Establishing 3-Dimensional Spheroids from Patient-Derived Tumor Samples and Evaluating their Sensitivity to Drugs
Summary
The present protocol describes generating 3D tumor culture models from primary cancer cells and evaluating their sensitivity to drugs using cell-viability assays and microscopic examinations.
Abstract
Despite remarkable advances in understanding tumor biology, the vast majority of oncology drug candidates entering clinical trials fail, often due to a lack of clinical efficacy. This high failure rate illuminates the inability of the current preclinical models to predict clinical efficacy, mainly due to their inadequacy in reflecting tumor heterogeneity and the tumor microenvironment. These limitations can be addressed with 3-dimensional (3D) culture models (spheroids) established from human tumor samples derived from individual patients. These 3D cultures represent real-world biology better than established cell lines that do not reflect tumor heterogeneity. Furthermore, 3D cultures are better than 2-dimensional (2D) culture models (monolayer structures) since they replicate elements of the tumor environment, such as hypoxia, necrosis, and cell adhesion, and preserve the natural cell shape and growth. In the present study, a method was developed for preparing primary cultures of cancer cells from individual patients that are 3D and grow in multicellular spheroids. The cells can be derived directly from patient tumors or patient-derived xenografts. The method is widely applicable to solid tumors (e.g., colon, breast, and lung) and is also cost-effective, as it can be performed in its entirety in a typical cancer research/cell biology lab without relying on specialized equipment. Herein, a protocol is presented for generating 3D tumor culture models (multicellular spheroids) from primary cancer cells and evaluating their sensitivity to drugs using two complementary approaches: a cell-viability assay (MTT) and microscopic examinations. These multicellular spheroids can be used to assess potential drug candidates, identify potential biomarkers or therapeutic targets, and investigate the mechanisms of response and resistance.
Introduction
In vitro and in vivo studies represent complementary approaches for developing cancer treatments. In vitro models allow for the control of most experimental variables and facilitate quantitative analyses. They often serve as low-cost screening platforms and can also be used for mechanistic studies1. However, their biological relevance is inherently limited, as such models only partially reflect the tumor microenvironment1. In contrast, in vivo models, such as patient-derived xenografts (PDX), capture the complexity of the tumor microenvironment and are more suitable for translational studies and individualizing treatment in patients (i.e., investigating the response to drugs in a model derived from an individual patient)1. However, in vivo models are not conducive to high-throughput approaches for drug screening, as the experimental parameters cannot be controlled as tightly as in in vitro models and because their development is time-consuming, labor intensive, and costly1,2.
In vitro models have been available for over 100 years, and cell lines have been available for over 70 years3. During the last several decades, however, the complexity of the available in vitro models of solid tumors has increased dramatically. This complexity ranges from 2-dimensional (2D) culture models (monolayer structures) that are either tumor-derived established cell lines or primary cell lines to the more recent approaches involving 3-dimensional (3D) models1. Within the 2D models, a key distinction is between the established and primary cell lines4. Established cell lines are immortalized; therefore, the same cell line can be used globally over many years, which from a historical perspective, facilitates collaboration, the accumulation of data, and the development of many treatment strategies. However, genetic aberrations in these cell lines accumulate with every passage, thus compromising their biological relevance. Furthermore, the limited number of available cell lines does not reflect the heterogeneity of tumors in patients4,5. Primary cancer cell lines are derived directly from resected tumor samples obtained via biopsies, pleural effusions, or resections. Therefore, primary cancer cell lines are more biologically relevant as they preserve elements of the tumor microenvironment and tumor characteristics, such as intercellular behaviors (e.g., cross-talk between healthy and cancerous cells) and the stem-like phenotypes of cancer cells. However, the replicative capacity of primary cell lines is limited, which leads to a narrow culture time and limits the number of tumor cells that can be used for analyses4,5.
Models using 3D cultures are more biologically relevant than 2D culture models since the in vivo conditions are retained. Thus, 3D culture models preserve the natural cell shape and growth and replicate elements of the tumor environment, such as hypoxia, necrosis, and cell adhesion. The most commonly used 3D models in cancer research include multicellular spheroids, scaffold-based structures, and matrix-embedded cultures4,6,7.
The present protocol generates 3D tumor culture models (multicellular spheroids) from primary cancer cells and evaluates their sensitivity to drugs using two complementary approaches: a cell-viability assay (MTT) and microscopic examinations. The representative results presented herein are from breast and colon cancer; however, this protocol is widely applicable to other solid tumor types (e.g., cholangiocarcinoma, gastric, lung, and pancreatic cancer) and is also cost-effective, as it can be performed in its entirety in a typical cancer research/cell biology lab without relying on specialized equipment. The multicellular spheroids generated using this approach can be used to assess potential drug candidates, identify potential biomarkers or therapeutic targets, and investigate the mechanisms of response and resistance.
This protocol is divided into three sections: (1) the generation, collection, and counting of the spheroids in preparation for their use as a model for testing drug efficacy; (2) MTT assay to assess drug efficacy on the spheroids; and (3) the microscopic evaluation of morphological changes following the treatment of the spheroids with drugs as another approach for evaluating drug efficacy (Figure 1).
Protocol
The collection of human tumor samples used for the primary tumor cell cultures was performed as per institutional review board (IRB)-approved protocols at the Rabin Medical Center with written informed consent from the patients. Patients eligible for participation in the study included male and female adult and pediatric cancer patients with non-metastatic breast, colon, liver, lung, neuroendocrine, ovary, or pancreatic cancer, any pediatric cancer, or any metastatic cancer. The only exclusion criterion was the lack of capacity to provide informed consent.
1. Generation and collection of spheroids
NOTE: The isolation of primary tumor cells can be performed as described by Kodak et al.8. Importantly, primary tumor cells used for generating the spheroids can be derived directly from patient samples obtained by biopsy, resection, etc., or indirectly using tumor samples from patient-derived xenograft (PDX) models, as described by Moskovits et al.9.
- Prepare a single-cell suspension of adherent primary tumor cell cultures of 75%-100% confluence by taking a small flask (T25) with a single-cell adherent primary cell culture, removing the cell culture media, washing with PBS, and then adding 1 mL of 1x Accutase (a cell detachment solution) (see the Table of Materials) for 3 min at 37 °C.
- Neutralize the Accutase solution by adding 5 mL of cell culture medium (RPMI-1640 culture medium supplemented with 10% FBS, 1:100 penicillin-streptomycin, 1% non-essential amino acids, and 1% L-glutamine; see the Table of Materials).
- Aspirate the cells with a 10 mL serological pipette, and deposit them in a 15 mL conical tube. Centrifuge the tube at 800 x g for 5 min at room temperature.
- Remove the cell culture medium, add 5 mL of fresh cell culture medium on top of the cell pellet, and mix gently.
- Count the viable cells with a hemocytometer10. To do this, take an aliquot of 50 μL of the cell suspension, and mix it with 50 μL of trypan blue. Count the live cells (the cells that are negative for blue color), and calculate the total number of live cells in the suspension.
- Prepare a "3D culture medium" (a cell culture medium supplemented with 5% basement membrane matrix; see the Table of Materials).
NOTE: The "3D culture medium" is liquid-like at room temperature. At 37 °C, the consistency becomes more gel-like (so that the cells stay together), although it can still be pipetted (since it contains only 5% basement membrane matrix). - Calculate the number of cells needed for the assay and the total volume required; each well must contain 2,000-8,000 cells in 200 μL of medium.
- Prepare a cell suspension with the desired number of cells (e.g., 4,000 cells) in 200 μL of the "3D culture medium" and mix gently with the pipette to ensure a homogenous distribution.
- Transfer the suspension to a pipetting reservoir. Using a multi-channel pipette, add 200 μL of the cell suspension to each well of an ultra-low attachment 96-well plate (see the Table of Materials). Before each collection of the cells, mix the suspension well.
- Centrifuge the plate at 300 x g for 10 min at room temperature to enforce the clustering of the cells, thereby improving the cell aggregation, and incubate the plate at 37 °C in a 5% CO2 humidified incubator.
- Every 2-3 days, refresh the "3D culture medium". Centrifuge the plate at 300 x g for 10 min at room temperature, gently remove and discard 50% of the medium (100 μL), and add 100 μL of fresh "3D culture medium" to replace the existing solution. Repeat step 1.11, and place the plate back in the 37 °C, 5% CO2 humidified incubator.
NOTE: The removal of the medium must be performed when the plate is held at 45°, and the medium must be collected only from the upper part to avoid collecting the cells. A vacuum suction system should not be used. The fresh medium needs to be added slowly and gently in order not to disrupt the spheroids, which will have already started to form. - Inspect the cells under a microscope every 1-2 days to monitor spheroid formation. Measure the diameter of the spheroids formed using the "scale" tool in the imaging software (see the Table of Materials).
NOTE: Microscopic inspection should first reveal irregular round-to-oval bodies that assume a spheroid shape as time progresses11,12.- When the spheroid diameter reaches 100-200 μm, perform the drug efficacy experiments.
NOTE: The drug efficacy experiments are performed when the spheroids reach this diameter as, at that time, the majority of the cells are proliferating, which is conducive to evaluating the response to treatment. Larger diameter spheroids contain a necrotic core and a quiescent layer11,12, resulting in a smaller proportion of proliferating cells that respond to treatment.
- When the spheroid diameter reaches 100-200 μm, perform the drug efficacy experiments.
- For spheroid collection, use a 1,000 μL pipette to collect the spheroids from each well, and deposit them into a 15 mL conical tube.
NOTE: Spheroids are inherently fragile and require gentle handling. When transferring the medium with the spheroids into the conical tube, hold the tube at 45°, and pipette slowly on the wall of the tube. Spheroids are visible to the eye. - Centrifuge the conical tube at 300 x g for 5 min at room temperature, and carefully aspirate and discard the supernatant using a pipette.
- Add 0.5 mL of the cell culture medium, and resuspend the pellet well but gently. Avoid making bubbles.
- Perform spheroid counting following the steps below.
- Use a 96-well plate and draw a plus sign on the underside of a well to divide the well into quadrants (to help track the counting).
- Add 50 μL of the suspension to the well, and count the spheroids manually under the microscope (using a 10x objective lens).
- Count the spheroids in each quadrant, be careful not to double count, and calculate the total number of spheroids in the well.
NOTE: For accuracy of the counting, count at least 50 spheroids in a well. If there are less than 50 spheroids, gently mix the suspension to ensure homogenous distribution, and count again using a larger volume of the suspension added to a new well. Alternatively, if there are less than 50 spheroids in a well, the spheroids can be centrifuged and resuspended gently in a volume of <0.5 mL. If there are over 100 spheroids, add cell culture medium to the spheroid suspension, mix gently, and recount. - Calculate the spheroid concentration (spheroid count/counting volume [μL]), and then calculate the total number of spheroids in the suspension (spheroid concentration × total volume).
2. Drug efficacy assay (MTT assay)
NOTE: For details, please see van Meerloo et al.13. Also, for the MTT assay, only the cell culture medium and not the "3D culture medium" must be used (adding the basement membrane matrix is not necessary and could potentially interfere with the MTT assay).
- In a new tube, prepare a spheroid suspension at a concentration of 200 spheroids per 200 μL of cell culture medium (RPMI-1640 culture medium supplemented with 10% FBS, 1:100 penicillin-streptomycin, 1% non-essential amino acids, and 1% L-glutamine).
- For each drug treatment, prepare a stock of spheroids in a 15 mL tube. Calculate the amount needed for each drug by the number of wells needed for repeats: (5-8) × 200 μL. Add the drug to the tube to the final concentration needed.
NOTE: Please see the representative results section regarding the drugs used for the present study and their dosage. Also, the commercial details of the drugs are listed in the Table of Materials.
- For each drug treatment, prepare a stock of spheroids in a 15 mL tube. Calculate the amount needed for each drug by the number of wells needed for repeats: (5-8) × 200 μL. Add the drug to the tube to the final concentration needed.
- Transfer 200 μL of the spheroid suspension into the wells of an ultra-low attachment 96-well plate, and incubate the plate at 37 °C in a 5% CO2 humidified incubator. Do not use the external rows and columns of the 96-well plate for the assay, as these wells are characterized by increased evaporation, which could lead to increased variability between the repeat experiments; instead, add PBS to these wells.
NOTE: It is key that the culture of the spheroids is homogeneous before the culture is aliquoted into the 96-well plate. Also, a control condition (i.e., untreated spheroids) must be included in every experiment. - After incubating the spheroids with the study drug for 24-72 h, centrifuge the plate at 300 x g for 5 min at room temperature, and gently remove 170 μL of the cell culture medium, leaving 30 μL (which includes the spheroids) at the bottom the well.
NOTE: The removal of the 170 μL of cell culture medium must be done carefully, so as not to remove the spheroids. Hold the plate at 45°, and place one hand (wearing a dark glove for contrast) under the wells so that the spheroids are visible (white dots). - Prepare the MTT solution (0.714 mg/mL in phenol-free RPMI, see the Table of Materials).
- Add 70 μL of MTT solution to each well to a final volume of 100 μL per well (the final MTT concentration in the well will be 0.05 mg per 100 μL). In addition, prepare "Blank" wells with MTT solution without cells.
NOTE: MTT is light-sensitive. Therefore, the light in the hood must be switched off, and the tube containing the MTT solution must be covered with aluminum foil. - Incubate the plate at 37 °C in a 5% CO2 humidified incubator for 3-4 h until a change in the color of the solution in the wells (purple color represents live cells) is observed.
- When a change is observed, add 100 μL of stop solution (0.1N HCl in isopropanol) to each well, and gently mix the content of the wells without creating bubbles.
- Read the absorbance of the plate in a Fluorometer-ELISA reader (see the Table of Materials) at a wavelength of 570 nm and a background wavelength of 630-690 nM.
NOTE: If the available Fluorometer-ELISA reader cannot read the ultra-low attachment 96-well plate, transfer the content of each well to a corresponding flat-bottom 96-well plate. - Calculate the cell viability following the steps below.
- For each well, calculate the "specific signal" ("specific signal" = the signal at 570 nm – the signal at 630-690 nm). Then, calculate the average value of the "Blank" wells, and subtract this value from each well.
- Calculate the average of the "specific signals" in the control wells that contain cells that were not treated with the study drug ("AV-SS-unt").
- Calculate the viability (percentage) of cells in each well relative to the wells with untreated cells.
NOTE: Viability = (specific signal in each well/"AV-SS-unt") × 100
3. Monitoring and analyzing the morphological changes in the spheroids
NOTE: As for the MTT assay, only the cell culture medium and not the "3D culture medium" should be used in this evaluation (adding the basement membrane matrix is not necessary and could potentially interfere with the analysis).
- After counting the spheroids, dilute the suspension in cell culture medium (RPMI-1640 culture medium supplemented with 10% FBS, 1:100 penicillin-streptomycin, 1% non-essential amino acids, and 1% L-glutamine) to approximately 1-2 spheroids per 20 μL.
- Put 80 μL of cell culture medium into the wells of an ultra-low attachment 96-well plate, and then add 20 μL of the spheroid suspension to the wells.
NOTE: The wells will, therefore, contain 1-2 spheroids in a volume of 100 μL. - Carefully check the wells under the microscope, and mark the wells that contain one spheroid, as they will be used for this analysis.
- Prepare the study drug in a concentration that is two-fold the concentration of interest. Add 100 μL of the drug solution to the relevant wells (the total concentration in the well will then be 1x).
- Capture an image of each well at Day 0 (before adding the study drug) and use the "scale" tool in the imaging software (see the Table of Materials) to determine the diameters of the spheroids.
- Incubate the plate at 37 °C in a 5% CO2 humidified incubator, and examine the morphology and measure the diameter of the spheroids (using the "scale" tool) daily for 3-7 days, depending on when the drug effect is observed (e.g., cell dissemination, invasive pods, structure destruction, etc.).
- At the end of the experiment, plot the changes in spheroid diameters (relative to Day 0) over time.
NOTE: Change (percentage) = (spheroid diameter at a specific day/the diameter of that spheroid at Day 0) × 100
Representative Results
This protocol presents procedures for generating a homogenous culture of spheroids from primary tumor cells, quantitatively evaluating drug efficacy on spheroid culture (MTT assay), and determining the effect of study drugs on spheroid morphology. Data from the representative experiments in spheroids generated from colon and breast cancer cell cultures are presented. Similar experiments were performed using other tumor types, including cholangiocarcinoma, gastric, lung, and pancreatic cancer (data not shown). All the experiments presented herein were performed in triplicate.
Figure 2 shows the spheroids that were generated from the primary colon cancer cell culture. As seen in Figure 2, the number of spheroids generated depends on the number of cells initially seeded in each well. The growth of the spheroids to over 100 μm in diameter took 10-14 days. The origin of the tumor cells (e.g., different patients and different origins) determined the growth rate. Seeding the wells with more cells did not shorten the time required for spheroid generation but rather increased the number of spheroids formed. Notably, upon prolonged culture of the colon cancer spheroids, they started attaching to each other and formed clusters of spheroids in grape-like structures (Figure 3), which prevented a homogenous culture and, thus, prohibited the use of the spheroids in the MTT assays.
Figure 4 presents the effect of three treatments (10 μM palbociclib, 10 μM sunitinib, and their combination at 10 μM each) on the viability of spheroids derived from two primary cancers. In this case, a PDX model was established first, and the tumor cells used for the spheroid analysis were derived from the PDX model9. The first PDX model was established using a colon cancer sample from a 50 year old male patient, and the second using a breast cancer sample from a 62 year old female. As demonstrated in Figure 4A,B, after 3 days of treatment, the combination of palbociclib plus sunitinib led to a significant reduction in viability as measured by the MTT assay. As demonstrated in Figure 4C,D, the morphological changes occurring with treatment were very clear. On Day 0, all the spheroids were intact. In contrast, on Day 3, the spheroids treated with the control (DMSO) were still intact, whereas the spheroids treated with the combination were disassembled, and their morphology was "open", with cells detaching from the solid structure, suggesting the destruction of the spheroid structure.
Figure 5 presents the follow-up of the spheroids over time. These spheroids, generated from breast cancer cells derived from a 44 year old female patient, were treated with one of two combinations (trastuzumab [10 μg/mL] plus vinorelbine [1 μg/mL], or 5-fluorouracil [200 μM] plus cisplatin [300 μM]). As shown in Figure 5A, the size of the spheroids treated with 5-fluorouracil plus cisplatin was reduced by Day 3, and the spheroids were completely destroyed by Day 7. In contrast, the treatment with trastuzumab plus vinorelbine had only a minor effect on the morphology of the spheroids (e.g., some level of an "open" structure), but the effect was not significant. Figure 5B presents the average change in the diameter of the spheroids relative to Day 0 (five spheroids were followed in each treatment group).
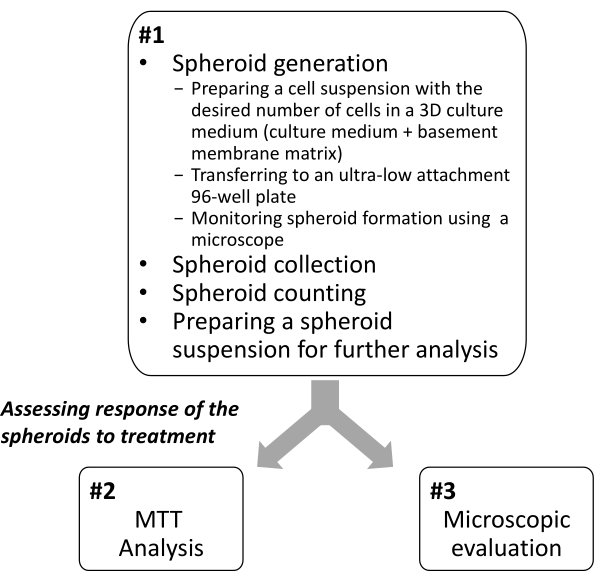
Figure 1: Overview of the protocol for establishing 3D spheroids from patient-derived tumor samples and evaluating their sensitivity to drugs. Please click here to view a larger version of this figure.
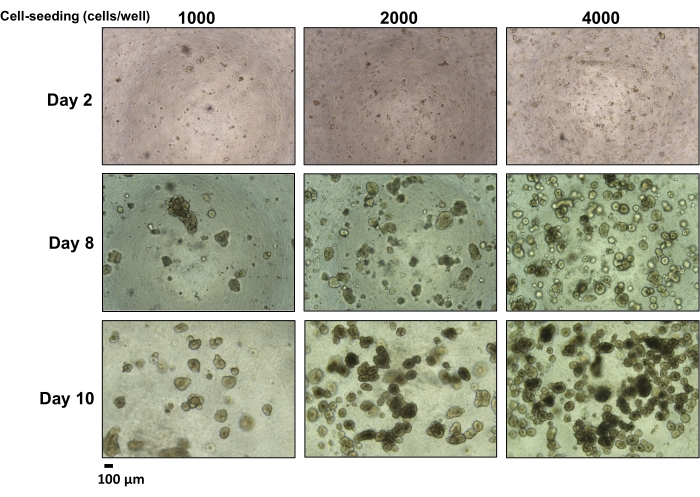
Figure 2: Formation of spheroids from a primary colon cancer cell culture over time by the number of initially seeded cells. Different numbers of cells were seeded in "3D culture medium" in an ultra-low attachment 96-well plate and observed under the microscope (4x magnification). Scale bar = 100 µm. Please click here to view a larger version of this figure.
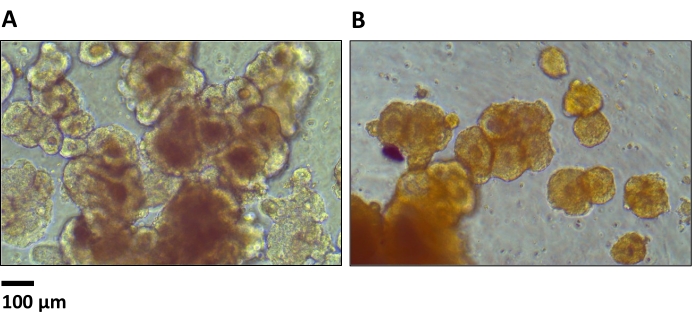
Figure 3: Spheroids from primary colon cancer cells (with initial cell seeding of 2,000 per well) after 12 days in culture. The two examples (A,B) show clusters created by the attachment of spheroids to each other (10x magnification). Scale bar = 100 µm. Please click here to view a larger version of this figure.
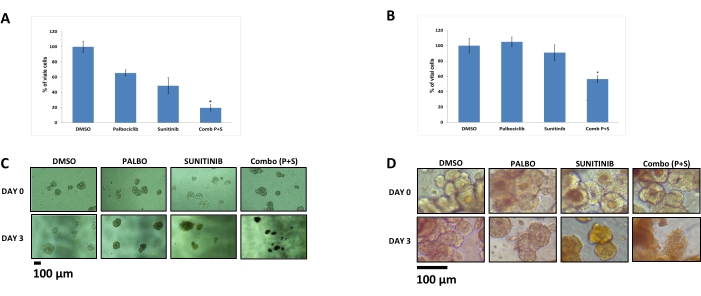
Figure 4: The effects of palbociclib (10 μM), sunitinib (10 μM), and their combination (10 μM each) on spheroids from primary tumor cells, including colon and breast cancer (PDX-derived). An MTT assay was conducted on spheroids derived from (A) colon and (B) breast cancer cells. The MTT signals were normalized to values from DMSO-treated cells. The values represent the means from four to eight replicates. The error bars represent SEM. *p < 0.05 versus a single agent (t-test). The effects of the various treatments on cell growth were also evaluated microscopically at Day 0 and after 3 days of treatment of the spheroids derived from (C) colon and (D) breast cancer cells (10x magnification). Scale bar = 100 µm. The figure is adapted from Moskovits et al.9. Please click here to view a larger version of this figure.
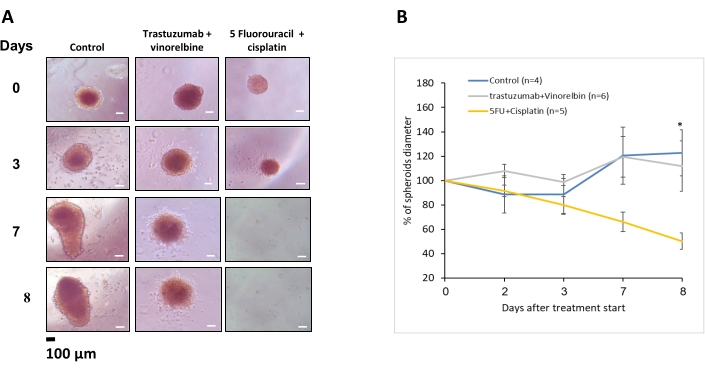
Figure 5: The effects of trastuzumab (10 μg/mL) plus vinorelbine (1 μg/mL) and 5-fluorouracil (200 μM) plus cisplatin (300 μM) on spheroids derived from breast cancer over time. (A) Each well included one spheroid and was monitored over time under the microscope (10x magnification). Scale bar = 100 µm. (B) The change in the diameter of the spheroids (relative to Day 0) by the duration of treatment. *p = 0.05 for 5-fluorouracil plus cisplatin versus controls (t-test). Each treatment group included four to six wells, with one spheroid in each well. The average change is presented. The error bars represent SEM. Please click here to view a larger version of this figure.
Discussion
The present protocol describes a simple method for generating 3D primary cell cultures (spheroids) derived from human tumor samples. These spheroids can be used for various analyses, including evaluating potential drug candidates and drug combinations, identifying potential biomarkers or therapeutic targets, and investigating the mechanisms of response and resistance. The protocol uses either primary tumor cells derived directly from patient samples or tumor cells from PDX models, which can be established using patient samples. The latter approach allows for conducting in vitro and in vivo experiments with the same primary tumor. Consistency has previously been shown in the results of drug sensitivity experiments between PDX models and 3D cultures derived from these models9, thus supporting the relevance of this in vitro/in vivo approach.
The main advantages of the current protocol include its wide applicability to most solid tumors and its cost-effectiveness, which stems from its compatibility with the typical capabilities/equipment of cancer research/cell biology labs (i.e., no need for specialized equipment or outsourcing). Additionally, the current protocol generates a homogenous spheroid population, which allows the use of high-throughput quantitative viability assays (e.g., MTT). Generating a homogenous spheroid population is important for obtaining meaningful results, as studies have demonstrated that the spheroid size affects the response to treatment. Larger spheroids, unlike smaller spheroids, are characterized by a necrotic core. However, most cells are in the linear growth stage in smaller spheroids. Furthermore, the size of the spheroid also affects the stiffness of its tissue structure, which could impact the diffusion of compounds (such as those used for the viability assays) into the spheroid14. The main limitation of the current protocol is that, even with its wide applicability, there are instances when the approach fails to generate spheroids. Importantly, such a failure is not tumor-type specific but rather patient-specific. Additional studies are required to explore why tumor samples from certain patients do not form spheroids using this protocol.
The current protocol is based on two key principles: (1) having a cell suspension with no cell clusters (i.e., a single-cell suspension) and (2) using an ultra-low attachment plate with a medium containing 5% basement membrane matrix (a solubilized extracellular matrix). The initial number of seeded cells affects the number of spheroids that are formed but not the time required for spheroid formation, suggesting that each spheroid is generated from a single tumor cell. Notably, the clustering of spheroids occurs, particularly after prolonged incubation. This clustering disrupts the homogenous culture and prohibits the use of MTT (due to the difficulty in dispensing an equal number of spheroids into each well). This clustering can be avoided by diluting the culture and transferring the spheroids to larger wells. If a homogenous culture cannot be achieved, MTT assays cannot be used, although the morphological assessment and the measurement of the spheroid diameters can still be performed. It must be noted that the morphological assessment is more labor-intensive since it requires the allocation of one spheroid per well and the monitoring of each spheroid under the microscope.
Determining the appropriate number of spheroids for an MTT assay is important for its interpretation. Thus, it is recommended to first generate a standard curve with known numbers of spheroids (e.g., 50, 100, 200, and 400 per well, in replicates) in order to determine the optimal number of spheroids for the MTT assay. The middle of the linear range of the plot must be used for the analysis so that there are enough spheroids to detect a signal but not too many (i.e., so that the plateau phase of the signal is not reached). Furthermore, using the middle range allows for keeping the signal within the linear range in cases of response to the drug (i.e., reduced signal), as well as non-response (i.e., the continued growth of the spheroid and increased signal). Lastly, since the MTT assay assesses cell metabolic activity, which could be different between tumors from different patients, a standard curve should be generated for each primary tumor sample.
In summary, this protocol for generating 3D tumor culture models from primary cancer cells and evaluating their sensitivity to drugs using a cell-viability assay (MTT) and morphologic examination under the microscope represents a valuable, biologically relevant tool that complements the current 2D in vitro approaches as well as the in vivo approaches.
Disclosures
The authors have nothing to disclose.
Acknowledgements
None.
Materials
| 5 Fluorouracil | TEVA Israel | lot 16c22NA | Fluorouracil, Adrucil |
| Accutase | Gibco | A1110501 | StemPro Accutase Cell Dissociation |
| Cisplatin | TEVA Israel | 20B06LA | Abiplatin, |
| Cultrex | Trevigen | 3632-010-02 | Basement membrane matrix, type 3 |
| DMSO (dimethyl sulfoxide) | Sigma Aldrich | D2650-100ML | |
| Fetal bovine serum (FBS) | Thermo Fisher Scientific | 2391595 | |
| Flurometer ELISA reader | Biotek | Synergy H1 | Gen5 3.11 |
| Hydrochloric acid (HCl) | Sigma Aldrich | 320331 | for stop solution |
| ImageJ | National Institutes of Health, Bethesda, MD, USA | Version 1.52a | Open-source software ImageJ |
| Isopropanol | Gadot | P180008215 | for stop solution |
| L-glutamine | Gibco | 1843977 | |
| MTT | Sigma Aldrich | M5655-1G | 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide |
| Non-essential amino acids | Gibco | 11140050 | |
| Palbociclib | Med Chem Express | CAS # 571190-30-2 | |
| PBS | Gibco | 14190094 | Dulbecco's Phosphate Buffered Saline (DPBS)*Without Calcium and Magnesium |
| Penicillin–streptomycin | Invitrogen | 2119399 | |
| Phenol-free RPMI 1640 | Biological industries, Israel | 01-103-1A | |
| Pippeting reservoir | Alexred | RED LTT012025 | |
| RPMI-1640 culture medium | Gibco | 11530586 | |
| Sunitinib | Med Chem Express | CAS # 341031-54-7 | |
| Trastuzumab | F. Hoffmann – La Roche Ltd, Basel, Switherland | 10172154 IL | Herceptin |
| Trypan blue 0.5% solution | Biological industries, Israel | 03-102-1B | |
| Ultra-low attachment 96 well plate | Greiner Bio-one | 650970 | |
| Vinorelbine | Ebewe | 11733027-03 | Navelbine |
References
- Katt, M. E., Placone, A. L., Wong, A. D., Xu, Z. S., Searson, P. C. In vitro tumor models: Advantages, disadvantages, variables, and selecting the right platform. Frontiers in Bioengineering and Biotechnology. 4, 12 (2016).
- Yoshida, G. J. Applications of patient-derived tumor xenograft models and tumor organoids. Journal of Hematology & Oncology. 13 (1), 4 (2020).
- Ledur, P. F., Onzi, G. R., Zong, H., Lenz, G. Culture conditions defining glioblastoma cells behavior: What is the impact for novel discoveries. Oncotarget. 8 (40), 69185-69197 (2017).
- Richter, M., et al. From donor to the lab: A fascinating journey of primary cell lines. Frontiers in Cell and Developmental Biology. 9, 711381 (2021).
- Esparza-Lopez, J., Martinez-Aguilar, J. F., Ibarra-Sanchez, M. J. Deriving primary cancer cell cultures for personalized therapy. Revista de Investigación Clínica. 71 (6), 369-380 (2019).
- Choi, J. R., et al. In vitro human cancer models for biomedical applications. Cancers. 14 (9), 2284 (2022).
- Eglen, R. M., Randle, D. H. Drug discovery goes three-dimensional: Goodbye to flat high-throughput screening. Assay and Drug Development Technologies. 13 (5), 262-265 (2015).
- Kodack, D. P., et al. Primary patient-derived cancer cells and their potential for personalized cancer patient care. Cell Reports. 21 (11), 3298-3309 (2017).
- Moskovits, N., et al. Palbociclib in combination with sunitinib exerts a synergistic anti-cancer effect in patient-derived xenograft models of various human cancers types. Cancer Letters. 536, 215665 (2022).
- Ricardo, R., Phelan, K. Counting and determining the viability of cultured cells. Journal of Visualized Experiments. (16), e752 (2008).
- Brajša, K., Trzun, M., Zlatar, I., Jelić, D. Three-dimensional cell cultures as a new tool in drug discovery. Periodicum Biologorum. 118 (1), 59-65 (2016).
- Han, S. J., Kwon, S., Kim, K. S. Challenges of applying multicellular tumor spheroids in preclinical phase. Cancer Cell International. 21 (1), 152 (2021).
- van Meerloo, J., Kaspers, G. J., Cloos, J. Cell sensitivity assays: The MTT assay. Methods in Molecular Biology. 731, 237-245 (2011).
- Walzl, A., et al. The resazurin reduction assay can distinguish cytotoxic from cytostatic compounds in spheroid screening assays. Journal of Biomolecular Screening. 19 (7), 1047-1059 (2014).

