fill it with nitrogen gas, add a magnetic stir bar, cap it with a rubber septum, and attach a nitrogen balloon.
- Inject 7 mL of dry pyridine and 3 mL of acetic anhydride (Ac2O) into the flask placed on an ice bath at 0 oC. Stir the resulting reaction for 12 h at room temperature using a magnetic stirrer at 600 – 700 rpm. Evaporate the solvents using a rotary evaporator at a vacuum pressure of 0 – 10 mbar at 50 oC.
- Dilute the crude mixture using 30 mL of dichloromethane (DCM) and extract with saturated aqueous sodium hydrogen carbonate (NaHCO3) (2 x 20 mL) into a 50 mL separatory funnel.
- Re-extract the aqueous layer with DCM (3 x 15 mL) and dry the combined organic layers over sodium sulfate. Remove the sodium sulfate by filtration and wash with DCM (5 mL). Evaporate the solvent using a rotary evaporator at 400 mbar vacuum pressure at 40 oC.
- Load the solution of crude mixture into 2 mL of DCM on top of the silica bed.
- Elute the column with a mixture containing toluene and acetone from 0 – 20% acetone in toluene and collect fractions of 5 – 10 mL.
- Monitor the collected fractions by TLC (steps 1.1.4 and 1.1.6), developing with toluene/acetone (7/3), and staining with a solution of cerium ammonium molybdate followed by heating on a hot plate.
- Evaporate the product containing fractions using a rotary evaporator to get desired products as white foam in 72% and 85% yields, respectively.
- Dissolve the products into 7 mL of acetonitrile: toluene: H2O in a 4:2:1 ratio in a 25 mL round-bottom flask.
- Add cerium ammonium nitrate (2 equivalents) while cooling to 0 oC using an ice bath. Stir the reaction at room temperature for 3 h at ~500 – 800 rpm.
- After TLC indicates total consumption of starting material (TLC developing with toluene/acetone (7/3) and visualizing by UV absorbance at 254 nm or by staining with a solution of cerium ammonium molybdate followed by heating), dilute the reaction mixture with ethyl acetate (30 mL) and extract with H2O (15 x 2 mL).
- Extract the combined organic layers with 5 mL of saturated sodium chloride (NaCl) solution.
- Evaporate the solvent using a rotary evaporator at 240 mbar vacuum pressure at 40 oC.
- Load the solution of crude product into 2 mL of DCM on top of the silica bed. Elute the column with a mixture containing toluene and acetone (0 – 20% acetone in toluene) and collect fractions of 5 – 10 mL. Monitor the collected fractions by TLC, developing with toluene/acetone (7/3).
- Evaporate the product-containing fractions using a rotary evaporator at 77 mbar vacuum and 40 oC to get desired products as white foams.
- Dissolve the respective alcohols into 10 mL of DCM in a 25-mL single-neck round-bottom flask and cool to -30 oC.
- Add diethylaminosulfur trifluoride (DAST, 2 equivalents). Stir the reaction mixture at -30 oC for 2 h.
- After TLC indicates consumption of starting material (TLC developing with toluene/acetone (4/1) and visualizing by UV absorbance at 254 nm or by staining with a solution of cerium ammonium molybdate followed by heating), dilute the reaction mixture with DCM (30 mL), and wash with saturated NaHCO3 (15 x 2 mL).
- Extract the combined organic layer with 5 mL of saturated NaCl solution, dry it by adding anhydrous magnesium sulfate, filter the mixture following a wash with DCM (5 mL), and collect it into a 100-mL single-neck round-bottom flask.
- Evaporate the solvent using a rotary evaporator at 400 mbar vacuum pressure at 40 oC.
- Load the solution of crude product in 2 mL of DCM on top of the silica bed. Elute the column with a mixture of toluene and acetone (0- 20% acetone in toluene) and collect fractions of 5 – 10 mL.
- Monitor the collected fractions by TLC, developing with toluene/acetone (7/3).
- Evaporate the product containing fractions using a rotary evaporator to get desired products 4, [2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[3,6-O-diacetyl-2-acetamido-2- deoxy-β-D-glucopyranosyl]-(1→2)-3,4,6-O-triacetyl-α-D-mannopyranosyl fluoride (54% over 2 steps) and 5, [2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[2,3,4-O-triacetyl-α-L- fucopyranosyl-(1→3)-3,6-O-diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-3,4,6-O-triacetyl-α-D-mannopyranosyl fluoride (67% over 2 steps) as white solids.
- Characterize the product by NMR and mass spectroscopy (See Supplementary Data File).
2. Preparation of Glycan 10
- Preparation of intermidiate 7
- Weigh silver triflate (AgOTf) (0.039 g, 0.155 mmol), bis(cyclopentadienyl)hafnium(IV) dichloride (Cp2HfCl2) (0.041 g, 0.108 mmol) into a 25-mL single-neck round-bottom flask, dry it under schlenk line vacuum for 30 min, remove it from the vacuum, and fill it with nitrogen gas.
- Transfer the freshly dried 4 Å molecular sieves (0.2 g) into the flask containing the AgOTf and Cp2HfCl2, add a magnetic stir bar, cap with the septum immediately, and add a nitrogen balloon.
- Transfer 3 mL dry toluene into the flask with a dry glass syringe. Stir the reaction mixture for 1 h at room temperature and then cool to 0 oC.
- Inject a solution of donor 4 (Figure 2, 0.043 g, 0.046 mmol) and acceptor 6, 5-Azidopentyl-O-2-O-acetyl-4,6-O-benzylidine-β-D-mannopyranosyl-(1→4)-O-(3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranosyl)-(1→4)-O-3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranoside (Figure 3, 0.045 g, 0.031 mmol) in 3 mL toluene into the flask through the septum using a 10 mL syringe at 0 oC. Stir the reaction mixture at room temperature for 3 h.
- After TLC indicates consumption of starting materials (TLC, developing with DCM/acetone (8.5/1.5) and visualizing by UV absorbance at 254 nm or by staining with a solution of cerium ammonium molybdate followed by heating), quench the reaction by injecting 10 equivalents of triethyl amine.
- Filter the molecular sieves through a celite bed into a 50-mL round-bottomed flask, and further wash with 10 mL of ethyl acetate. Extract the combined organic layers with a saturated solution of aqueous NaHCO3 (2 x 20 mL) in a 50 mL separatory funnel. Extract the aqueous layer with ethyl acetate (3 x 15 mL).
- Extract the combined organic layers with saturated NaCl solution (5 mL), dry it by using anhydrous magnesium sulfate, filter the mixture to remove the magnesium sulfate, wash with ethyl acetate (3 x 15 mL), and collect the filtrate in a 100-mL round-bottomed flask.
- Evaporate the solvent using a rotary evaporator at 240 mbar vacuum pressure at 40 oC.
- Load the solution of crude product in DCM on top of the silica bed column. Elute the column with a mixture containing DCM and acetone (0-10% acetone in DCM) and collect fractions of 5-10 mL.
- Monitor the collected fractions by TLC, developing with DCM/acetone (8.5/1.5). Evaporate the fractions containing product using a rotary evaporator to obtain intermidiate 7, 5-Azidopentyl-O-{[2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[3,6-O-diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-[3,4,6-O-triacetyl-α-D-mannopyranosyl]}-(1→3)-[2-O-acetyl-4,6-O-benzylidine-β-D-mannopyranosyl-(1→4)-O-(3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranosyl)-(1→4)-O-3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranoside (0.052 g, 70%), as white foam.
- Characterize the product by NMR and mass spectroscopy (See Supplementary Data File).
- Preparation of intermidiate 8
- Weigh hexasaccharide 7 (Figure 3, 0.040 g, 0.016 mmol) into a 25-mL single-neck round-bottom flask, dry it under vacuum for 1 h, remove it from vacuum, fill it with nitrogen gas, add a magnetic stir bar, and cap it with a rubber septum.
- Transfer 3 mL of acetonitrile:methanol (MeOH) (2:1 ratio) into the flask.
- Add p-toluene sulfonic acid monohydrate (0.001 g, 0.008 mmol) into the reaction flask and stir the reaction at 800 rpm for 5 h.
- After TLC indicates comsumption of starting material (TLC, developing with DCM/acetone (8.5/1.5), and visualizing by UV absorbance at 254 nm or by staining with a solution of cerium ammonium molybdate followed by heating), quench the reaction by injecting 10 equivalents of triethyl amine.
- Evaporate the solvent using a rotary evaporator at 337 mbar vacuum pressure at 40 oC.
- Dissolve the crude mixture with approximately 2 mL of DCM and load it on top of the silica bed.
- Elute the column with a mixture of DCM and acetone (0 – 10 % acetone in DCM) and collect fractions of 5 – 10 mL. Monitor the collected fractions by TLC, developing with DCM/acetone (8.5/1.5). Evaporate the solvent using a rotary evaporator at 400 mbar vacuum and 40 oC.
- Dry the residue under reduced pressure to give intermediate 8, 5-Azidopentyl-O-(2-O- acetyl-3,4,6-tri-O-benzyl-ced pressure to give →3)-2-O-acetyl-4, 6-O-benzylidine-β-D- mannopyranosyl-(1→4)-O-(3,6-di-O-benzyl-2-deoxy-2-phthalimido-β-D-glucopyranosyl)-(1→4)-O-3,6 -di-O-benzyl-2-deoxy-2-phthalimido-nzyl-2-deoxy-2-phth. (Figure 3, 0.022 g, 57%) as a white solid. Characterize the product by NMR and mass spectroscopy (See Supplementary Data File).
- Preparation of intermidiate 9
- Preparation of intermidiate 9 (Figure 3) was accomplished using donor 5 (0.015 g, 13.2 µmol) and acceptor 8 (Figure 3, 0.020 g, 8.80 µmol).
- AgOTf (0.011 g, 44.1 µmol) and Cp2HfCl2 (0.012 g, 30.8 µmol) were used as a promotors.
- Perform the steps 2.1.1 to 2.1.10 to get intermediate 9, 5-Azidopentyl-O-{[2,3,4,6-O- tetraacetyl-β-D-galactopyranosyl]-(1→4)-[3,6-O-diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-[3,4,6-O-triacetyl-α-D-mannopyranosyl]}-(1→3)-{2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[2,3,4-O-triacetyl-α-LS160fucopyranosyl-(1→3)-3,6-O-diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-3,4,6-O-triacetyl-α-D-mannopyranosyl}-(1→6)-[2-O-acetyl-β-D-mannopyranosyl-(1→4)-O-(3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranosyl)-(1O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranosyl)-D-glucopyra (0.010 g, 34%), as white foam.
- Global deprotection of intermidiate
- Weigh compound 9 (0.010 g, 2.9 µmol) into a 25-mL single-neck round-bottom flask, add a magnetic stir bar, cap with a rubber septum, and attach a nitrogen balloon.
- Transfer 2 mL of 1, 4 dioxane: H2O (4:1) into the flask. Add lithium hydroxide (LiOH) (0.005 g, 50% by wt.) into the flask. Stir the reaction mixture at 90 – 100 oC for 12 h and allow it to cool at RT.
- Evaporate the solvents using a rotary evaporator and dry the flask under vacuum for 1 h, fill it with nitrogen, remove it from the vacuum manifold, and cap it with a rubber septum.
- Inject 4 mL of dry pyridine and 2 mL of acetic anhydride into the flask through the septum using a 10 mL syringe at 0 oC. Stir the reaction mixture for 12 h at RT.
- Evaporate the solvents using a rotary evaporator at 0-10 mbar vacuum pressure at 50 oC.
- Dissolve the crude mixture using 30 mL of DCM and extract the solution with saturated aqueous NaHCO3 (2 x 20 mL) in a 50 mL separatory funnel.
- Re-extract the aqueous layer with DCM (3 x 15 mL). Evaporate the solvent using a rotary evaporator.
- Load a solution of product in approximately 2 mL of DCM on top of the C18 silica bed. Elute the column with a mixture of water and methanol (0 – 100% of methanol in water) and collect fractions of 5 – 10 mL.
- Collect the product fractions and evaporate under reduced pressure to get desired product as white foam.
- Dissolve the product into 5 mL of dry methanol in a 25 mL round bottom flask and cap it with a rubber septum.
- Transfer 0.1 mL of sodium methoxide (NaOMe) in methanol and cool to 0 oC using an ice bath and stir the mixture for 12 h.
- Remove the solvent using a rotary evaporator and dry the product under high vacuum.
- Dissolve the crude products in 5 mL of methanol: H2O: acetic acid (6:3:1).
- Add palladium hydroxide (Pd(OH)2) (50% by wt) and stir the reaction under hydrogen atmosphere using a hydrogen balloon for 15 h.
- Filter the reaction through a celite bed and wash with 2 mL methanol followed by 2 mL of dd water.
- Evaporate the solvents using a rotary evaporator.
- Dissolve the crude mixture with approximately 1 mL of water and load it on top of the polyacrylamide gel bed (see Table of Materials). Elute the product with water and collect fractions of 1 – 2 mL.
- Monitor the collected fractions by TLC, develop with n-butanol: H2O: acetic acid (1:1:1).
- Collect the product fractions and lyophilize to get the desired product 10, 5-Aminopentyl-β-D-galactopyranosyl-(1→4)-2-acetamido-2-deoxy-β-D-glucopyranosyl-(1→2)-α-D-mannopyranosyl]-(1→3),-[β-D-galactopyranosyl-(1→4)-(α-L-fucopyranosyl-(1→3)-2-2-acetamido-2-deoxy-β-D-glucopyranosyl)-(1→2)-α-D-mannopyranosyl]-(1→6)-β-D-mannopyranosyl-(1→4)-2-acetamido-2-deoxy-β-D-glucopyranosyl-(1→4)-2-acetamido-2-deoxy-β-D-glucopyranoside (0.002 g, 36%), as a white powder.
- Characterize the product by NMR and mass spectroscopy (See Supplementary Data File).
3. Preparation of Glycans with Phosphonic Acid Tail19,22
- Weigh the respective glycan (2 – 5 µmol) in a 5 mL round-bottom flask, add a magnetic stir bar, and cap it with a rubber septum.
- Transfer 400 µL of freshly dried dimethylformamide.
- Add [2 (2 (2 (bis (benzyloxy) phosphoryl) ethoxyethoxy) ethyl (2, 5-dioxopyrrolidin1-yl) carbonate] (10 – 25 µmol, prepared in house) to the glycan solution. Stir the mixture at 800 rpm for 5 h.
- Evaporate the solvents using a rotary evaporator at 5 – 10 mbar vacuum pressure at 40 oC.
- Dissolve the reaction mixture in 0.5 mL of water and load at the top of the polyacrylamide gel bed (see Table of Materials). Elute the product using water and collect fractions of 1-2 mL.
- Monitor the collected fractions by TLC26. Spot the reaction mixture on a TLC plate, and add stain solution of 0.25 M cerium ammonium molybdate; follow by heating using a hot plate. Collect the product containing fractions and lyophilize to get the desired product as a white powder.
- Dissolve the product in 2 mL of water and add Pd (OH)2 (50% by weight). Stir the reaction at 800 rpm under hydrogen atmosphere using a hydrogen balloon for 15 h.
- Filter the reaction through a flux-calcined bed and wash with 2 mL methanol followed by 2 mL of distilled de-ionized water. Evaporate the solvents using a rotary evaporator at 300 mbar vacuum pressure at 40 oC.
- Dissolve the crude mixture with approximately 1 mL of water and load it on top of the polyacrylamide gel bed (see Table of Materials).
- Elute the product with water and collect fractions of 1 – 2 mL. Monitor the collected fractions by TLC26. Lyophilize the fractions (freeze the product using liquid nitrogen then place on lyophilizer under a vacuum) to get the desired product as a white powder. Characterize the products by NMR and mass spectroscopy (See Supplementary Data File) using D2O as a solvent.
4. Glycan Array
- Preparation of aluminum coated glass slides (ACG slide)19,20,22
NOTE: Fabrication of the aluminum-coated glass slides was done at Thin Film Technology Division, Instrument Technology Research Center and National Applied Research Laboratories, Hsinchu Science Park, Taiwan.- Pack coated slides, vacuum-seal them to prevent the formation of NAO, and keep sealed until the electrochemical reaction for surface anodization.
- Surface anodization of aluminum coated glass slides19,20,22
- Set the temperature-controlled incubator at 4 °C. Prepare 0.3 M oxalic acid aqueous solution and keep it in an ice bath. Take the 500 mL beaker, add a magnetic stirrer bar.
- Transfer the 0.3 M oxalic acid to the 500 mL beaker, and then place a 10 cm long platinum rod as a cathode into the solution. Keep stirring (at 300 rpm) the oxalic acid throughout the anodization process.
- Turn on lab tracer software, click on the "set up" button then "function choice sweep voltage." Set the start and stop voltage to 25.8 V, number of points to 100, compliance to 1, and sweep delay to 1,200 ms and click "ok."
- Clamp the aluminum-coated glass side facing toward the cathode. Click "run test" button. Observe the current measurement (~8 – 10 mA).
- After surface anodization, wash the slide thoroughly with double distilled water, purge dry with nitrogen gas, and then store in a 30% relative humidity chamber till further use.
- Fabrication of ACG glycan microarray22
- Prepare 100 µL of all monovalent glycans (I-XI) in ethylene glycol at 10 mM concentration individually.
- Dilute the above glycan with printing buffer (80% ethylene glycol and 20% de-ionized water) to make 100 µM concentration.
- For hetero-ligand study, prepare 5 µL of individual I-XI glycans, and to each add 5 µL of the Man5GlcNAc2 (1:1 ratio).
- Print microarrays by robotic pin (see Table of Materials) by the deposition of 0.6 nL of the previously prepared glycans onto ACG slides31.
- Store the printed slides in a humidity controlled dry box before the binding assay.
- Mapping glycan epitopes of HIV-1 broadly neutralizing antibody PG922
- Prepare 1 mL BSA contained PBST buffer, 3% w/v.
- Prepare 70 µL of PG9 (50 µg/mL) in PBST buffer (BSA contained PBST buffer, 3% w/v).
- Prepare 120 µL of secondary fluorescent tag antibody Donkey Anti-Human IgG (Alexa Fluor 647 conjugated) 50 µg/mL in PBST buffer (BSA contained PBST buffer, 3% w/v) in the dark.
- Mix primary antibody (PG9) and secondary fluorescent tag antibody in 1:1 ratio (60 µL each). Incubate premixed antibodies for 30 min at 4 °C.
- Load the ACG slide into the slide incubation chamber which is divided into 16 wells. Transfer 100 µL of premixed antibodies to the glycan array and incubate at 4 °C for 16 h.
- Pipet out premixed antibodies. Remove the slide incubation chamber.
- Wash the slide first in PBST buffer (PBS and 0.05% Tween-20), followed by de-ionized water and spin dry at 2,000 x g.
- Open the microarray image analysis software (see Table of Materials). Insert a slide with the arrayed features facing down.
- Click on the "settings" menu, set the image resolution to 5 µm per pixel, and the wavelength at 635 nm with PMT450 and Power to 100%.
- Click on the "scan" button to start imaging the slide.
- Click the "file" icon to save the scan image in tif format.
- Perform image analysis by following the software user's guide32.
- Calculate the total intensity of fluorescence and illustrate using image processing software33 (see Table of Materials).
NOTE: Here, the average percentage error for all data points is presented by error bars.
Representative Results
A modular chemo-enzymatic strategy for the synthesis of a wide array of N-glycans is presented in Figure 1. The strategy is based on the fact that diversity can be created at beginning by chemo-enzymatic synthesis of the three important modules, followed by the α-specific mannosylation at the 3-O and/or 6-O position of the mannose residue of the common core trisaccharide of N-glycans. Considering the structural diversity of bi-, tri-, and tetra-antennary complex type N-glycan structures, we believed that a set of oligosaccharyl donors and the core trisaccharides acceptor with preinstalled alkyl handlecould be used as starting materials to generate the desired structural diversity (Figure 1). The applicability of glycosyl fluoride donors in building a complex glycan library has been proven previously21.
To demonstrate the effectiveness of our strategy, a bi-antennary isomeric structure (10, Figure 3) was selected for synthesis. The D1 arm antennae 4 and D2 arm antennae 5 of glycan 10 were generated on large scales by chemo-enzymatic methods. In particular, the disaccharide acceptor 1 was enzymatically galactosylated by using β-1, 4-galactosyltransferase and uridine 5'-diphosphogalactose (UDP-Gal) to form trisaccharide 2. Next, the intermediate 2 was fucosylated at GlcNAc 3-O position in presence of α-1, 3-fucosyltransferase from Helicobacter pylori (Hpα1,3FT) to afford the desired tetrasaccharide 3. For chemical ligation to core, building blocks 2 and 3 were first peracetylated, and the reducing end p-methoxy phenol group was removed. Finally, fluoride was installed in the presence of DAST to get the desired modules 4 and 5, respectively (Figure 2).
Having the desired modules in hand, we next proceed to the stereoselective 3-O glycosylation of 4 to the core trisaccharide 6 under catalysis of silver triflate and hafnocene dichloride to provide the respective hexasaccharide 7 (Figure 3). The benzylidene protection that masking 4, 6-OH was removed using catalytic p-toluene sulfonic acid (p-TSA). Taking advantage of its reactivity, primary 6-OH of 8 was reacted with fluoride module 5 under similar experimental conditions to achieve the required decasaccharide 9. At last, global deprotection was performed to get glycan 10, which was further characterized using NMR and mass spectroscopy (See the Supplementary Data File).
Glycans containing a pentyl amine tail at the reducing end were modified with phosphonic acid linkers and attached to ACG surface through phosphonate chemistry (Figure 4). At last, HIV-1 bNAb PG9 was screened for its glycan specificity using homo- and hetero-glycans arrays to demonstrate for the first time that PG9 interacted with adjacent heteroglycans in the V1/V2 loop of HIV-1 gp120 surface (Figure 5).
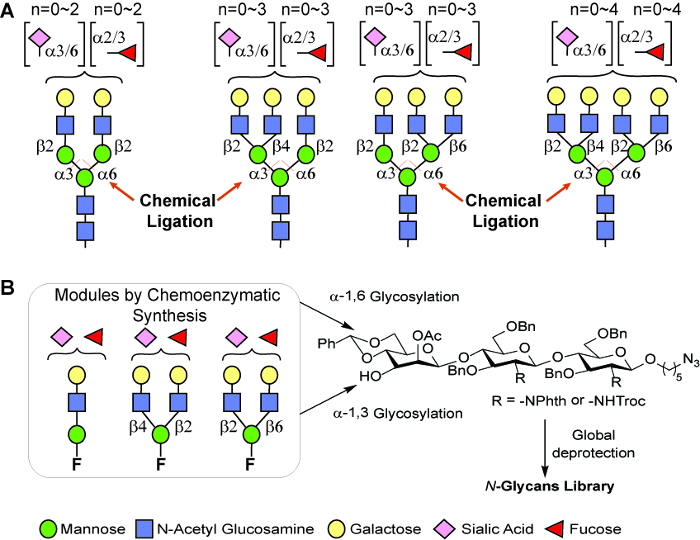
Figure 1: A general modular strategy for the preparation of N-glycans. (A) The number of N-glycans generated by this strategy that commonly occur on human glycoproteins is estimated to exceed 20,000. (B) Three types of modules prepared by this chemo-enzymatic approach that can be used for α-seletive glycosylations. Please click here to view a larger version of this figure.
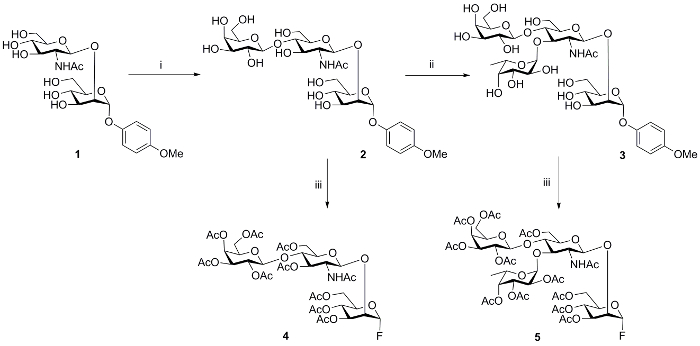
Figure 2: Chemoenzymatic synthesis of modules. i, UDP-galactose, β 1, 4-GalT, 15 h, 86%; ii, GDP-fucose, α 1, 3-FucT, 15 h, 84%; iii, (1) Ac2O, pyridine, RT, 12 h; (1) CAN, ACN: toluene: H2O, (3) DAST, CH2Cl2, -30 oC. CAN: Cerium ammonium nitrate; DAST: Diethylaminosulfur trifluoride.
Product nomenclature : 1, p-methoxyphenyl-O-2-acetamido-2-deoxy-β-D- glucopyranosyl-(1→ 2)-α-D-mannopyranoside; 2, p-methoxyphenyl-O-β-D-galactopyranosyl-(1→4)-2-acetamido-2- deoxy-β-D-glucopyranosyl-(1→2)-α-D-mannopyranoside; 3, p-methoxyphenyl-O-β-D- galactopyranosyl-(1→4)-[α-L-fucopyranosyl-(1→3)-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-α-D-mannopyranoside; 4, [2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[3,6-O- diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-3,4,6-O-triacetyl-α-D-mannopyranosyl fluoride; 5, [2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[2,3,4-O-triacetyl-α-L- fucopyranosyl(1→3)-3,6-O-diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-3,4,6-O-triacetyl-α-D-mannopyranosyl fluoride Please click here to view a larger version of this figure.
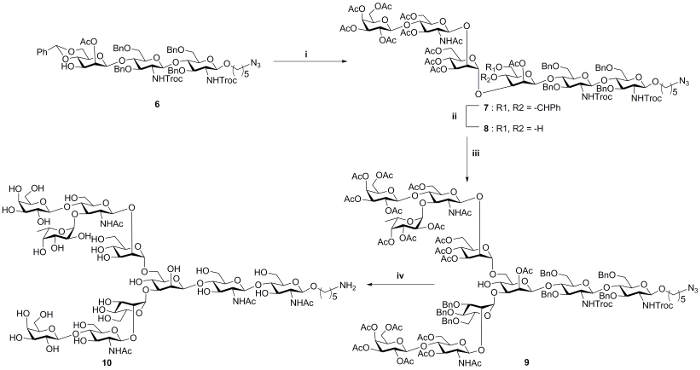
Figure 3: Chemical glycosylation of D1/D2 arm modules to core acceptor. i, 4, AgOTf, Cp2HfCl2, toluene, 4 Å MS, 0 oC to RT, 70%; ii, p-TSA, acetonitrile, RT, 57%; iii, 5, AgOTf, Cp2HfCl2, toluene, 4 Å MS, 0 oC to RT, 34%; iv, (1) LiOH, 1,4-dioxane: H2O; 90 oC, 12 h; (2) Ac2O, pyridine, 12 h; (3) NaOMe, MeOH, 12 h; (4) Pd(OH)2, MeOH : H2O : HCOOH (5:3:2), H2, 36%. AgOTf: Silver trifluromethanesulfonate; Cp2HfCl2: Bis(cyclopentadienyl)hafnium dichloride, MS: molecular sieves, Product nomenclature : 7, 5-Azidopentyl-O-{[2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[3,6-O-diacetyl-2- acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-[3,4,6-O-triacetyl-α-D-mannopyranosyl]}-(1→3)-[2-O-acetyl-4,6-O-benzylidine-β-D-mannopyranosyl-(1→4)-O-(3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranosyl)-(1→4)-O-3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranosid. 8, 5-Azidopentyl-O-(2-O- acetyl-3,4,6-tri-O -benzyl-α-D-mannopyranosyl-(1→3)-2-O-acetyl-4,6-O-benzylidine-β-D-mannopyranosyl-(1→4)-O-(3,6-di-O-benzyl-2-deoxy-2-phthalimido-β-D-glucopyranosyl)-(1→4)-O-3,6-di-O-benzyl-2-deoxy-2-phthalimido-β-D-glucopyranoside. 9, 5-Azidopentyl-O-{[2,3,4,6-O-tetraacetyl-β-D- galactopyranosyl]-(1→4)-[3,6-O-diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-[3,4,6-O-triacetyl-α-D-mannopyranosyl]}-(1→3)-{2,3,4,6-O-tetraacetyl-β-D-galactopyranosyl]-(1→4)-[2,3,4-O-triacetyl-α-LS160fucopyranosyl-(1→3)-3,6-O-diacetyl-2-acetamido-2-deoxy-β-D-glucopyranosyl]-(1→2)-3,4,6-O-triacetyl-α-D-mannopyranosyl}-(1→6)-[2-O-acetyl-β-D-mannopyranosyl-(1→4)-O-(3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-D-glucopyranosyl)-(1→4)-O-3,6-di-O-benzyl-2-deoxy-2-(2,2,2-trichloroethoxy)carbonylamino-β-Dglucopyranoside 10, 5-Aminopentyl-β-D-galactopyranosyl-(1→4)-2-acetamido-2-deoxy-β-D-glucopyranosyl- (1→2)-α-D-mannopyranosyl]-(1→3),-[β-D-galactopyranosyl-(1→4)(α-L-fucopyranosyl-(1→3)-2-2-acetamido-2-deoxy-β-D-glucopyranosyl)-(1→2)-α-D-mannopyranosyl]-(1→6)-β-D-mannopyranosyl-(1→4)-2-acetamido-2-deoxy-β-D-glucopyranosyl-(1→4)-2-acetamido-2-deoxy-β-D-glucopyranoside Please click here to view a larger version of this figure.
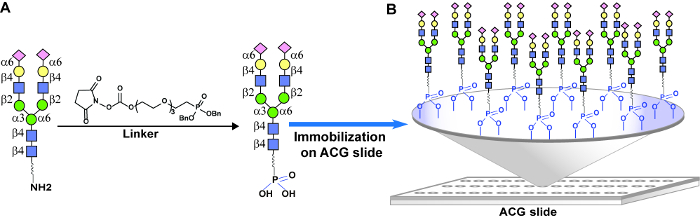
Figure 4: Glycan immobilization on an ACG array. (A) Chemical modification of glycan with amino tail into a phosphonic acid tail for covalent attachment to the ACG slide through phosphonate chemistry. (B) Distribution of glycans on an ACG surface. Please click here to view a larger version of this figure.
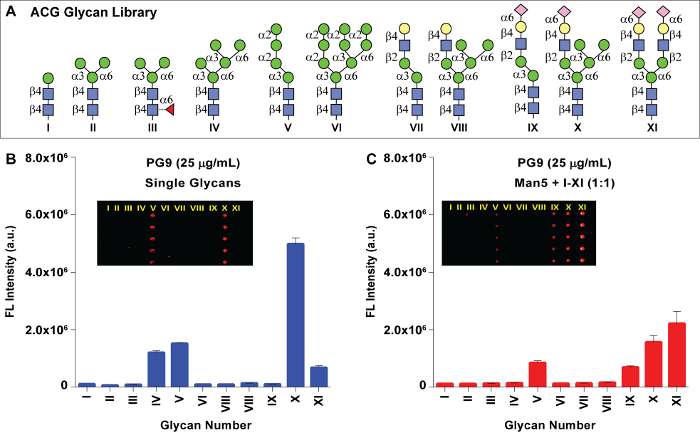
Figure 5: Glyan array analysis. (A) Structures of synthetic N-glycans that are printed on an ACG array. (B) Binding analysis of PG9 to individual glycans I-XI printed on ACG array (left panel) and to glycan mixtures of Man5 mixed with glycans I-XI (right panel) with 100 µM concentration. The molar concentrations in µM for PG9 are given in the legend. The mean signal intensities and standard error calculated for five independent replicates on the array are shown. Insets show fluorescence images. Please click here to view a larger version of this figure.
Discussion
A class of HIV-1 bNAbs including PG9, PG16, and PGTs 128, 141 – 145 were reported to be highly potent in neutralizing 70 – 80% of circulating HIV-1 isolates. The epitopes of these bNAbs are highly conserved among the variants of the entire HIV-1 group M, therefore they may guide the effective immunogen design for an HIV vaccine to elicit neutralizing antibodies23,24,25. As a part of our ongoing efforts to identify the glycan epitopes of HIV-1 broadly neutralizing antibodies, we reported the chemical and enzymatic synthesis of a panel of high mannose, hybrid, and complex type oligosaccharides to form a glycan array21,22. However, the commonly used glycan arrays on N-hydroxysuccinimide-coated (NHS) glass slides are unable to detect low affinity carbohydrate-protein interactions, probably due to heterogeneous glycan distribution resulting from hydrolysis of reactive N-hydroxysuccimide functional groups on its surface. To overcome the limitations of conventional array formats, we have developed a glycan array on an aluminum oxide coated-glass (ACG) slide. We further performed a homogeneity comparison between the ACG array and the commonly used NHS array to prove that the ACG array offered a more homogeneous glycan presentation on its surface22. Our preliminary binding analysis was not able to observe binding of PG9 to any of the glycans on NHS array (data not shown). Therefore, to define the binding specificity of PG9, the glycans I-XI were attached to a phosphonic acid tail and printed on the ACG slide with 100 µM of individual glycans (Figure 4). The glycan immobilization on an ACG slide through phosphonate chemistry offered a more stable and homogenous distribution. Each of the glycans was printed with five replicates and slide images were obtained from a fluorescence scan after DyLight649-conjugated donkey anti-Human IgG antibody incubation. The binding analysis of PG9 towards individual glycans printed on ACG array (Figure 5, left panel) suggests that PG9 interacted strongly with hybrid-type glycan (X). The interactions were also detected for high mannose type Man5 (glycan IV) and the complex-type glycan (XI).
The molecular level interactions between PG9 and hybrid-type glycan (X) are difficult to determine due to the lack of their co-crystal structure information. The hybrid-type glycan X is composed of a complex-type arm at the 3-O position of the core and a trimannose arm linked to the 6-O position of the central trisaccharide. Binding of PG9 to hybrid glycan X suggests that PG9 requires both mannose and sialic acid residues in close vicinity for high affinity interactions. To identify the glycan combination that best fits into the PG9 binding pocket, we performed a mixed-glycan array study. Glycan IV was mixed with every glycan from I-XI in a 1:1 mole ratio and spotted on an ACG array. The binding analysis of PG9 to each of the mixtures indicated that a combination of Man5 and a SCT (IV+XI) resulted in the highest affinity towards PG9 compared to IV or XI alone. In addition, mixtures containing Man5 and those containing sialylated antennae such as glycans VIII and X were also detected22. For the first time, these results offered an array based proof for hetero-glycans' binding behavior of PG9 that was suggested by crystal structure studies of PG9 with the HIV-1 gp120 V1/V2 domain23.
In conclusion, we have demonstrated an efficient modular chemo-enzymatic strategy for the preparation of highly diverse N-linked oligosaccharides that occur on human glycoproteins. In addition, the development of an ACG array provides an effective means to determine extremely weak protein-carbohydrate interactions and, more importantly, the interactions that occur through hetero-glycans.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
The authors thank the Thin Film Technology Division, Instrument Technology Research Center (ITRC) and National Applied Research Laboratories, Hsinchu Science Park, Taiwan. This work was supported by the National Science Council (grant no. MOST 105-0210-01-13-01) and Academia Sinica.
Materials
| Acetic acid | Sigma Aldrich | 64197 | |
| Acetonitrile | Sigma Aldrich | 75058 | |
| Acetic anhydride | Sigma Aldrich | 108247 | |
| Anhydrous magnesium sulfate | Sigma Aldrich | 7487889 | |
| Boron trifluoride ethyl etherate | Sigma Aldrich | 109637 | |
| Bovine serum albumin | Sigma Aldrich | 9048468 | |
| Bio-Gel P2 polyacrylamide | Bio-Rad | 1504118 | |
| Bis(cyclopentadienyl)hafnium(IV) dichloride | Sigma Aldrich | 12116664 | |
| β-1, 4 Galactosyl transferases from bovine milk | Sigma Aldrich | 48279 | |
| BioDot Cartesion technology with robotic pin SMP3 (Stealth Micro Spotting Pins) | Arrayit | ||
| Cerium ammonium molybdate | TCI | C1794 | |
| Cerium ammonium nitrate | Sigma Aldrich | 16774213 | |
| Clean glass slide | Schott | ||
| Cytidine-5′-monophospho-N-acetylneuraminic acid | Sigma Aldrich | 3063716 | |
| Deuterated chloroform | Sigma Aldrich | 865496 | |
| Donkey Anti-Human IgG (Alexa Fluor647 conjugated | Jackson Immuno Research, USA | 709605098 | |
| Dichloromethane | Sigma Aldrich | 75092 | |
| Diethylaminosulfur trifluoride | Sigma Aldrich | 38078090 | |
| Dimethylformamide | Sigma Aldrich | 68122 | |
| Ethyl acetate | Sigma Aldrich | 141786 | |
| Ethylene glycol | Acros Organic | 107211 | |
| FAST frame slide incubation chambers | Sigma Aldrich | ||
| Guanosine 5'-diphospho-b-L-fucose disodium salt | Sigma Aldrich | 15839700 | |
| Lab tracer 2.0 software | Section 4 of the Protocol | ||
| GenePix Pro 4300A reader (microarray image analysis) | moleculardevices | www.moleculardevices.com | |
| GraphPad Prism Software (Image processing ) | GraphPad Software, Inc | http://www.graphpad.com/guides/prism/6/user-guide/ | |
| Lithium hydroxide | Sigma Aldrich | 1310652 | |
| Manganese chloride | Sigma Aldrich | 7773015 | |
| Methanol | Sigma Aldrich | 67561 | |
| N-butanol | Sigma Aldrich | 71363 | |
| Oxalic acid | Acros Organic | 144627 | |
| Palladium hydroxide | Sigma Aldrich | 12135227 | |
| Phosphate Buffered Saline | Thermo Fisher Scientific | 10010023 | |
| Pyridine | Sigma Aldrich | 110861 | |
| P-Toluene sulfonic acid monohydrate | Sigma Aldrich | 773476 | |
| Silver triflate | Sigma Aldrich | 2923286 | |
| Sodium bicarbonate | Sigma Aldrich | 144558 | |
| Sodium chloride | Sigma Aldrich | 7647145 | |
| Sodium hydrogen carbonate | Sigma Aldrich | 144558 | |
| Sodium methoxide | Sigma Aldrich | 124414 | |
| Sodium sulfate | Sigma Aldrich | 7757826 | |
| Toluene | Sigma Aldrich | 108883 | |
| Tris buffer | Amresco | N/A | Ultra-pure grade |
| Tween-20 | Amresco | 9005645 | |
| Uridine diphosphate galactose (UDP-galactose) | Sigma Aldrich | 137868521 |
Riferimenti
- Kim, P. J., Lee, D. Y., Jeong, H. Centralized modularity of N-linked glycosylation pathways in mammalian cells. PloS one. 4, e7317 (2009).
- Townsley, S., Li, Y., Kozyrev, Y., Cleveland, B., Hu, S. L. Conserved Role of an N-Linked Glycan on the Surface Antigen of Human Immunodeficiency Virus Type 1 Modulating Virus Sensitivity to Broadly Neutralizing Antibodies Against the Receptor and Coreceptor Binding Sites. J.virol. 90, 829-841 (2015).
- Wang, Z., et al. A General Strategy for the Chemoenzymatic Synthesis of Asymmetrically Branched N-Glycans. Science. 341, 379-383 (2013).
- Li, L., et al. Efficient Chemoenzymatic Synthesis of an N-glycan Isomer Library. Chem Sci. 6, 5652-5661 (2015).
- Pritchard, L. K., Harvey, D. J., Bonomelli, C., Crispin, M., Doores, K. J. Cell- and Protein-Directed Glycosylation of Native Cleaved HIV-1 Envelope. J.Virol. 89, 8932-8944 (2015).
- Behrens, A. J., et al. Composition and Antigenic Effects of Individual Glycan Sites of a Trimeric HIV-1 Envelope Glycoprotein. Cell Rep. 14, 2695-2706 (2016).
- Barouch, D. H. Challenges in the development of an HIV-1 vaccine. Nature. 455, 613-619 (2008).
- Horiya, S., MacPherson, I. S., Krauss, I. J. Recent Strategies Targeting HIV Glycans in Vaccine Design. Nat Chem Bio. 10, 990-999 (2014).
- Wang, L. X. Synthetic carbohydrate Antigens for HIV Vaccine Design. Curr Opin Chem Biol. 17, 997-1005 (2013).
- Tian, J., et al. Effect of Glycosylation on an Immunodominant Region in the V1V2 Variable Domain of the HIV-1 Envelope gp120 Protein. PLoS Comput Biol. 12, e1005094 (2016).
- Geyer, H., Holschbach, C., Hunsmann, G., Schneider, J. Carbohydrates of human Immunodeficiency Virus. Structures of Oligosaccharides Linked to the Envelope Glycoprotein 120. The J Bio Chem. 263, 11760-11767 (1988).
- Lee, J. H., Ozorowski, G., Ward, A. B. Cryo-EM Structure of A Native, Fully Glycosylated, Cleaved HIV-1 Envelope Trimer. Science. 351, 1043-1048 (2016).
- Lonardi, E., Balog, C. I., Deelder, A. M., Wuhrer, M. Natural GlycanMicroarrays. Expert Rev Proteomics. 7, 761-774 (2010).
- Paulson, J. C., Blixt, O., Collins, B. E. Sweet Spots in Functional Glycomics. Nat. Chem. Bio. 2, 238-248 (2006).
- Dotsey, E. Y., et al. A High Throughput Protein Microarray Approach to Classify HIV Monoclonal Antibodies and Variant Antigens. PLoS One. 10, e0125581 (2015).
- Wu, C. Y., Liang, P. H., Wong, C. H. New Development of Glycan Arrays. Org Biomol Chem. 7, 2247-2254 (2009).
- Scurr, D. J., et al. Surface Characterization of Carbohydrate Microarrays. Langmuir. 26, 17143-17155 (2010).
- Blixt, O., et al. Printed Covalent Glycan Array for Ligand Profiling of Diverse Glycan Binding Proteins. Proc Natl Acad Sci U S A. 101, 17033-17038 (2004).
- Chang, S. H., et al. Glycan Array on Aluminum Oxide-Coated Glass Slides Through Phosphonate Chemistry. J. Am. Chem. Soc. 132, 13371-13380 (2010).
- Tseng, S. Y., et al. Preparation of Aluminum Oxide-Coated Glass Slides for Glycan Microarrays. ACS Omega. 1, 773-783 (2016).
- Shivatare, S. S., et al. Efficient Convergent Synthesis of Bi-, Tri-, and Tetra-Antennary Complex Type N-Glycans and Their HIV-1 Antigenicity. J. Am. Chem. Soc. 135, 15382-15391 (2013).
- Shivatare, S. S., et al. Modular synthesis of N-Glycans and Arrays for the Hetero-Ligand Binding Analysis of HIV Antibodies. Nat Chem. 8, 338-346 (2016).
- McLellan, J. S., et al. Structure of Hiv-1 gp120 V1/V2 Domain with Broadly Neutralizing Antibody PG9. Nature. 480, 336-343 (2011).
- Julien, J. P., et al. Asymmetric Recognition of the HIV-1 Trimer by Broadly Neutralizing Antibody PG9. Proc Natl Acad Sci U S A. 110, 4351-4356 (2013).
- Willis, J. R., et al. Long Antibody HCDR3s from HIV-Native Donors Presented on a PG9 Neutralizing Antibody Background Mediate HIV Neutralization. Proc Natl Acad Sci U S A. 113, 4446-4451 (2016).
- JoVE Science Education Database. Essentials of Organic Chemistry. Performing 1D Thin Layer Chromatography. JoVE. , (2017).
- de Castro, M. D., et al. A useful approach to the automation of analytical processes?. J Automat Chem. 12, 267-279 (1990).
- JoVE Science Education Database. Essentials of Organic Chemistry. Nuclear Magnetic Resonance (NMR) Spectroscopy. JoVE. , (2017).
- JoVE Science Education Database. Essentials of Analytical Chemistry. Introduction to Mass Spectrometry. JoVE. , (2017).
- Tseng, S. Y., et al. Preparation of Aluminum Oxide-Coated Glass Slides for Glycan Microarrays. ACS Omega. 1 (5), 773-783 (2016).
- Blixt, O., et al. Printed covalent glycan array for ligand profiling of diverse glycan binding proteins. Proc Natl Acad Sci U S A. 101, 17033-17038 (2004).
- . . GenePix Pro 6.0 Microarray Acquisition and Analysis Software for GenePix Microarray Scanners User’s Guide & Tutorial. , (2017).

