An Efficient and Fast Method for Labeling and Analyzing Mouse Glomeruli
Summary
This study presents an easy-to-use, complete, and simple set of methods to label and analyze glomeruli from CUBIC-cleared mouse kidneys. Data such as glomerulus number and volume can be obtained easily and reliably using fluorescein isothiocyanate (FITC)-Dextran, light sheet fluorescence microscopy (LSFM), or common confocal microscopy and software such as Imaris.
Abstract
The glomeruli are fundamental units in the kidney; hence, studying the glomeruli is pivotal for understanding renal function and pathology. Biological imaging provides intuitive information; thus, it is of great significance to label and observe the glomeruli. However, the glomeruli observation methods currently in use require complicated operations, and the results may lose label details or three-dimensional (3D) information. The clear, unobstructed brain imaging cocktails and computational analysis (CUBIC) tissue clearing technology has been widely used in renal research, allowing for more accurate detection and deeper detection depth. We found that mouse glomeruli can be rapidly and effectively labeled by tail vein injection of medium molecular weight FITC-Dextran followed by the CUBIC clearing method. The cleared mouse kidney could be scanned by a light-sheet microscope (or a confocal microscope when sliced) to obtain three-dimensional image stacks of all the glomeruli in the entire kidney. Processed with appropriate software, the glomeruli signals could be easily digitized and further analyzed to measure the number, volume, and frequency of the glomeruli.
Introduction
The number and volume of glomeruli are very important for the diagnosis and treatment of various kidney diseases1,2,3,4,5. The golden standard of glomeruli number estimation is the physical dissector/fractionator combination. However, this method requires special reagents and equipment, making it slow and expensive6,7,8,9. Biopsy provides a wealth of information, but obviously, this method is only suitable for rough estimations10,11. Medical imaging technologies, including magnetic resonance imaging (MRI), computed tomography (CT), and X-ray, are also widely used in glomerular detection12,13,14,15, but such technologies require bulky instruments. New methods, such as matrix-assisted laser desorption/ionization (MALDI) imaging mass spectrometer16 or the thick and thin section method17, have also been used in glomerular detection, though they remain tedious and laborious.
With the help of transparency technologies, it is possible to observe deeper depths and obtain richer and more complete information from thick tissues or even whole organs18,19,20,21,22,23. Therefore, transparency technologies have been widely used in kidney research24. The observation and detection of glomeruli in the cleared kidneys are also involved. However, these published articles either only briefly referred to glomerular detection25 or used difficult-to-achieve labeling methods such as transgenic animals26, self-produced dyes13, or high-concentration antibody incubation27 to label the glomeruli. In addition, although studies had analyzed glomeruli in cleared kidneys, the analyses were always limited13 or relied on analysis algorithms established by the authors themselves26.
We have previously demonstrated a more convenient way to label the glomeruli in mice kidneys28. By using Imaris, we found that glomeruli count, frequency, and volume could be quickly obtained. Thus, here we present a more accessible, comprehensive, and simplified set of methods to label and analyze the glomeruli of mice kidneys.
Protocol
Adult C57BL/6 mice (6 weeks of age, 25-30 g) were used in this study. All procedures were performed in compliance with local regulations of animal welfare and experimental ethics. The study was approved by the West China Hospital of Sichuan University Biomedical Research Ethics Committee.
1. Glomeruli labeling and tissue preparation
- Glomeruli labeling
- Dissolve FITC-dextran (10 mg) in 1x phosphate-buffered saline (PBS) at a ratio of 1:1 (1 mg: 1 mL) to prepare the working probe solution.
NOTE: The working solution can be stored at 4 °C for 1 month. - Place the C57 mouse in a mouse tail vein fixator. Wet the gauze with hot water, wrap the gauze around the middle part of the mouse tail, and warm the tail for 1 min.
- Wipe the mouse tail with 75% ethanol until the left and right tail veins are visible.
- Draw 100 µL of the working probe solution with a 1 mL insulin syringe and slowly inject the solution into the mouse tail vein.
- After injection, let the probe solution circulate for 30 min. Allow the mice to move freely in their cages.
- Once sufficient circulation is achieved, deeply anesthetize the miceby intraperitoneal injection of sodium pentobarbital (1%, 60-80 µg/kg).
- Dissolve FITC-dextran (10 mg) in 1x phosphate-buffered saline (PBS) at a ratio of 1:1 (1 mg: 1 mL) to prepare the working probe solution.
- Kidney sampling and fixation
- Fix the anesthetized mice to the anatomical plate in a supine position. Secure their paws with medical tape.
- After skin preparation, euthanize the mice with a lethal dose of sodium pentobarbital (1%, 10 mg/kg).
- Make a midline abdominal incision to expose the abdominal cavity.
- Reveal the kidneys and remove them with a scalpel and scissors in the fume hood. Remove the renal envelope gently.
- Collect the kidneys and fix them in 4% paraformaldehyde (PFA) at 4 °C overnight.
- Wash the samples with PBS three times 1 h each with shaking at room temperature (RT).
- Prepare the kidney for microscopy.
- Slice the kidney for confocal microscopy: Cut the kidney into 1 mm thick slices using the brain matrix (1 mm steel, 40-75 g) to standardize the slice thickness. Transfer the slices into 4% PFA and continue the fixation at 4 °C overnight.
- Whole kidney for light-sheet microscopy: Fix the kidneys in 4% PFA at 4 °C for 48 h.
2. Clearing
- Reagent preparation
- Prepare CUBIC-L reagent by mixing 10% (wt/wt) N-butyldiethanolamine, 10% (wt/wt) Triton X-100 and 80% ddH2O.
- Prepare CUBIC-R reagent by mixing 45% (wt/wt) antipyrine, 30% (wt/wt) nicotinamide, 0.5% (vol/vol) N-butyldiethanolamine and 24.5% ddH2O. Ensure the solution has a pH of 9.6-9.8 and a refractive index (RI) of 1.522.
NOTE: CUBIC-L is used for delipidation, and CUBIC-R is used for refractive index matching.
- Clearing procedure
- Collect the kidney samples in a 50 mL centrifuge tube and then clear them in CUBIC-L with shaking at 60 rpm at 37 °C. Replace CUBIC-L every 4 h (for kidney slices) or every 24 h (for a whole kidney) until achieving satisfactory optical transparency.
NOTE: This process takes 1 day for kidney slices and 5 days for a whole kidney. - Wash the samples three times in PBS with gentle shaking at RT for 1 h each.
- Apply CUBIC-R to the samples, shaking samples at 37 °C, 60 rpm for 6 h.
- Observe the cleared samples directly or store them in black tubes filled with CUBIC-R at RT for 1 week.
- Collect the kidney samples in a 50 mL centrifuge tube and then clear them in CUBIC-L with shaking at 60 rpm at 37 °C. Replace CUBIC-L every 4 h (for kidney slices) or every 24 h (for a whole kidney) until achieving satisfactory optical transparency.
3. Image acquisition
- Confocal imaging
NOTE: Image the sliced kidney samples with confocal microscopy (Lenses: A Plan Apo λ 4×/0.2 N1 objective or a Plan Apo VC 10×/0.45 DIC N1 objective). Capture large images (e.g., 6052 µm x 6052 µm for the kidney slice) with z-stacks.- Turn on the confocal microscope in the following sequence: laser power, confocal control switch, computer, microscope, and confocal hardware. After running the self-test, turn on the confocal software.
- Select the appropriate lens. Remove the sample from the CUBIC-R reagent, place it in the confocal dish, and cover it to prevent the CUBIC-R reagent from drying out.
- Move the sample to the middle of the field of view and adjust the focal plane until it is in focus.
- Apply 3D reconstruction (step 3.1.4.1) and large image reconstruction (steps 3.1.4.2-3.1.4.3).
- 3D reconstruction (confocal): Switch the zoom plane in one direction until no fluorescence is visible; this position is defined as the top. Then, switch the zoom plane in the opposite direction until no fluorescence is visible; this position is defined as the bottom. Click the Run button in the lower-right corner of the dialog box to start the scanning.
- Large image reconstruction (confocal): Move the field of view to the middle of the sample, and set the current position as the center. Select a 2 x 2 field of view.
- In the ND multifunction interface, select the Z stack to reconstruct large images and set different options in the panel. Then press the Run butto n to start the scanning.
- Light sheet fluorescence microscopy (LSFM) imaging
NOTE: Image the whole kidneys with a light sheet fluorescence microscope (Lens: EC Plan-Neofluar 5x/0.16 (Medium n = 1.529) objective).- Sample attaching (LSFM):
- Glue the transparent kidney to the sample fixing adapter and wait until the kidney is firmly attached.
- Soak the kidney in the sample bin and push the sample bin into the microscope system. Close the hatch.
- Observe and scan (LSFM):
- In the Locate interface, move the sample to a suitable observation position. Switch to the Acquisition interface, set the optical path, select 488 nm laser, LBF 405/488/561/640, optical splitting filter block SBS LP 490, select Camera 2, and change the pseudo-color to green.
- Fill in the Left/Right Z Offset value recorded earlier in the Channel submenu. Select the smallest zoom (0.36) and fine-tune it for maximum clarity.
- Find the XY boundary and Z range of the entire organ. Adjust the laser intensity and exposure time (preferably less than 10 ms), and click Run to start.
- If the tissue is too big, merge two or more images with image stitching. Open the captured file using the microscopy software (here, ZEISS ZEN 3.7). Select Processing > Method > Stitching > Method Parameter > Apply to complete image stitching.
- Sample attaching (LSFM):
4. Data processing and quantification
NOTE: Process the image stacks with Imaris (image analysis) software, using the Surface function to label the glomeruli and perform analysis.
- Count of glomeruli
- Import large images (confocal microscopy) or stitched images (light-sheet microscopy) into the image analysis software.
- Open the files in the image analysis software. Click the Surface button to create a new Surface. Follow the guide at the bottom left part of the screen.
- Click the Next button (right arrow button) without ticking anything.
- Tick the box before the Smooth option and enter a proper number (e.g., 14.5) to control the detail of the Surface. Select Background Subtraction in Thresholding and fill the box with the approximate average diameter of the glomeruli. Click the Next button.
- Adjust if needed, and click the Next button. Select the proper range and click the Next button to create a Surface.
- Click Filter, then Add button to add Sphericity to filter out non-spherical objects. Click the Duplicate button to copy this new surface.
- Click the Statistics button, then click the Overall button; the software will present the number of glomeruli as the Total Number of Surface.
- Volume of glomeruli
- Create a surface of the glomeruli as above.
- When the Surface is completed, click the Selection button to select the target objects and present the volume of every object. Click the Save button, and export statistics to a spreadsheet for further analysis.
- Volume of the slice or whole kidney
- Import large images (confocal microscopy) or stitched images (light-sheet microscopy) into the image analysis software.
- Open files in the image analysis software. Click the Surface button to create a new Surface. Follow the guide at the bottom left part of the screen.
- Click the Next button (right arrow button) without ticking.
- Tick the box before the Smooth option and enter a proper number (e.g., 50) to control the detail of the Surface. Select Absolute Intensity in Thresholding. Click the Next button.
- Adjust the Surface until it covers the whole tissue, and click the Next button. Click the Next button to create the Surface.
- Click the Selection button to select the Surface of the whole tissue and present the volume of the tissue. Click the Save button to export statistics to a spreadsheet.
Representative Results
This study provides a simple and efficient method for labeling and analyzing the glomeruli in mice kidneys.
Glomeruli (blood vessels) can be well labeled by intravascularly injected FITC-Dextran. After the clearing process, the kidney became transparent (Figure 1A), and the glomeruli could be clearly observed by using light-sheet microscopy (Figure 1B) or confocal microscopy (Figure 1C). Confocal microscopy has a limited scanning depth, so kidneys should be cut into approximately 1 mm-thick slices. If a light-sheet microscope is used, the whole kidney can be scanned directly.
With the clear signals, it was easy to count the number of glomeruli. In fact, the labeling was so effective that the glomeruli could be counted with the naked eye. The volume of glomeruli could also be directly measured. Of course, software such as Imaris greatly accelerated this process. Using the Surface function, all the glomeruli in a kidney slice (Figure 2), in an entire kidney (Figure 3), or in a strip (Figure 4) could be selected, and the number and volume of the glomeruli could be obtained directly (Figure 2C, Figure 3C, Figure 4C). The volume of the selected region could also be measured (Figure 2B, Figure 3B, Figure 4B), so the glomeruli volume ratio and frequency in a certain region or in the whole kidney could be calculated (Figure 2D, Figure 3D, Figure 4D). An area of the kidney could be selected to perform calculations in specific regions. We presented a strip similar to a biopsy. The number, volume, and frequency of the glomeruli in the strip could also be obtained easily (Figure 4).
As shown in the figures, the results presented in this study are (N = number, F = frequency, V = volume, V(a) = average volume, V(t) = total volume):
Nglomeruli in slice = 1128, Nglomeruli in whole kidney = 14006, Fglomerular in slice = 65 per mm3, Fglomerular in whole kidney = 105 per mm3, Vtotal glomeruli in slice/Vslice = 2.24%, Vtotal glomeruli in whole kidney/Vkidney = 5.54%, V(a)glomeruli in slice = 373654 µm3, V(a)glomeruli in whole kidney = 521627 µm3, V(t)glomeruli in slice = 421481724 µm3, V(t)glomeruli in whole kidney = 7305386256 µm3 (Figure 2D, Figure 3D).
Nglomeruli in strip = 63, Fglomeruli in strip = 72 per mm3, Vtotal glomeruli in strip/Vstrip = 2.23%, V(a)glomeruli in strip = 307698 µm3, V(t)glomeruli in strip = 19384977 µm3 (Figure 4D).
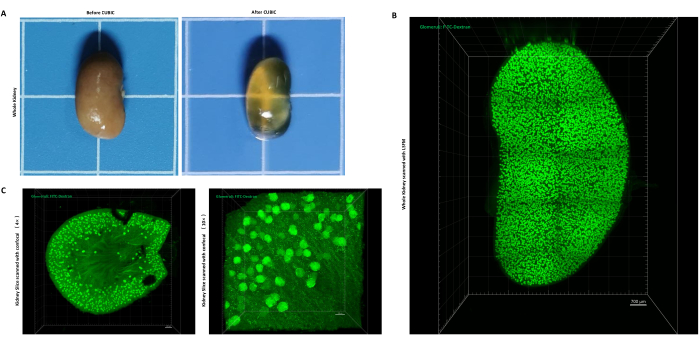
Figure 1: Kidney tissue transparency and FITC-dextran-labeled glomeruli. (A) Kidney before and after transparency treatment. (B) The image of a whole kidney scanned with LSFM. (C) The image of a slice of kidney scanned with the confocal microscope (left: 4x, right: 10x). scale bar = 300 µm (4x, confocal); scale bar = 100 µm (10x, confocal).; scale bar = 700 µm (5x, zoom 0.36, LSFM). Glomeruli were labeled with FITC-Dextran. Please click here to view a larger version of this figure.
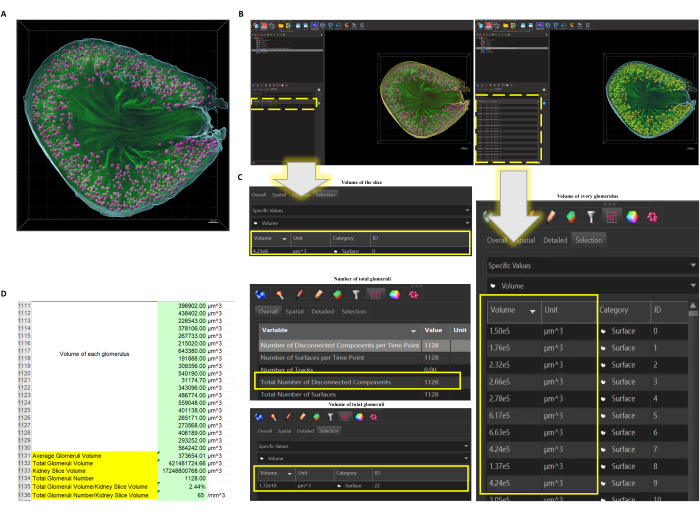
Figure 2: The function Surface applied to the confocal scanned kidney slice. (A) Surface of the slice and all glomeruli are created. Pink = glomeruli, blue = out Surface of the kidney slice, green = original label of vessels (glomeruli and some big vessels). Scale bar = 300 µm. (B) When the Surface of the slice (left) or the glomeruli (right) are selected (selected objects would turn yellow), data could be obtained directly (dotted box highlights where it shows the data). (C) The number, the volume of every single glomerulus, and the volume of total glomeruli could be directly obtained as well as the volume of the slice. (D) Exported data could be further analyzed so the calculation, such as the average volume of the glomeruli and volume ratio of glomeruli and selected area, could be worked out. Please click here to view a larger version of this figure.
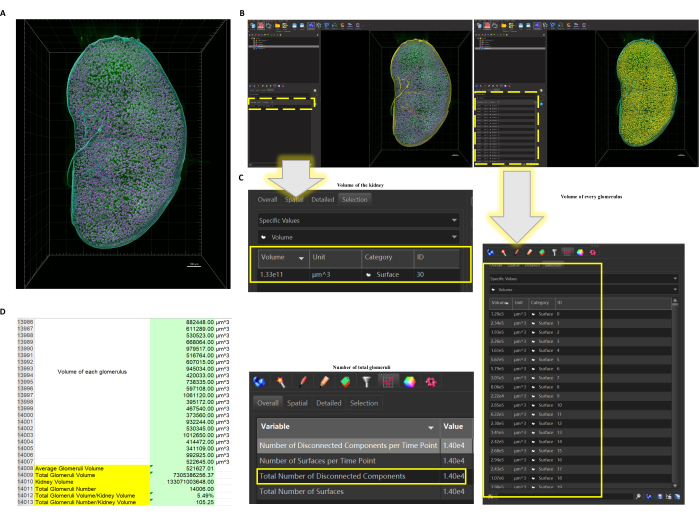
Figure 3: The function Surface applied to the LSFM scanned whole kidney. (A) Surface of the whole kidney and all glomeruli are created. Pink = glomeruli, blue = out Surface of the whole kidney, green = original label of vessels (glomeruli and some big vessels). Scale bar =1000 µm. (B) When the Surface of the kidney (left) or the glomeruli (right) are selected (selected objects would turn yellow), data could be obtained directly (dotted box highlights where it shows the data). (C) The number, the volume of every single glomerulus, and the number of the glomeruli could be directly obtained, as well as the volume of the kidney. (D) Exported data could be further analyzed so the calculation, such as the average volume of glomeruli and volume ratio of glomeruli and the whole kidney, could be worked out. Please click here to view a larger version of this figure.
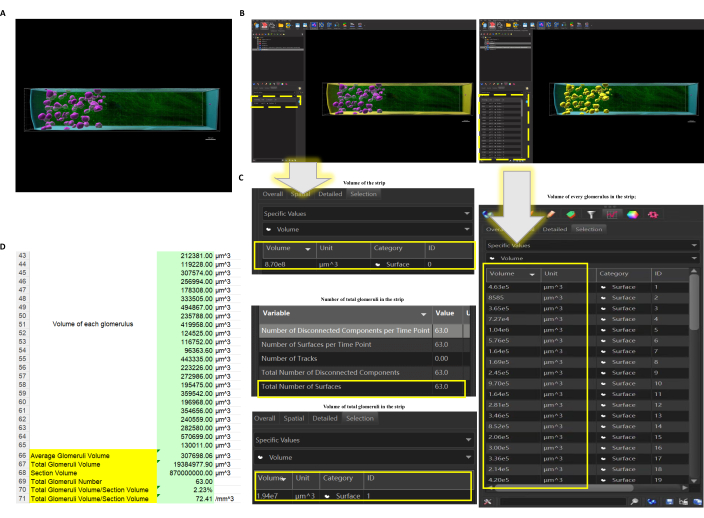
Figure 4: The function Surface applied to the selected kidney strip. (A) Surface of the strip and all glomeruli are created. Pink = glomeruli, blue = out Surface of the kidney strip, green = original label of vessels (glomeruli and some big vessels). Scale bar =150 µm. (B) When the Surface of the strip (left) or the glomeruli (right) are selected (selected objects would turn yellow), data could be obtained directly (dotted box highlights where it displays the data). (C) The number, the volume of every single glomerulus, and the volume of total glomeruli could be directly obtained, as well as the volume of the strip. (D) Exported data could be further analyzed so the calculation, such as the average volume of the glomeruli and volume ratio of glomeruli and selected area, could be worked out. Please click here to view a larger version of this figure.
Discussion
Tissue-clearing technologies can be classified into 3 or 4 groups29,30,31. Organic solvent-based tissue clearing (e.g., DISCO and PEGASOS), aqueous-based tissue clearing (e.g., CUBIC), and hydrogel embedding tissue clearing (e.g., CLARITY) have all been applied in kidney clearing 25,26,28,32. CUBIC, as we have demonstrated, works best when the glomeruli (vessels) are labeled with FITC-Dextran. In addition, CUBIC has a relatively more convenient operation process and requires no more than ordinary microscopic lenses27, as the processed samples do not need to be soaked in organic reagents33. However, different clearing methods and vessel label methods have their own advantages and disadvantages; researchers may need to experiment with different approaches to meet their specific needs.
Previous studies noticed that the glomeruli could be labeled in cleared kidneys when blood vessels in the kidneys were labeled13,25,26,28. A thorough explanation for this phenomenon still needs to be further investigated, but it can be deduced that a good labeling method for the vascular system also effectively labels the glomeruli. Based on the experimental results, we would recommend the intravascular injection of FITC-Dextran with a medium molecular weight (such as 150 kDa). However, since labeling relies on blood circulation, this method may fail to label the glomeruli when they lack circulation.
LSFM permits large-scale tissue observation, making it more suitable for whole kidney glomeruli detection. However, light-sheet microscopes may not be available in every laboratory. So, we are providing an alternative set of protocols using ordinary confocal microscopes. Although the kidney needs to be sliced into multiple sections, obtaining data such as the counts and volume of glomeruli in the whole kidney is feasible. Besides, confocal microscopy provides a smaller volume of data, which is more data processing friendly.
Imaris provides direct data output. However, care should be taken to match the analog signal to the original closely. Additionally, the software may take a long time to process large files, which can be frustrating. As long as the glomeruli can be clearly labeled and visualized, there is more than one unique solution for data analysis. Researchers from different laboratories could work with their own familiar software and programming languages.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This study was supported by grants from the National Natural Science Foundation of China (82204951) and Sichuan Science and Technology Program (2020JDRC0102).
Materials
| 4% PFA | Biosharp | 7007171800 | Fixation reaagen |
| 502 Glue | Deli | 7146 | For fixing the kidney to the sample fixing adapter |
| Antipyrine | Aladdin | A110660 | Clearing reagent |
| Brain Matrix | RWD Life Science | 1mm 40-75 | Tissue slicing |
| Confocal microscopy | Nikon | A1plus | Image acquisition |
| FITC-Dextran | Sigma-Aldrich | FD150S | Labeling reagent |
| Light sheet fluorescence microscopy | Zeiss | Light sheet 7 | Image acquisition |
| Mice | Ensiweier | Adult C57BL/6 mice (6 weeks of age, 25–30 g) | |
| N-Butyldiethanolamine | Aladdin | B299095 | Clearing reagent |
| Nicotinamide | Aladdin | N105042 | Clearing reagent |
| Pentobarbital Natriumsalz | Sigma-Aldrich | P3761 | |
| Tail vein fixator | JINUOTAI | JNT-FS35 | Fix the mouse for vail injection |
| Triton X-100 | Sigma-Aldrich | T8787 | Clearing reagent |
References
- Hoy, W. E., et al. Nephron number, glomerular volume, renal disease and hypertension. Current Opinion in Nephrology and Hypertension. 17 (3), 258-265 (2008).
- Bertram, J. F., Douglas-Denton, R. N., Diouf, B., Hughson, M. D., Hoy, W. E. Human nephron number: implications for health and disease. Pediatric Nephrology. 26 (9), 1529-1533 (2011).
- Nyengaard, J. R., Bendtsen, T. F. Glomerular number and size in relation to age, kidney weight, and body surface in normal man. The Anatomical Record. 232 (2), 194-201 (1992).
- Rasch, R. Prevention of diabetic glomerulopathy in streptozotocin diabetic rats by insulin treatment. Kidney size and glomerular volume. Diabetologia. 16 (2), 125-128 (1979).
- Puelles, V. G., et al. Glomerular number and size variability and risk for kidney disease. Current Opinion in Nephrology and Hypertension. 20 (1), 7-15 (2011).
- Bertram, J. F., et al. Why and how we determine nephron number. Pediatric Nephrology. 29, 575-580 (2014).
- Bertram, J. F., Soosaipillai, M. C., Ricardo, S. D., Ryan, G. B. Total numbers of glomeruli and individual glomerular cell types in the normal rat kidney. Cell and Tissue Research. 270 (1), 37-45 (1992).
- Nyengaard, J. R. Stereologic methods and their application in kidney research. Journal of the American Society of Nephrology. 10 (5), 1100-1123 (1999).
- Bertram, J. F. Analyzing renal glomeruli with the new stereology. International Review of Cytology. 161, 111-172 (1995).
- Lødrup, A. B., Karstoft, K., Dissing, T. H., Pedersen, M., Nyengaard, J. R. Kidney biopsies can be used for estimations of glomerular number and volume: a pig study. Virchows Archiv. 452 (4), 393-403 (2008).
- Lane, P. H., Steffes, M. W., Mauer, S. M. Estimation of glomerular volume: a comparison of four methods. Kidney International. 41 (4), 1085-1089 (1992).
- Baldelomar, E. J., Charlton, J. R., deRonde, K. A., Bennett, K. M. In vivo measurements of kidney glomerular number and size in healthy and Os(/+) mice using MRI. American Journal of Physiology-Renal Physiology. 317 (4), F865-F873 (2019).
- Huang, J., et al. A cationic near infrared fluorescent agent and ethyl-cinnamate tissue clearing protocol for vascular staining and imaging. Scientific Reports. 9 (1), 521 (2019).
- Beeman, S. C., et al. Measuring glomerular number and size in perfused kidneys using MRI. American Journal of Physiology-Renal Physiology. 300 (6), F1454-F1457 (2011).
- Basgen, J. M., Steffes, M. W., Stillman, A. E., Mauer, S. M. Estimating glomerular number in situ using magnetic resonance imaging and biopsy. Kidney International. 45 (6), 1668-1672 (1994).
- Prentice, B. M., Caprioli, R. M., Vuiblet, V. Label-free molecular imaging of the kidney. Kidney International. 92 (3), 580-598 (2017).
- Sanden, S. K., Wiggins, J. E., Goyal, M., Riggs, L. K., Wiggins, R. C. Evaluation of a thick and thin section method for estimation of podocyte number, glomerular volume, and glomerular volume per podocyte in rat kidney with Wilms’ tumor-1 protein used as a podocyte nuclear marker. Journal of the American Society of Nephrology. 14 (10), 2484-2493 (2003).
- Hama, H., et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nature Neuroscience. 14 (11), 1481-1488 (2011).
- Susaki, E. A., et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell. 157 (3), 726-739 (2014).
- Lloyd-Lewis, B., et al. Imaging the mammary gland and mammary tumours in 3D: optical tissue clearing and immunofluorescence methods. Breast Cancer Research. 18 (1), 127 (2016).
- Ren, Z., et al. CUBIC-plus: An optimized method for rapid tissue clearing and decolorization. Biochemical and Biophysical Research Communications. 568, 116-123 (2021).
- Azaripour, A., et al. A survey of clearing techniques for 3D imaging of tissues with special reference to connective tissue. Progress in Histochemistry and Cytochemistry. 51 (2), 9-23 (2016).
- Matsumoto, K., et al. Advanced CUBIC tissue clearing for whole-organ cell profiling. Nature Protocols. 14 (12), 3506-3537 (2019).
- Puelles, V. G., Moeller, M. J., Bertram, J. F. We can see clearly now: optical clearing and kidney morphometrics. Current Opinion in Nephrology and Hypertension. 26 (3), 179-186 (2017).
- Zhu, J., et al. Optimal combinations of fluorescent vessel labeling and tissue clearing methods for three-dimensional visualization of vasculature. Neurophotonics. 9 (4), 045008 (2022).
- Klingberg, A., et al. Fully automated evaluation of total glomerular number and capillary tuft size in nephritic kidneys using lightsheet microscopy. Journal of the American Society of Nephrology. 28 (2), 452-459 (2017).
- Renier, N., et al. iDISCO: A simple, rapid method to immunolabel large tissue samples for volume imaging. Cell. 159 (4), 896-910 (2014).
- Bai, L., et al. A simple and effective vascular network labeling method for transparent tissues of mice. Journal of Biophotonics. 16 (7), e202300042 (2023).
- Richardson, D. S., Lichtman, J. W. Clarifying tissue clearing. Cell. 162 (2), 246-257 (2015).
- Kolesová, H., Olejníčková, V., Kvasilová, A., Gregorovičová, M., Sedmera, D. J. I. Tissue clearing and imaging methods for cardiovascular development. Iscience. 238 (2), 489-507 (2021).
- Tian, T., Yang, Z., Li, X. Tissue clearing technique: Recent progress and biomedical applications. Journal of Anatomy. 238 (2), 489-507 (2021).
- Du, H., Hou, P., Zhang, W., Li, Q. Advances in CLARITY-based tissue clearing and imaging. Experimental and Therapeutic. 16 (3), 1567-1576 (2018).
- Ertürk, A., Lafkas, D., Chalouni, C. Imaging cleared intact biological systems at a cellular level by 3DISCO. Journal of Visualized Experiments: JoVE. 89, e51382 (2014).

