Dissection of Adult Mouse Stria Vascularis for Single-Nucleus Sequencing or Immunostaining
Summary
The stria vascularis is vital to the generation of endocochlear potential. Here, we present the dissection of the adult mouse stria vascularis for single-nucleus sequencing or immunostaining.
Abstract
Endocochlear potential, which is generated by the stria vascularis, is essential to maintain an environment conducive to appropriate hair cell mechanotransduction and ultimately hearing. Pathologies of the stria vascularis can result in a decreased hearing. Dissection of the adult stria vascularis allows for focused single-nucleus capture and subsequent single-nucleus sequencing and immunostaining. These techniques are used to study stria vascularis pathophysiology at the single-cell level.
Single-nucleus sequencing can be used in the setting of transcriptional analysis of the stria vascularis. Meanwhile, immunostaining continues to be useful in identifying specific populations of cells. Both methods require proper stria vascularis dissection as a prerequisite, which can prove to be technically challenging.
Introduction
The cochlea consists of three fluid filled chambers, the scala vestibuli, scala media, and scala tympani. The scala vestibuli and scala tympani each contain perilymph, which has a high concentration of sodium (138 mM) and a low concentration of potassium (6.8 mM)1. The scala media contains endolymph, which has a high concentration of potassium (154 mM) and a low concentration of sodium (0.91 mM)1,2,3. This difference in ion concentration can be referred to as the endocochlear potential (EP), and is primarily generated by the movement of potassium ions through various ion channels and gap junctions in the stria vascularis (SV) along the lateral wall of the cochlea4,5,6,7,8,9,10,11. The SV is a heterogenous, highly vascularized tissue that lines the medial aspect of the lateral wall of the cochlea and contains three main cell types: marginal, intermediate, and basal cells12 (Figure 1).
Marginal cells are connected by tight junctions to form the most medial surface of the SV. The apical membrane faces the endolymph of the scala media and contributes to potassium ion transport into the endolymph using various channels, including KCNE1/KCNQ1, SLC12A2, and Na+–K+-ATPase (NKA)5,10,13,14. Intermediate cells are pigmented cells that reside between marginal and basal cells and facilitate potassium transport through the SV using KCNJ10 (Kir 4.1)15,16. Basal cells lie in close proximity to the lateral wall of the cochlea and are closely associated with fibrocytes of the spiral ligament to promote potassium recycling from the perilymph12. Pathology of the SV has been implicated in numerous otologic disorders17,18. Mutations in genes expressed in the major SV cell types, such as Kcnq1, Kcne1, Kcnj10, and Cldn11, can cause deafness and SV dysfunction, including the loss of EP19,20,21,22,23. In addition to the three major cell types, there are other less-studied cell types in the SV, such as spindle cells22, root cells12,24, macrophages25, pericytes26, and endothelial cells27, that have incompletely defined roles involving ionic homeostasis and the generation of EP28.
In comparison to bulk RNA-sequencing, single-nuclei RNA-sequencing (sNuc-Seq) provides information about cell heterogeneity, rather than an average of mRNA across a group of cells29, and can be particularly useful when studying the heterogenous SV30. For example, sNuc-Seq has produced transcriptional analysis that suggests there may be a role for spindle and root cells in EP generation, hearing loss, and Meniere's disease18. Further transcriptional characterization of the various SV cell types can provide us with invaluable information on the pathophysiology underlying different mechanisms and subtypes of SV-related hearing fluctuation and hearing loss. The harvest of these delicate inner ear structures is of paramount importance to optimal tissue analysis.
In this study, the microdissection approach to access and isolate the stria vascularis from the adult mouse cochlea for sNuc-Seq or immunostaining is described. Dissection of the adult mouse SV is required to understand various SV cell types and further characterize their role in hearing.
Protocol
All animal experiments and procedures were performed according to protocols approved by the Animal Care and Use Committee of the National Institute of Neurological Diseases and Stroke and the National Institute on Deafness and Other Communication Disorders, National Institutes of Health. All experimental protocols were approved by the Animal Care and Use Committee of the National Institute of Neurological Diseases and Stroke and the National Institute on Deafness and Other Communication Disorders, National Institutes of Health. All methods were carried out in accordance with relevant guidelines and regulations of the Animal Care and Use Committee of the National Institute of Neurological Diseases and Stroke and the National Institute on Deafness and Other Communication Disorders, National Institutes of Health.
1. Animal euthanasia
- Use C57BL/6J or Kcnj10-ZsGreen transgenic mice, postnatal day 30+ (P30+), that weigh between 15-20 g.
- Euthanize an adult mouse (age P30 or older) using an approved protocol (here CO2 asphyxiation). Perform decapitation as a secondary step.
NOTE: If immunostaining, some antibodies may have background signal due to nonspecific binding to blood in the capillaries of the SV. Cardiac perfusion with fixative can address this issue by removing blood from the capillaries of the SV. However, most antibodies can achieve good results without this method, and it will not be covered in this protocol.
2. Exposing bony labyrinth
- Clean the mouse by spraying with 70% ethanol and wiping with a paper towel. Pay particular attention to the head region to reduce the risk of contamination. Remove the skin from the skull by pulling the skin toward the nose.
- Using sharp scissors, split the skull along the midsagittal plane to separate the left and right portions.
- Remove the brain from the skull to expose the bony labyrinth.
3. Inner ear extraction
- Identify the inner ear within the temporal bone (Figure 2A) and scrape away the cranial nerves using #55 forceps.
NOTE: The user should stabilize the specimen with their nondominant hand using a second set of #55 forceps. Different areas of the specimen may be grabbed with the stabilizing forceps throughout the dissection if critical areas of the specimen are avoided (e.g., do not crush the cochlea). - Using #55 forceps, dissect out the bulla and capsule, detaching them from surrounding bone. Remove any remaining soft tissue from the inner ear surface.
- Transfer the specimen to a clear tissue culture dish containing 1x cold PBS (phosphate-buffered saline), pH 7.4, and place under a dissection scope (Figure 2B).
4. SV dissection
NOTE: With practice, it is possible to dissect the SV as one long piece resembling a ribbon. The SV is fragile, so if it breaks into pieces, these may be stored together. Alternatively, these can be stored in separately labeled wells according to their turn (e.g., basal, middle, apical).
- If the dissection scope light sources can be moved, place one light above the dish and the second light source from the side, parallel to the dissecting surface. This allows for a better contrast of the tissue under the cochlear bone.
- Using #55 forceps, hold the specimen by the vestibular portion with the cochlea facing up.
- Using #55 forceps, pierce through the cochlear bone at the apex.
NOTE: #5 forceps are thicker than #55 and may be used for breaking bone, but the increase in size/strength sacrifices the fineness of the forceps and the ability to manipulate small tissues. Consider using forceps made with a material such as inox or dumostar to avoid damage that may occur to more delicate forceps. Avoid pushing the forceps too deep under the bone to avoid damaging the SV. - Using #55 forceps, scrape along the apical turn (Figure 2C), applying gentle force to break away small pieces of the outer bone layer. Gently lift the bone and detach it from the lateral wall.
NOTE: It may be helpful to think about this dissection as "removing everything away from the SV", rather than "dissecting the SV out of the cochlea". - Continue using #55 forceps to remove cochlear bone pieces from the apical and middle turns (Figure 2D,E). Gently pushing down the lateral wall, pry and remove bone wall pieces toward the middle turn to expose the lateral wall of the apical and middle turns.
- Continuing using #55 forceps, gently push the apical turn lateral wall aside to expose the spiral ganglion. Detach the lateral wall of apical and middle turns from the spiral ganglion along the outer hair cell layer. Remove pieces of spiral ganglion from inside the cochlea.
NOTE: #55 forceps are smaller and provide an advantage when manipulating small and delicate tissues. Extra care should be taken to avoid bending or damaging them. - To have better access to the lateral wall of the basal turn, detach the cochlea from the vestibular portion of the temporal bone using #55 forceps. Put the #55 forceps into the round and oval window and push down toward the vestibule. The cochlea will detach. Remove the vestibular portion of the inner ear.
- Continuing using #55 forceps, now remove pieces of basal cochlear bone, starting from the middle turn area. Gently push the lateral wall layer under the bone to detach it.
- Remove the farthest basal part of the lateral wall by prying and gently pulling the tissue. The bone in this region may be too thick to be removed without damaging the soft tissue below.
- Now, with the basal part of the lateral wall detached, separate as much of the lateral wall from the cochlea as possible. This can be done by tracing the #55 forceps along the lateral wall. Gently brush the forceps between the bone and the remaining lateral wall. This can be done all the way to the apex in one piece if the user is experienced. Move the lateral wall to fresh PBS.
NOTE: Although larger pieces of SV may be easier to image, given the delicate nature of the tissues, smaller pieces can be taken (Figure 2F,G), and depending on the size, can also work for imaging. - Use #55 forceps to lay the lateral wall flat. The SV should be visible as a darker layer of tissue on the internal side of the lateral wall.
NOTE: Users may find, particularly closer to the apical end, that some of the SV tissue incidentally detaches from the lateral wall throughout the dissection. - Gently pry the SV layer to detach it from the lateral wall at its basal end (Figure 2H). Gently push aside the detached SV and move the forceps further toward the apical end of the lateral wall, detaching the SV as one long ribbon if possible (Figure 2I). Do not squeeze the SV with forceps to pull it. If the tissue is too fragile to be grabbed, it can alternatively be collected using a tissue scoop, such as a chalazion curette.
NOTE: Moving the SV should always be done under light microscopy to ensure it is not lost. Always be careful when grabbing using forceps due to the risk of damage. Users may find that using chalazion curette mitigates the risk of damage to SV tissue. For single-nucleus sequencing, proceed to section 5. For SV immunostaining, proceed to section 7.
5. SV single-nucleus suspension
NOTE: This protocol is adapted for SV tissue specifically from a published manufacturer's single-nucleus suspension protocol. Platforms from different manufacturers may be used31. Given platform-specific variation, it is recommended to review the manufacturer-specific protocol provided with the equipment. To achieve optimal results for sNuc-Seq and minimize RNA degradation, the faster the tissue dissection the better (recommended within 15-20 min from euthanasia). It may be helpful to euthanize one animal at a time and only when ready to dissect. Having multiple people simultaneously work on the dissections can also eliminate degradation time (e.g., one lab personnel working on the left ear while another works on right ear).
- If the SV tissue will be used for sNuc-Seq, place the tissue in chilled lysis buffer (10 mM Tris-HCl, 10 mM NaCl, 3 mM MgCl2, 0.005% nonidet P40 in nuclease free water).
NOTE: If sequencing is to be done immediately following dissection, continue the protocol. If it is desired to pause the protocol, SV preservation is possible by placing the SV into RNAlater. Store overnight at 4 °C, then flash freeze in liquid nitrogen and store at -80 °C until ready to use. When ready to use, thaw and wash twice in PBS for 5 min each, then proceed with homogenization (step 5.2). - Homogenize the tissue in a 2 mL dounce homogenizer with 10-20 strokes on ice.
NOTE: The glass cylinder will manually contact the bottom of the glass tube, fragmenting the SV. The SV is delicate, so 20 strokes usually achieves successful homogenization. - Lyse the tissue on ice for 25 min.
NOTE: The lysis time varies according to the tissue type. A 25 min lysis in SV tissue may achieve a low cell viability (less than 4%), but the time may be adapted if the cell viability is too high (step 5.8). - Filter through a 30 µm filter and spin the filtrate at 500 x g for 5 min at 4 °C.
- Remove the supernatant and resuspend the cell pellet in 1 mL of nuclei wash and resuspension buffer (1x PBS, 1% bovine serum albumin [BSA], 0.2U/µL RNase inhibitor).
- Filter the cells through a 10 µm filter and centrifuge at 500 x g for 5 min at 4 °C.
- Remove the supernatant and resuspend in 50 µL of nuclei wash and resuspension buffer.
- Count the nuclei using a cell counter and trypan blue staining. Dilute 5 µL of the cell suspension into 50 µL of 1x PBS for cell counting; The viability should be minimal (3%-4%) at this point. If the viability is found to be higher, adapt the protocol for step 5.3 (lysis) and standardize the lysis time to ensure adequate lysis of the cells.
NOTE: A high cell viability in a single nucleus preparation means there are more intact cells than desired, and that additional digestion may be required. - Load the sample with the desired nuclear density onto the manufacturer's apparatus/chip (see manufacturers guide). Single-nuclei captures are now ready.
NOTE: From one adult mouse SV, 50 µL of the suspension at a concentration of 150,000 nuclei/mL is usually achieved.
6. SV single-nucleus sequencing
- Send the samples to the core facility for sequencing.
NOTE: This part of the protocol follows the general steps of (1) gel beads-in emulsion (GEM) generation and barcoding, (2) post-GEM-RT cleanup and cDNA amplification, and (3) 3' gene expression library construction. The remainder of the sNuc-Seq protocol is relatively lengthy, and it is recommended that readers use their manufacturer-specific protocol. The manufacturer guide will likely describe steps with specific details and accompanying images. There may also be 'how-to' videos on the manufacturer website. Datasets generated may be represented in a dimensional reduced fashion, such as a uniform manifold approximation and projection (UMAP; Figure 3). Many labs utilize a sequencing core that may perform this protocol for users.
7. SV immunostaining and tissue mounting
- If the tissue will be used for immunostaining, transfer it to a 24-well plate, submerged in 200 µL of 4% paraformaldehyde (PFA) in 1x PBS, and incubate for 20 min at room temperature.
NOTE: It is helpful to perform all liquid removal, washing, and immunostaining underneath a microscope. Visualization while pipetting the tissue will ensure it is not accidently lost during liquid removal. The volumes of the antibodies and washes may be adjusted as long as the volume is large enough to submerge the SV tissue within the solution. - After the fixation step, remove the PFA by performing two short washes in 1x PBS (approximately 500 µL) at 4 °C for 5 min each on an orbital shaker at low (30-50) RPM. If storing, the tissue can be stored in 1x PBS at 4 °C.
- Remove the PBS and permeabilize and block for a minimum of 1 h at room temperature in approximately 300 µL of PBS-T solution (0.05 Tween20 in 1x PBS with 10% fetal bovine serum).
NOTE: Primary and secondary antibodies should be diluted in this blocking solution. - Stain the tissue with primary antibody according to the specifications of the particular antibody used, usually overnight at 4 °C . Remove the remaining PBS and add the diluted primary antibody.
NOTE: The dilution should be chosen based on the manufacturer suggestions, published data, previous experience, etc. Usually, the first concentration to test is a 1:100-200 dilution. For the example presented in this paper, the GS-IB4 and DAPI antibodies were each diluted at 1:200. If staining for more than one protein, multiple primary antibodies can be combined into the same solution, as long as each primary antibody has a different host to avoid cross reactivity of secondary antibodies. - Wash the primary antibody off the tissue using approximately 500 µL of 1x PBS twice for 10 min each on an orbital shaker at low RPM.
NOTE: Continue to ensure that the SV tissue is submerged in the solution, and not stuck on the side of the well. - Stain with a secondary antibody according to the specifications of the particular antibody used, usually for 2 h at room temperature on an orbital shaker at low RPM. Cover the 24-well plate so the fluorescently tagged secondary antibody is protected from light.
NOTE: If more than one secondary antibody is needed, combine the secondary antibodies into the same solution. Ensure to avoid cross labeling. - Wash the secondary antibody off the tissue using approximately 500 µL of 1x PBS four times for 5 min each on an orbital shaker at low RPM. Keep the 24-well plate covered to protect it from light.
- Prepare the larger microslide (75 mm x 25 mm) by placing two small streaks of glue along the 25 mm axis, just smaller than the 18 mm x 18 mm smaller glass coverslip. Place one drop of mounting reagent (approximately 50 µL) in between the glue streaks.
NOTE: The glue creates a small amount of vertical space so that when the second coverslip is placed on top, the SV tissue sample can safely exist, but continue to remain in place. While the stria thickness is 30-40 microns, squeezing the tissue between the glass surfaces should be avoided to preserve morphology. Alternatively, clear tape may also be used. - Using #55 forceps, grab as little tissue as possible on one end of the SV and transfer from the PBS to the drop of mounting reagent on the larger microslide (75 mm x 25 mm). Place one end of a smaller glass coverslip (18 mm x 18 mm) on the slide where one streak of glue was placed and gently release to avoid creating air bubbles. The coverslip will fall to the other glue streak and will cover the SV tissue.
- Seal the mount by dabbing a drop of transparent nail polish on each corner of the mount. Label the specimen accordingly on the glass coverslip away from the visualization field. It is now ready for confocal microscopy (Figure 4).
- Visualize the SV tissue using a dissection microscope to ensure it is laying flat and not near any air bubbles. If the SV tissue is not oriented correctly, the user can attempt to push out air bubbles or carefully remove the smaller glass coverslip and reposition the SV. This must be done gently to avoid breaking the glass coverslips.
Representative Results
We present a method to isolate the SV to be used for either sNuc-Seq or immunostaining. The relevant anatomy (Figure 1) of the cochlea relative to the SV can help users better understand the organization of the SV and steps of the dissection protocol.
Each step of this microdissection of SV from a P30 mouse is detailed in the associated video, and snapshots of the key steps of this dissection and isolation of SV are presented in Figure 2.
sNuc-Seq is useful in investigating the transcriptional profile of the various cells in the heterogenous SV. One way this may be visualized is by clustering on a 2D UMAP (Figure 3). The dataset generated from sNuc-Seq may be evaluated using different data analysis techniques, further outlined in the discussion.
SV whole mounting with GS-IB4 and DAPI immunolabeling, along with Kcnj10-ZsGreen fluorescence, are presented in Figure 4. Using florescence, the SV can be visualized after removing the surrounding tissues and mounting. In these mice, ZsGreen is expressed particularly in the SV intermediate cells. The vasculature of the SV can be visualized in red (endothelial cells, GS-IB4).
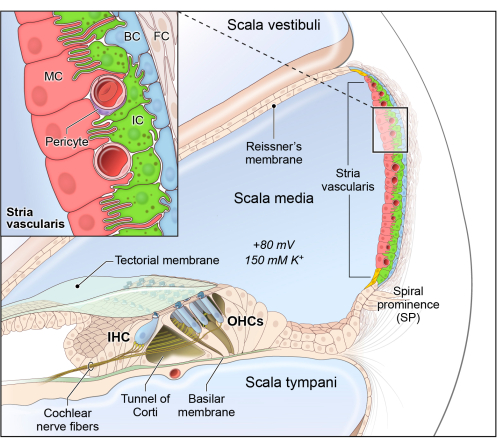
Figure 1: Schematic of the stria vascularis cellular heterogeneity and organization. Stria vascularis cellular heterogeneity and organization. Schematic of the stria vascularis (SV) and its relationship to structures in the cochlea. The SV is composed of three layers of cells and is responsible for generating EP and high potassium concentration in the endolymph-containing scala media. The relationship between the marginal, intermediate, and basal cells are demonstrated with the marginal extending basolateral projections to interdigitate with the intermediate cells, which have bidirectional cellular projections that interdigitate with both the marginal and basal cells. In addition to these cell types, other cell types, including spindle cells (yellow), endothelial cells, pericytes, and macrophages (not shown) are present in the SV. Used with permission from Korrapati et al.30. Please click here to view a larger version of this figure.
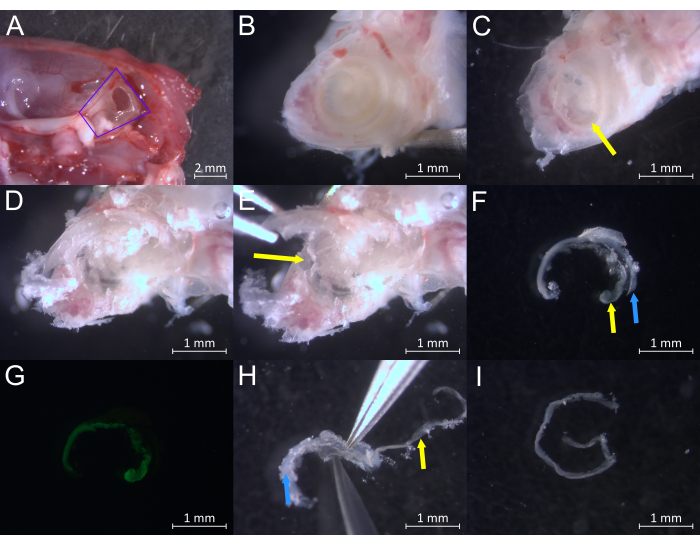
Figure 2: Dissection of an adult mouse cochlea at greater than or equal to postnatal day 30 (P30+). (A) Specimen during inner ear extraction from the surrounding temporal bone during step 3.2. The inner ear is outlined in purple. (B) Specimen during step 4.2 showing the surface of the cochlea and underlying pigmented SV. (C) Specimen during step 4.4 showing bone removed from the apical turn and the exposed SV (yellow arrow). (D) Specimen during step 4.5 showing the cochlea with more bone removed from the apical and middle turns. (E) Specimen during step 4.5 showing the removal of bone covering the middle turn, showing the underlying SV (yellow arrow). (F) Example of smaller pieces of SV (yellow arrow) and the lateral wall (blue arrow) during step 4.10. (G) Example of fluorescent ZsGreen SV while the remaining lateral wall remains dark. Kcnj10-ZsGreen is particularly expressed in the intermediate cells of the SV. (H) Example of larger pieces of SV (yellow arrow) and the lateral wall (blue) being separated during step 4.12. (I) The SV fully separated from the lateral wall during step 4.12. Please click here to view a larger version of this figure.
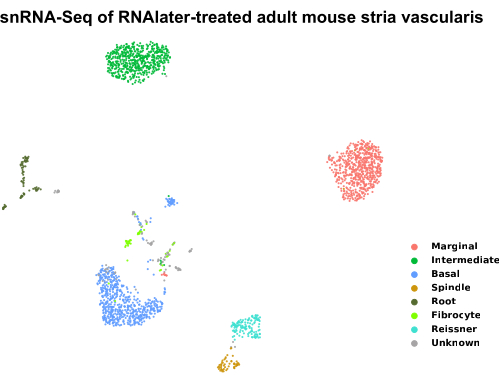
Figure 3: UMAP plot of sNuc-Seq datasets from an RNAlater-treated adult stria vascularis. Samples preserved with RNAlater treatment of adult SV tissue with subsequent nuclei isolation were clustered by a modularity-based clustering method with the Leiden optimization algorithm and visualized in a dimensionally reduced fashion by a 2D UMAP plot. Each dot represents a single cell, and cells with similar transcriptional profiles are clustered together. Cell types are colored based on their expression of known genes expressed by adult SV cell types. The samples were stored in RNAlater for no more than 6 months. The SV from about five CBA/J P30 mice were used to generate this dataset. Please click here to view a larger version of this figure.
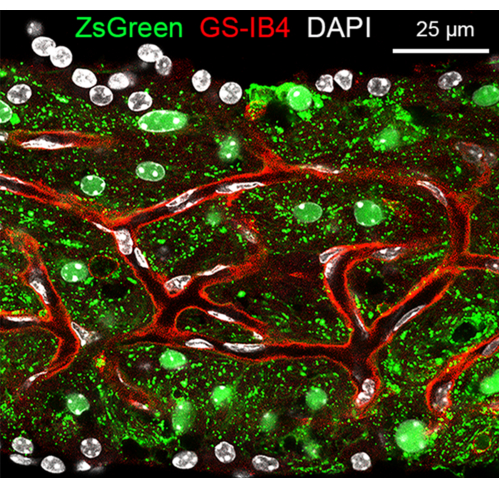
Figure 4. Stria vascularis whole mounting from a P30 Kcnj10-ZsGreen mouse. Confocal image of SV whole mounting from a P30 mouse, demonstrating ZsGreen expression in the intermediate cells, GS-IB4 (red) labeling endothelial cells, and DAPI (white) labeling nuclei. Please click here to view a larger version of this figure.
Discussion
Prior to the advent of single-cell sequencing, many researchers used bulk tissue analysis, which only made it possible to analyze transcriptomes averaged across cells. In particular, single-cell and sNuc-Seq made it possible to isolate the transcriptome of a single cell or single nucleus, respectively32. In this instance, single-nucleus transcriptomes can be identified for marginal, intermediate, and basal cells, as well as spindle cells30. This enables the investigation of transcriptional heterogeneity amongst SV cell types and can be used in the future to investigate the contribution of these cell types to SV function, including the generation and maintenance of EP. Datasets generated using sNuc-Seq can be displayed in a dimensionally reduced fashion to facilitate the understanding of overall transcriptional differences and compare different cell populations using a UMAP plot Figure 4. There are numerous ways to conduct bioinformatic analysis for the interpretation of sNuc-Seq datasets. These include but are not limited to: differential expression analysis, single-cell regulatory network inference and clustering (SCENIC), heatmap or grin violin plot construction, and regulon analysis18. Further computational analysis techniques for single-cell RNA sequencing are similar to sNuc-Seq, and are discussed in more detail in reviews by Hwang et al.32, Shafer33, and Chen et al.34.
The other major application of this dissection technique is immunostaining to better study selected structures within the SV. We can observe this in the setting of an SV whole mount that was imaged using confocal microscopy (Figure 3). The nuclei of all cells are visualized with DAPI labeling. The capillaries are seen via endothelial cell staining with GS-IB4 in red. Intermediate cells, in this case, have been genetically modified using an intermediate cell-specific promoter to express ZsGreen.
Limitations of sNuc-Seq include: (1) lack of spatial information, (2) biases against immune cell populations, and (3) exclusion of cytoplasmic RNA. Spatial transcriptomic techniques can provide more specific information about cellular subpopulations within tissue35. Given protocol differences between sNuc-Seq and single-cell RNA sequencing, such as tissue dissociation and storage, the sNuc-Seq-generated transcriptional profile may bias against immune cells while being more powerful at characterizing attached cell types36. Also, single-cell RNA sequencing includes cytoplasmic RNA such as mitochondrial RNA, while sNuc-Seq does not. Despite these instances of transcriptional differences between the two methods, it has been demonstrated that, by and large, sNuc-Seq has a similar transcriptional profile to whole single-cell RNA sequencing37, and in the SV in particular30.
For successful microdissection, there are several critical steps to be aware of. In the first few steps, one must properly extract the inner ear to avoid crushing the underlying cochlea. Observation of proper temporal bone extraction is shown in Figure 2A. Once the cochlea is properly exposed, care must be taken to gently scrape and break the cochlear bone to expose the underlying lateral wall and SV. Once the lateral wall and SV are extracted successfully, the user must carefully detach the SV from the lateral wall. Enough force in the correct plane should be applied to pry the SV from the lateral wall, while avoiding damage to the fragile SV. For SV microdissection, it may be helpful to think of “dissecting other structures away from the SV”, rather than “dissecting the SV out of the cochlea”.
This protocol can be modified for different ages of mice and for different mammalian species, such as rats. For younger mice, tissues will differ in texture and structures will be smaller in general. For different mammalian species, the cochlear anatomy will be relatively similar, with the size of the animal determining the majority of the difference.
In general, drawbacks of this protocol are similar to other cochlear microdissection techniques38. Isolating the delicate SV can be technically challenging and it can be broken into smaller pieces.
This technique is not optimal for saving the organ of Corti for whole mounting as it is often damaged with this protocol. There exists other dissection techniques specifically aimed at preserving the organ of Corti for whole mounting and immunostaining38, but these have their own drawbacks, considering sNuc-Seq must be done with fresh samples to avoid RNA degradation, not in the setting of tissue fixation and decalcification.
With time and repetition, users will become more comfortable with this SV microdissection protocol.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This research was supported in part by the Intramural Research Program of the NIH, NIDCD to M.H. (DC000088)
Materials
| 10-µm filter (Polyethylenterephthalat) | PluriSelect | #43-50010-01 | Filter tissue during sNuc-Seq |
| 18 x 18 mm cover glass | Fisher Scientific | 12-541A | Cover slip to mount SV |
| 30-µm filter (Polyethylenterephthalat) | PluriSelect | #43-50030-03 | Filter tissue during sNuc-Seq |
| 75 x 25 mm Superfrost Plus/Colorforst Plus Microslide | Daigger | EF15978Z | Microslide to mount SV on |
| C57BL/6J Mice | The Jackson Laboratory | RRID: IMSR_JAX:000664 | General purpose mouse strain that has pigment more easily seen in the intermediate cells of the SV. |
| Cell Counter | Logos Biosystems | L20001 | Used for cell counting |
| Chalizon curette 5'', size 3 2.5 mm | Biomedical Research Instruments | 15-1020 | Used to transfer SV |
| Chromium Next GEM single Cell 3' GEM Kit v3.1 | Chromium | PN-1000141 | Generates single cell 3' gene expression libraries |
| Clear nail polish | Fisher Scientific | NC1849418 | Used for sealing SV mount |
| Corning Falcon Standard Tissue Culture Dishes, 24 well | Corning | 08-772B | Culture dish used to hold specimen during dissection |
| DAPI | Invitrogen | D1306, RRID: AB_2629482 | Stain used for nucleus labeling |
| Dounce homogenizer | Sigma-Aldrich | D8938 | Used to homogenize tissue for sNuc-seq |
| Dumont #5 Forceps | Fine Science Tools | 11252-30 | General forceps for dissection |
| Dumont #55 Forceps | Fine Science Tools | 11255-20 | Forceps with fine tip that makes SV manipulation easier |
| Fetal Bovine Serum | ThermoFisher | 16000044 | Used for steps of sNuc-Seq |
| Glue stick | Fisher Scientific | NC0691392 | Used for mounting SV |
| GS-IB4 Antibody | Molecular Probes | I21411, RRID: AB-2314662 | Antibody used for capillary labeling |
| KCNJ10-ZsGreen Mice | n/a | n/a | Transgenic mouse that expresses KCNJ10-ZsGreen, partiularly in the intermediate cells of the SV. |
| MgCl2 | ThermoFisher | AM9530G | Used for steps of sNuc-Seq |
| Mounting reagent | ThermoFisher | #S36940 | Mounting reagent for SV |
| Multiwell 24 well plate | Corning | #353047 | Plate used for immunostaining |
| NaCl | ThermoFisher | AAJ216183 | Used for steps of sNuc-Seq |
| Nonidet P40 | Sigma-Aldrich | 9-16-45-9 | Used for steps of sNuc-Seq |
| Nuclease free water | ThermoFisher | 4387936 | Used for steps of sNuc-Seq |
| Orbital shaker | Silent Shake | SYC-2102A | Used for steps of immunostaining |
| PBS | ThermoFisher | J61196.AP | Used for steps of immunostaining and dissection |
| RNA Later | Invitrogen | AM7021 | Used for preservation of SV for sNuc-Seq |
| Scizzors | Fine Science Tools | 14058-09 | Used for splitting mouse skull |
| Tris-HCl | Sigma-Aldrich | 15506017 | Used for steps of sNuc-Seq |
| Trypan blue stain | Gibco | 15250061 | Used for cell counting |
| Tween20 | ThermoFisher | AAJ20605AP | Used for steps of sNuc-Seq |
| Zeiss STEMI SV 11 Apo stereomicroscope | Zeiss | n/a | Microscope used for dissections |
References
- Bosher, S. K., Warren, R. L. Observations on the electrochemistry of the cochlear endolymph of the rat: a quantitative study of its electrical potential and ionic composition as determined by means of flame spectrophotometry. Proceedings of the Royal Society of London. Series B. Biological Sciences. 171 (1023), 227-247 (1968).
- Patuzzi, R. Ion flow in stria vascularis and the production and regulation of cochlear endolymph and the endolymphatic potential. Hearing Research. 277 (1-2), 4-19 (2011).
- Wangemann, P. K+ cycling and the endocochlear potential. Hearing Research. 165 (1-2), 1-9 (2002).
- Adachi, N., et al. The mechanism underlying maintenance of the endocochlear potential by the K+ transport system in fibrocytes of the inner ear. The Journal of Physiology. 591 (18), 4459-4472 (2013).
- Hibino, H., Nin, F., Tsuzuki, C., Kurachi, Y. How is the highly positive endocochlear potential formed? The specific architecture of the stria vascularis and the roles of the ion-transport apparatus. Pflugers Archiv. 459 (4), 521-533 (2010).
- Lang, F., Vallon, V., Knipper, M., Wangemann, P. Functional significance of channels and transporters expressed in the inner ear and kidney. American Journal of Physiology. Cell Physiology. 293 (4), C1187-C1208 (2007).
- Liu, W., Schrott-Fischer, A., Glueckert, R., Benav, H., Rask-Andersen, H. The human "cochlear battery"-claudin-11 barrier and ion transport proteins in the lateral wall of the cochlea. Frontiers in Molecular Neuroscience. 10, 239 (2017).
- Marcus, D. C., Wu, T., Wangemann, P., Kofuji, P. KCNJ10 (Kir4.1) potassium channel knockout abolishes endocochlear potential. American Journal of Physiology. Cell Physiology. 282 (2), C403-C407 (2002).
- Spicer, S. S., Schulte, B. A. Differentiation of inner ear fibrocytes according to their ion transport related activity. Hearing Research. 56 (1-2), 53-64 (1991).
- Wangemann, P., Liu, J., Marcus, D. C. Ion transport mechanisms responsible for K+ secretion and the transepithelial voltage across marginal cells of stria vascularis in vitro. Hearing Research. 84 (1-2), 19-29 (1995).
- Yoshida, T., et al. The unique ion permeability profile of cochlear fibrocytes and its contribution to establishing their positive resting membrane potential. Pflugers Archiv. 468 (9), 1609-1619 (2016).
- Johns, J. D., Adadey, S. M., Hoa, M. The role of the stria vascularis in neglected otologic disease. Hearing Research. 428, 108682 (2023).
- Kim, J., Ricci, A. J. In vivo real-time imaging reveals megalin as the aminoglycoside gentamicin transporter into cochlea whose inhibition is otoprotective. Proceedings of the National Academy of Sciences. 119 (9), e2117846119 (2022).
- Zdebik, A. A., Wangemann, P., Jentsch, T. J. Potassium ion movement in the inner ear: insights from genetic disease and mouse models. Physiology. 24, 307-316 (2009).
- Chen, J., Zhao, H. B. The role of an inwardly rectifying K+ channel (Kir4.1) in the inner ear and hearing loss. Neuroscience. 265, 137-146 (2014).
- Steel, K. P., Barkway, C. Another role for melanocytes: their importance for normal stria vascularis development in the mammalian inner ear. Development. 107 (3), 453-463 (1989).
- Ito, T., Nishio, A., Wangemann, P., Griffith, A. J. Progressive irreversible hearing loss is caused by stria vascularis degeneration in an Slc26a4-insufficient mouse model of large vestibular aqueduct syndrome. Neuroscience. 310, 188-197 (2015).
- Gu, S., et al. Characterization of rare spindle and root cell transcriptional profiles in the stria vascularis of the adult mouse cochlea. Scientific Reports. 10 (1), 18100 (2020).
- Gow, A., et al. Deafness in claudin 11-null mice reveals the critical contribution of basal cell tight junctions to stria vascularis function. The Journal of Neuroscience. 24 (32), 7051-7062 (2004).
- Chang, Q., et al. Virally mediated Kcnq1 gene replacement therapy in the immature scala media restores hearing in a mouse model of human Jervell and Lange-Nielsen deafness syndrome. EMBO Molecular Medicine. 7 (8), 1077-1086 (2015).
- Faridi, R., et al. Mutational and phenotypic spectra of KCNE1 deficiency in Jervell and Lange-Nielsen Syndrome and Romano-Ward Syndrome. Human Mutation. 40 (2), 162-176 (2019).
- Wangemann, P., et al. Loss of KCNJ10 protein expression abolishes endocochlear potential and causes deafness in Pendred syndrome mouse model. BMC Medicine. 2, 30 (2004).
- Kitajiri, S. -. I., et al. Expression patterns of claudins, tight junction adhesion molecules, in the inner ear. Hearing Research. 187 (1-2), 25-34 (2004).
- Jagger, D. J., Nevill, G., Forge, A. The membrane properties of cochlear root cells are consistent with roles in potassium recirculation and spatial buffering. Journal of the Association for Research in Otolaryngology. 11 (3), 435-448 (2010).
- Ito, T., Kurata, N., Fukunaga, Y. Tissue-resident macrophages in the stria vascularis. Frontiers in Neurology. 13, 818395 (2022).
- Zhang, J., et al. VEGFA165 gene therapy ameliorates blood-labyrinth barrier breakdown and hearing loss. JCI Insight. 6 (8), e143285 (2021).
- Shi, X. Pathophysiology of the cochlear intrastrial fluid-blood barrier (review). Hearing Research. 338, 52-63 (2016).
- Gu, S., et al. Identification of potential Meniere’s disease targets in the adult stria vascularis. Frontiers in Neurology. 12, 630561 (2021).
- Fischer, J., Ayers, T. Single nucleus RNA-sequencing: how it’s done, applications and limitations. Emerging Topics in Life Sciences. 5 (5), 687-690 (2021).
- Korrapati, S., et al. Single cell and single nucleus RNA-Seq reveal cellular heterogeneity and homeostatic regulatory networks in adult mouse stria vascularis. Frontiers in Molecular Neuroscience. 12, 316 (2019).
- Pyle, M. P., Hoa, M. Applications of single-cell sequencing for the field of otolaryngology: A contemporary review. Laryngoscope Investigative Otolaryngology. 5 (3), 404-431 (2020).
- Hwang, B., Lee, J. H., Bang, D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Experimental & Molecular Medicine. 50 (8), 1-14 (2018).
- Shafer, M. E. R. Cross-species analysis of single-cell transcriptomic data. Frontiers in Cell and Developmental Biology. 7, 175 (2019).
- Chen, G., Ning, B., Shi, T. Single-cell RNA-Seq technologies and related computational data analysis. Frontiers in Genetics. 10, 317 (2019).
- Longo, S. K., Guo, M. G., Ji, A. L., Khavari, P. A. Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics. Nature Reviews Genetics. 22 (10), 627-644 (2021).
- Kim, N., Kang, H., Jo, A., Yoo, S. A., Lee, H. O. Perspectives on single-nucleus RNA sequencing in different cell types and tissues. Journal of Pathology and Translational Medicine. 57 (1), 52-59 (2023).
- Grindberg, R. V., et al. RNA-sequencing from single nuclei. Proceedings of the National Academy of Sciences. 110 (49), 19802-19807 (2013).
- Montgomery, S. C., Cox, B. C. Whole mount dissection and immunofluorescence of the adult mouse cochlea. Journal of Visuazlied Experiments. (107), e53561 (2016).

