Methods for the Extraction of Endosymbionts from the Whitefly Bemisia tabaci
Summary
Here, we present a protocol to isolate endosymbionts from the whitefly Bemisia tabaci through dissection and filtration. After amplification, the DNA samples are suitable for subsequent sequencing and study of the mutualism between endosymbionts and whitefly.
Abstract
Bacterial symbionts form an intimate relationship with their hosts and confer advantages to the hosts in most cases. Genomic information is critical to study the functions and evolution of bacterial symbionts in their host. As most symbionts cannot be cultured in vitro, methods to isolate an adequate quantity of bacteria for genome sequencing are very important. In the whitefly Bemisia tabaci, a number of endosymbionts have been identified and are predicted to be of importance in the development and reproduction of the pests through multiple approaches. However, the mechanism underpinning the associations remains largely unknown. The obstacle partially comes from the fact that the endosymbionts in whitefly, mostly restrained in bacteriocytes, are hard to separate from the host cells. Here we report a step-by-step protocol for the identification, extraction and purification of endosymbionts from the whitefly B. tabaci mainly by dissection and filtration. Endosymbiont samples prepared by this method, although still a mixture of different endosymbiont species, are suitable for subsequent genome sequencing and analysis of the possible roles of endosymbionts in B. tabaci. This method may also be used to isolate endosymbionts from other insects.
Introduction
Bacteria forming an intimate symbiotic relationship with relative hosts are widespread in arthropods1. The endosymbionts have been demonstrated to affect aspects of hosts, such as nutrition metabolism, reproduction, responses to environmental stresses2,3,4 etc., in almost every developmental stage5. However, the mechanism underpinning the associations still remains largely unknown. Genomics is of priority and importance when studying the potential functions and roles of bacteria. Some fundamental information, i.e. the taxonomic status, functional genes, metabolism pathways, secretion systems, can be inferred from genome sequences, which sheds lights on the potential roles of symbionts in symbiosis. With the development of high-throughput sequencing, a vast number of bacterial genomes have been sequenced with diverse functions revealed6.
Endosymbionts are of vital importance in hemipterans, such as aphids7, bedbugs8, psyllids9, brown planthoppers10 and cicadas11. For instance, Buchnera in aphids, as the obligate symbiont, has been demonstrated to be involved in essential amino acids biosynthesis, along with the genes from aphid genome12. Furthermore, transcriptional regulation of Buchnera is also revealed13. In psyllids, Carsonella is sequenced and ranked the smallest bacterial genome ever found14. All these hallmarks of endosymbionts are based and inferred from the genome sequences. Because these endosymbionts cannot be cultured in vitro, several approaches have been applied to isolate adequate bacteria for sequencing. In aphids, endosymbionts are extracted through centrifugation and filtration, and subjected to further genomic and transcriptomic analysis5. In brown planthoppers, endosymbionts are sequenced along with the whole insect genome10.
Whitefly B. tabaci is a species complex containing more than 35 morphologically indistinguishable species (cryptic species), among which, two invasive species have invaded all over the world and caused tremendous harm to agricultural production15. Of note, endosymbionts within the B. tabaci species have shown importance in the development of the pests16. To date, eight endosymbionts have been identified in the whitefly, including the obligate symbiont, Candidatus Portiera aleyrodidarum, and seven secondary symbionts Hamiltonella, Rickettsia, Arsenophonus, Cardinium, Wolbachia, Fritschea and Hemipteriphilus defined17,18.
Unlike the hemipterans described previously, the whitefly B. tabaci is an extremely tiny insect only 1 mm in length. Most endosymbionts are confined to bacteriocytes19 (specialized cells containing symbionts, which further form bacteriome in B. tabaci). In addition, these endosymbionts cannot be cultured in vitro. The only way to obtain endosymbionts from B. tabaci is to dissect the bacteriome out. However, there is difficulty in the dissection. First, the fragile bacteriome always links with other tissues of the whitefly, which is hard to separate. Secondly, the tiny size of the whitefly limits the isolation of enough bacteriome. Thirdly, endosymbionts cluster in the bacteriome, making it extremely complicated to acquire a single species of bacterium.
Here, we report a simple and inexpensive protocol to isolate whitefly endosymbionts for subsequent metagenome sequencing. Through dissection, purification and amplification, adequate endosymbiont DNA could be obtained and the species of bacteria could be confirmed. The described protocol can be used similarly in other arthropods.
Protocol
1. Whitefly Rearing and Cryptic Species Identification
- Maintain the whitefly species on cotton Gossypium hirsutum (Malvaceae) (cv. Zhe-Mian 1973) in cages under standard conditions of 27 ± 1 °C, 70 ± 10% humidity and 14 h light: 10 h dark regime.
- Collect an individual adult whitefly and homogenize in 30 µL of lysis buffer (10 mM Tris, pH 8.4, 50 mM KCl, 0.45% [wt/vol] Tween-20, 0.2% [wt/vol] gelatin, 0.45% [vol/vol] Nonidet P 40, 60 g/mL Proteinase K).
- Incubate the homogenate at 65 °C for 1 h and then 100 °C for 10 min.
NOTE: The incubation time at 65 °C can be increased if necessary. The incubation time at 100 °C should be critically controlled to sufficiently inactivate Proteinase K and avoid DNA damage. - Perform polymerase chain reaction (PCR) using whitefly DNA (acquired in section 1.3) based on mitochondrial cytochrome oxidase I primers.
COI-F: 5'-TTGATTTTTTGGTCATCCAGAAGT-3'
COI-R: 5'-TAATATGGCAGATTAGTGCATTGGA-3'- Perform the PCR reactions in a final volume of 25 µL containing 1 U of Taq DNA polymerase, 2.5 µL 10x Buffer, 0.2 mM dNTP, 0.2 mM of each primer and 2 µL of whitefly DNA.
- Use the following PCR procedure conditions: initial denaturation 95 °C for 3 min followed by 35 cycles of 95 °C for 45 s (denaturation), 50 °C for 45 s (annealing) and 72 °C for 1 min (extension) and another 10 min at 72 °C for further extension.
- Clean the PCR amplified product using a DNA gel extraction kit with the recommended protocol.
- Sequence the purified DNA samples with a DNA sample sequencing system following the manufacturer's instruction.
- Analyze the sequences using the Basic Local Alignment Search Tool (BLAST) to identify the cryptic species of whitefly and avoid contamination of other species.
2. Endosymbiont Identification and Localization
- Perform PCR on the whitefly DNA (acquired in Step 1.3) to amplify the specific genes of each endosymbiont within the whitefly (the PCR recipe is described in section 1.4, and PCR procedures and primers are listed in Table 1).
- Subject the amplified sample to agarose gel electrophoresis according to previously published protocol20 (1% agarose gel, Tris-acetate-EDTA as running buffer, voltage 5 V/cm) to identify the specific gene of each bacterium.
- Recover and sequence each band to determine the bacterial species within whitefly (see sections 1.5-1.7).
- Perform fluorescence in situ hybridizations (FISH) on whiteflies to identify the location of endosymbionts according to previously published protocol19.
NOTE: Increase the concentration of the fluorescent probes if necessary.
3. Transmission Electron Microscopy (TEM)
- Immerse and fix whiteflies in phosphate buffer (0.1 M, pH 7.0) containing 2.5% [vol/vol] glutaraldehyde for 4 h.
- Wash the samples in phosphate buffer (0.1 M, pH 7.0) three times, 15 min each time.
- Immerse and fix the samples again in phosphate buffer (0.1 M, pH 7.0) containing 1% [wt/vol] OsO4 for 2 h.
- Dehydrate the samples by a graded series of ethanol (30%, 50%, 70%, 80%, 90%, 95% and 100%), 15 min each time.
- Transfer and immerse the samples into 100% acetone for 20 min.
- Place the samples in mixture of 100% acetone and 100% Spurr resin (1:1) for 1 h at room temperature.
- Transfer the samples to mixture of 100% acetone and 100% Spurr resin (1:3) for 3 h at room temperature.
- Transfer and immerse the samples into 100% Spurr resin (incubated at 70 °C) for more than 9 h.
- Section the whitefly samples by using a microtome.
- Stain the ultra-thin films (acquired in section 3.9) by immersion in absolute uranyl acetate for 5-10 min, followed by immersion in 50% alkaline lead citrate for another 5-10 min.
NOTE: The volumes of reagents utilized in section 3.1-3.10 depend on the samples. Make sure the samples are immersed in the reagents. - Observe samples under transmission electron microscopy (15,000X) to determine the localization, shape and size of bacteria for further use (see section 4.6).
4. Whitefly Bacteriome Dissection and Purification
- Add 100 µL of 1x PBS solution onto a microscope slide. Pick some 3rd or 4th instar nymphs from cotton leaves and immerse them into the PBS solution.
- Use fine entomological needles under a stereomicroscope and pull the bacteriomes out from the whitefly body.
- Gently cut a hole on one side of the nymph, and slightly press the other side to let the bacteriomes out.
- Mount a 20 µL microloader (with the leading end cut to meet the size of bacteriome) on a 0.5-10 µL pipette. Extract the individual bacteriome out of the PBS solution into the microloader.
- Wash the bacteriome to eliminate the contamination of other whitefly tissues by pipetting the bacteriome into the PBS solution and extracting the bacteriome. Repeat three times.
- Immediately pipette the washed bacteriome into a centrifuge tube containing 60 µL of 1x PBS solution.
- Syringe-filter the assembled bacteriome through a 5 µm filter membrane (determined by sizes of bacteria and nucleus of bacteriocyte in whitefly). Repeat multiple times to move the mixed liquid through the filter membrane thoroughly.
NOTE: To achieve a better result of amplification and sequencing, assemble over 100 bacteriomes. Wet the filter membrane first by using 1x PBS solution to decrease the loss of bacteria binding to the membrane.
5. Amplification of Endosymbiont Genomes
- Amplify the filtrate (mainly containing endosymbionts of the whitefly) using a DNA amplification kit by the recommended protocol with some modification.
- Choose the recommended protocol of amplification of genomic DNA from blood or cells and prepare buffer D2 for subsequent experiment.
- Directly add 1.5 µL of filtrate (see section 4.6) to the centrifuge tube, followed by 1.5 µL of the Buffer D2 and mix well.
- Incubate on ice for 10 min and add 1.5 µL of the stop solution.
- Add 15 µL of the reaction buffer and 1 µL of the DNA polymerase and mix gently.
- Incubate the mixture at 30 °C for 16 h, followed by 3 min at 65 °C.
- Perform PCR directly on the amplified filtrate (dilute the amplified samples if necessary) by using specific primers designed for bacteria to confirm the bacteria species (see section 2.1).
- Conduct PCR to confirm whether there is contamination of host genome by using whitefly gene beta-Actin and EF1 primers according to previously published method21 (the PCR recipe is described in section 1.4).
6. Endosymbiont Metagenome Sequencing
- Subject the amplified filtrate to a spectrophotometer and a fluorometer (according to the manufacturer's instructions) before sequencing to test the quality of samples and whether the samples meet the criteria for sequencing.
- Subject the amplified to a genome sequencer according to the manufacturer's instructions.
Representative Results
The Middle East Asia Minor 1 (MEAM1) species of the B. tabaci complex was taken as an example here for description. Cotton for rearing whiteflies and several developmental stages of whiteflies are shown in Figure 1 including a cotton plant, adult whitefly and the 1st, 2nd and 4th instar nymphs of whitefly (the 3rd instar nymph looks similarly as the 4th instar nymph). It was obvious that the 4th instar nymph is larger than the 1st and 2nd instar nymphs (Figure 1). FISH analysis of Portiera and Hamiltonella within MEAM1 is shown in Figure 2. These two endosymbionts are confined to the bacteriocytes of the whitefly, and overlapping of the two bacteria is also observed. Figure 3 shows transmission electron microscopy images of the Portiera endosymbiont of the whitefly, which indicates that Portiera may lose the its cell wall.
For sequencing, two libraries were built, with insert sizes of 200 bp and 2,000 bp, respectively. After sequencing, the two libraries generated 1,626 Mb and 1,302 Mb of raw data, respectively. After cleaning up the contamination data for adaptors and duplications, 1,484 Mb and 1,219 Mb of clean data were acquired. The assembly successfully resulted in a complete genome sequence for the obligate symbiont, Portiera. In addition, a draft genome of Hamiltonella was also obtained. The assembly based on both 200 bp and 2,000 bp libraries resulted in 138 contigs. According to paired-end relationships, the 138 contigs were further assembled into 89 scaffolds (Table 2).

Figure 1: Overview of the Plant-insect System Utilized in this Paper. (A) Whiteflies are reared on cotton plants (Gossypium hirsutum cv. Zhe-Mian 1793). (B) View of an adult whitefly. (C) Several developmental stages of instar nymph in the whitefly life cycle. Red arrow refers to the egg of whitefly; Black arrow refers to the 1st instar nymph of whitefly; White arrow refers to the 2nd nymph of whitefly; Blue arrow refers to the 4th instar nymph of whitefly. Please click here to view a larger version of this figure.
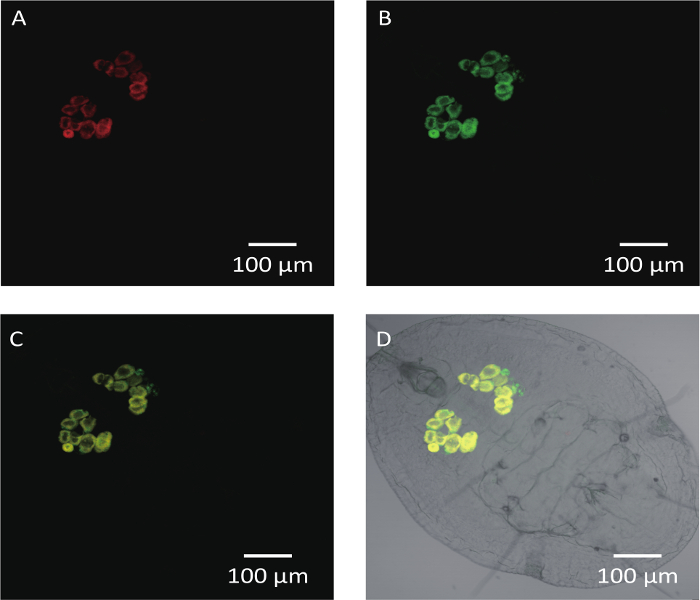

Figure 2: FISH Analysis of Portiera, Hamiltonella in the 4th Instar Nymph of MEAM1. Portiera-specific probe (red) conjugated to Cy3, Hamiltonella-specific probe (green) marked with Cy5 were used. Scale bars = 100 µm. (A) The red hybridization signals represent the Portiera under dark field. (B) The green hybridization signals represent the Hamiltonella under dark field. (C) Portiera and Hamiltonella merged signals under dark field. (D) Portiera and Hamiltonella merged signals under bright field. Please click here to view a larger version of this figure.
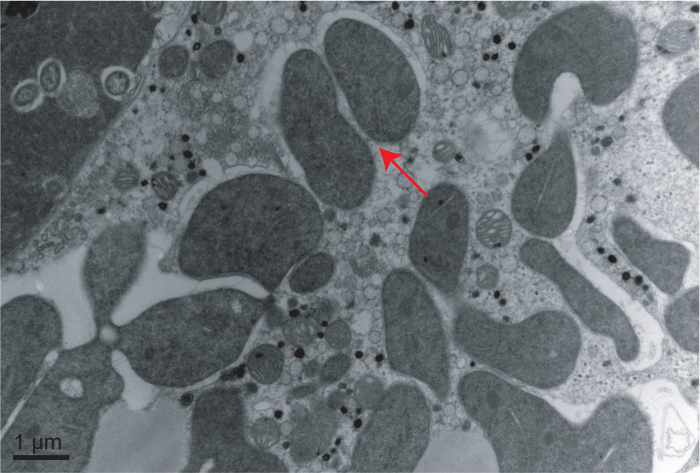
Figure 3: Transmission Electron Microscopy Image of Portiera in Whitefly. Arrow indicates the location of Portiera. Scale bar = 1 µm. Please click here to view a larger version of this figure.
| Target symbiont | Target gene | Primer sequences (5'-3') | PCR procedures | References | ||
| Portiera | 16S rRNA | Por-F: TGCAAGTCGAGCGGCATCAT | 95 °C 1 min, 60 °C 1 min, 72 °C 1 min, 5 cycles; | 27 | ||
| Por-R: AAAGTTCCCGCCTTATGCGT | 95 °C 1 min, 58 °C 1 min, 72 °C 1 min, 30 cycles; | |||||
| 72 °C 20 min | ||||||
| Hamiltonella | 16S rRNA | Ham-F: TGAGTAAAGTCTGGGAATCTGG | 95 °C 1 min, 60 °C 1 min, 72 °C 1 min, 5 cycles; | 27 | ||
| Ham-R: CCCGGGAACGTATTCACCGTAG | 95 °C 1 min, 58 °C 1 min, 72 °C 1 min, 30 cycles; | |||||
| 72 °C 20 min | ||||||
| Rickettsia | 16S rRNA | Ric-F: GCTCAGAACGAACGCTATC | 95 °C 2 min; | 24 | ||
| Ric-R: GAAGGAAAGCATCTCTGC | 92 °C 1 min, 58 °C 1 min, 72 °C 90 s, 30 cycles; | |||||
| 72 °C 5 min | ||||||
| Arsenophonus | 23S rRNA | Ars-F: CGTTTGATGAATTCATAGTCAAA | 95 °C 5 min; | 28 | ||
| Ars-R: GGTCCTCCAGTTAGTGTTACCCAAC | 95 °C 30 s, 60.5 °C 30 s, 72 °C 45 s, 30 cycles; | |||||
| 72 °C 10 min | ||||||
| Cardinium | 16S rRNA | Car-F: TACTGTAAGAATAAGCACCGGC | 95 °C 2 min | 29 | ||
| Car-R: GTGGATCACTTAACGCTTTCG | 92 °C 1 min, 57 °C 1 min, 72 °C 90 s, 30 cycles; | |||||
| 72 °C 5 min | ||||||
| Wolbachia | 16S rRNA | Wol-F: TTGTAGCCTGCTATGGTATAACT | 94 °C 5 min; | 30, 31 | ||
| Wol-R: GAATAGGTATGATTTTCATGT | 94 °C 1 min, 55 °C 1 min, 72 °C 1 min, 35 cycles | |||||
| Fritschea | 23S rRNA | Fri-F: GATGCCTTGGCATTGATAGGCGATGAAGGA | 95 °C 5 min; | 32 | ||
| Fri-R: TGGCTCATCATGCAAAAGGCA | 94 °C 1 min, 64 °C 1 min, 72 °C 90 s, 35 cycles; | |||||
| 72 °C 5 min | ||||||
| Hemipteriphilus | groEL | OR-groEL-F: CACCWAAAATTACTAAAGATGG | 94 °C 3 min; | 18 | ||
| OR-groEL-R: TAGAARTCCATWCCKCCCATWC | 94 °C 30 s, 52 °C 30 s, 72 °C 2 min, 34 cycles; | |||||
| 72 °C 10 min | ||||||
Table 1: PCR Primers and Procedures of Endosymbionts in Whitefly.
| Portiera | Hamiltonella | |
| Contig Number | 1 | 138 |
| Scaffold numbera | 1 | 89 |
| Total length, bp | 358,232 | 1,711,449 |
| N50, bp | / | 118,802 |
| N90, bp | / | 16,753 |
| Max length, bp | / | 390,511 |
| Min length, bp | / | 503 |
| GC content, % | 26.18 | 39.89 |
| a Information presented below are all based on scaffolds. | ||
Table 2: General Features of Portiera and Hamiltonella Draft Genome.
Discussion
Since the endosymbionts within whiteflies cannot be cultured in vitro, dissection and assembling bacteriocytes is an effective way to obtain enough genetic material of endosymbionts. Before dissection, the species of whitefly and endosymbionts involved should be explicitly confirmed. The whitefly B. tabaci is a species complex with more than 35 morphologically indistinguishable species and different cryptic species may contain different endosymbionts. Portiera is uniformly harbored as an obligate symbiont22, and secondary symbionts vary among species, even the individual whitefly17,21,23. Moreover, although many endosymbionts have been proven to be restrained within bacteriocytes, some may be distributed in the entire body of the whitefly24. Thus, analyzing the localization of different endosymbionts by FISH and TEM is necessary before bacteriome dissection.
For bacteriome dissection and any subsequent operations, the utilized materials must be strictly sterile to eliminate possible contamination of other bacteria, particularly the PBS solution, the filtration equipment, and the centrifuge tubes. Special care should be taken when filtering the PBS solution containing the bacteriomes to avoid environmental contamination. It is best to use a relatively aseptic environment. In addition, whitefly nymphs can be stored in absolute ethanol at room temperature before dissection, which may be useful to send samples to other places. Nevertheless, dead or environmental stress treated insects should not be used because there would be declination of bacteria25,26.
In this protocol, the 3rd or 4th instar nymphs were utilized to isolate bacteriome. Compared to other stages of the whitefly, there are several advantages of using the 3rd or 4th instar nymphs for dissection. Firstly, compared to the 1st and 2nd instar nymphs, whiteflies in this stage are obviously larger and not so fragile. Secondly, in the 3rd or 4th instar nymphs of whitefly, bacteriome can be easily distinguished from other tissues of the whitefly body with an obviously yellow color, and bacteriocytes always cluster firmly and can be conveniently separated from other tissues. For the bacteriome in adult whiteflies, once pulled out from the body, bacteriocytes would separate quite easily thus increasing the difficulty in assembly. Hence, the best choice for dissecting bacteriomes is the 3rd or 4th instar nymphs. Nevertheless, this protocol can be used with adult whiteflies for certain experiment needs, with more practice.
Compared to a previously published method using a single bacteriocyte21, this protocol assembled more bacteriocytes used in the amplification, which contributed to the richness and diversity of bacteriocytes. Furthermore, we utilized filtration to separate symbiont cells from whitefly bacteriocytes, which is more convenient than another method using a micromanipulator and a microinjector. Nevertheless, there were some limitations of this protocol. Firstly, symbionts not confined to bacteriocytes cannot be separated by this method, since we only dissect the bacteriome of the whitefly. Secondly, this protocol cannot absolutely eliminate the contamination of the whitefly genome, since some fragments of whitefly tissues would pass through the filter membrane. Thirdly, clustered symbionts cannot be further separated into individual symbiont species. Hence, obtaining a single species of bacterium accurately and purely requires further study.
In summary, we reported a convenient method for the extraction and purification of endosymbionts from the whitefly B. tabaci that is suitable for subsequent sequencing and analysis. The protocol described here can be easily adapted for the isolation of endosymbionts from other insects for genome sequencing.
Disclosures
The authors have nothing to disclose.
Acknowledgements
Financial support for this study was provided by the National Key Research and Development Program (2016YFC1200601) and the National Natural Science Foundation of China (31390421).
Materials
| Taq DNA polymerase | Takara | R001A | including rTaq, 10×Buffer and dNTP |
| Gel DNA extraction kit | Qiagen | 28704 | |
| DNA sample sequencing system | ABI | ABI-3730XL | |
| Microtome | Leica | EM UC7 | |
| Transmission electron microscopy | Hitachi | H-7650 TEM | |
| Stereo microscope | Zeiss | Stemi 2000-C | |
| 20 μL microloader | Eppendorf | F2771951 | |
| Filter holder | Millipore | SX0001300 | |
| Filter membrane filter | Millipore | SMWP001300 | 5.0 μm SMWP |
| REPLI-g UltraFast Mini Kit | Qiagen | 150033 | DNA amlification kit |
| NanoDrop | Thermo Scientific | NanoDrop 2000 | |
| Qubit Fluorometer | Thermo Fisher Scientific | Q33216 | |
| Genome Sequencer | Illumina | Hiseq 2000 |
References
- Buchner, P. Endosymbiosis of animals with plant microorganisms. J. Basic Microbiol. 7 (2), (1967).
- Sloan, D. B., Moran, N. A. Endosymbiotic bacteria as a source of carotenoids in whiteflies. Biol. Letters. 8 (6), 986-989 (2012).
- Stouthamer, R., Breeuwer, J. A., Hurst, G. D. Wolbachia pipientis: microbial manipulator of arthropod reproduction. Annu. Rev. Microbiol. 53, 71-102 (1999).
- Brumin, M., Kontsedalov, S., Ghanim, M. Rickettsia influences thermotolerance in the whitefly Bemisia tabaci B biotype. Insect Sci. 18 (1), 57-66 (2011).
- Hansen, A. K., Degnan, P. H. Widespread expression of conserved small RNAs in small symbiont genomes. ISME J. 8, 2490-2502 (2014).
- Moran, N. A., McCutcheon, J. P., Nakabachi, A. Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 42, 165-190 (2008).
- Shigenobu, S., Watanabe, H., Hattori, M., Sakaki, Y., Ishikawa, H. Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. APS. Nature. 407, 81-86 (2000).
- Nikoh, N., et al. Evolutionary origin of insect-Wolbachia nutritional mutualism. Proc. Natl. Acad. Sci. U. S. A. 111 (28), 10257-10262 (2014).
- Sloan, D. B., et al. Parallel histories of horizontal gene transfer facilitated extreme reduction of endosymbiont genomes in sap-feeding insects. Mol. Biol. Evol. 31 (4), 857-871 (2014).
- Xue, J., et al. Genomes of the rice pest brown planthopper and its endosymbionts reveal complex complementary contributions for host adaptation. Genome Biol. 15 (12), 521 (2014).
- Campbell, M. A., et al. Genome expansion via lineage splitting and genomereduction in the cicada endosymbiont Hodgkinia. Proc. Natl. Acad. Sci. U. S. A. 112 (33), 10192-10199 (2015).
- Wilson, A. C., et al. Genomic insight into the amino acid relations of the pea aphid, Acyrthosiphon pisum, with its symbiotic bacterium Buchnera aphidicola. Insect Mol. Biol. 19, 249-258 (2010).
- Degnan, P. H., Ochman, H., Moran, N. A. Sequence conservation and functional constraint on intergenic spacers in reduced genomes of the obligate symbiont Buchnera. PLoS Genet. 7 (9), e1002252 (2011).
- Nakabachi, A., et al. The 160-kilobase genome of the bacterial endosymbiont Carsonella. Science. 314 (5797), 267 (2006).
- De Barro, P. J., Liu, S. S., Boykin, L. M., Dinsdale, A. B. Bemisia tabaci: a statement of species status. Annu. Rev. Entomol. 56, 1-19 (2011).
- Himler, A. G., et al. Rapid spread of a bacterial symbiont in an invasive whitefly is driven by fitness benefits and female bias. Science. 332 (6026), 254-256 (2011).
- Bing, X. L., Ruan, Y. M., Rao, Q., Wang, X. W., Liu, S. S. Diversity of secondary endosymbionts among different putative species of the whitefly Bemisia tabaci. Insect Sci. 20 (2), 194-206 (2013).
- Bing, X. L., Yang, J., Zchori-Fein, E., Wang, X. W., Liu, S. S. Characterization of a newly discovered symbiont of the whitefly Bemisia tabaci (Hemiptera: Aleyrodidae). Appl. Environ. Microbiol. 79 (2), 569-575 (2013).
- Kliot, A., et al. Fluorescence in situ hybridizations (FISH) for the localization of viruses and endosymbiotic bacteria in plant and insect tissues. J. Vis. Exp. (84), e51030 (2014).
- Lee, P. Y., Costumbrado, J., Hsu, C., Kim, Y. H. Agarose gel electrophoresis for the separation of DNA fragments. J. Vis. Exp. (62), e3923 (2012).
- Rao, Q., et al. Genome reduction and potential metabolic complementation of the dual endosymbionts in the whitefly Bemisia tabaci. BMC Genomics. 16 (1), 226 (2015).
- Thao, L. L., Baumann, P. Evolutionary relationships of primary prokaryotic endosymbionts of whiteflies and their hosts. Appl. Environ. Microbiol. 70 (6), 3401-3406 (2004).
- Zhu, D. T., et al. Sequencing and comparison of the Rickettsia genomes from the whitefly Bemisia tabaci Middle East Asia Minor I. Insect Sci. 23 (4), 531-542 (2016).
- Gottlieb, Y., et al. Identification and localization of a Rickettsia sp. in Bemisia tabaci (HomopteraAleyrodidae). Appl. Environ. Microbiol. 72 (5), 3646-3652 (2006).
- Zhang, C. R., et al. Differential temporal changes of primary and secondary bacterial symbionts and whitefly host fitness following antibiotic treatments. Sci. Rep. 5, 15898 (2015).
- Shan, H. W., et al. Temporal changes of symbiont density and host fitness after rifampicin treatment in a whitefly of the Bemisia tabaci species complex. Insect Sci. 23 (2), 200-214 (2016).
- Zchori-Fein, E., Brown, J. K. Diversity of prokaryotes asscociated with Bemisia tabaci (Gennadius) (Hemiptera: Aleyrodidae). Ann. Entomol. Soc. Am. 95 (6), 711-718 (2002).
- Thao, M. L., Baumann, P. Evolutionary relationships of primary prokaryotic endosymbionts of whiteflies and their hosts. Appl. Environ. Microbiol. 70 (6), 3401-3406 (2004).
- Zchori-Fein, E., Perlman, S. J. Distribution of the bacterial symbiont Cardinium in arthropods. Mol. Ecol. 13 (7), 2009-2016 (2004).
- O’Neill, S. L., Giordano, R., Colbert, A. M., Karr, T. L., Robertson, H. M. 116S rRNA phylogenetic analysis of the bacterial endosymbionts associated with cytoplasmic incompatibility in insects. Proc. Natl. Acad. Sci. U. S. A. 89 (7), 2699-2702 (1992).
- Nirgianaki, A. Wolbachia infections of the whitefly Bemisia tabaci. Curr. Microbiol. 47 (2), 93-101 (2003).
- Everett, K. D., Thao, M. L., Horn, M., Dyszynski, G. E., Baumann, P. Novel chlamydiae in whiteflies and scale insects: endosymbionts ‘Candidatus Fritschea bemisiae’ strain Falk and ‘Candidatus Fritschea eriococci’ strain Elm. Int. J. Syst. Evol. Micr. 55, 1581-1587 (2005).

