Enrichment of Bacterial Lipoproteins and Preparation of N-terminal Lipopeptides for Structural Determination by Mass Spectrometry
Summary
The enrichment of bacterial lipoproteins using a non-ionic surfactant phase partitioning method is described for direct use in TLR assays or other applications. Further steps are detailed to prepare N-terminal tryptic lipopeptides for structural characterization by mass spectrometry.
Abstract
Lipoproteins are important constituents of the bacterial cell envelope and potent activators of the mammalian innate immune response. Despite their significance to both cell physiology and immunology, much remains to be discovered about novel lipoprotein forms, how they are synthesized, and the effect of the various forms on host immunity. To enable thorough studies on lipoproteins, this protocol describes a method for bacterial lipoprotein enrichment and preparation of N-terminal tryptic lipopeptides for structural determination by matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS). Expanding on an established Triton X-114 phase partitioning method for lipoprotein extraction and enrichment from the bacterial cell membrane, the protocol includes additional steps to remove non-lipoprotein contaminants, increasing lipoprotein yield and purity. Since lipoproteins are commonly used in Toll-like receptor (TLR) assays, it is critical to first characterize the N-terminal structure by MALDI-TOF MS. Herein, a method is presented to isolate concentrated hydrophobic peptides enriched in N-terminal lipopeptides suitable for direct analysis by MALDI-TOF MS/MS. Lipoproteins that have been separated by Sodium Dodecyl Sulfate Poly-Acrylamide Gel Electrophoresis (SDS-PAGE) are transferred to a nitrocellulose membrane, digested in situ with trypsin, sequentially washed to remove polar tryptic peptides, and finally eluted with chloroform-methanol. When coupled with MS of the more polar trypsinized peptides from wash solutions, this method provides the ability to both identify the lipoprotein and characterize its N-terminus in a single experiment. Intentional sodium adduct formation can also be employed as a tool to promote more structurally informative fragmentation spectra. Ultimately, enrichment of lipoproteins and determination of their N-terminal structures will permit more extensive studies on this ubiquitous class of bacterial proteins.
Introduction
Bacterial lipoproteins are characterized by a conserved N-terminal lipid-modified cysteine that anchors the globular protein domain to the cell membrane surface. They are universally distributed in bacteria, constituting 2 to 5% of all cellular genes within a typical genome1. Lipoproteins play critical roles in a wide variety of cellular processes, including nutrient uptake, signal transduction, assembly of protein complexes, and in maintaining cell envelope structural integrity2. In pathogenic bacteria, lipoproteins serve as virulence factors3,4. During an infection, recognition of N-terminal lipopeptides by Toll-like receptor (TLR) 2 incites an innate immune response to remove invading pathogens. Depending on the N-terminal acylation state, lipoproteins are generally recognized by alternate TLR2 heterodimeric complexes. TLR2-TLR1 recognizes N-acylated lipopeptides, while TLR2-TLR6 binds free lipopeptide α-amino termini. Once bound, the signaling pathways converge to induce secretion of proinflammatory cytokines3,4.
It was previously thought that lipoproteins from Gram-positive bacteria were diacylated and those from Gram-negative bacteria were triacylated, differing in the absence or presence of an amide-linked fatty acid on the conserved N-terminal cysteine residue. This assumption was supported by the lack of sequence orthologs in Gram-positive genomes to Lnt, the Gram-negative N-acyl transferase that forms triacylated lipoproteins5. However, recent studies have revealed lipoprotein triacylation in Gram-positive Firmicutes that lack lnt, as well as three novel N-terminal lipoprotein structures, termed the peptidyl, lyso, and N-acetyl forms6,7,8. These findings raise questions about possible yet-to-be-discovered lipoprotein forms, alongside fundamental questions about how these novel lipoproteins are made and what physiological purpose or advantage the various forms impart. Furthermore, they clearly demonstrate the current inability of genomics to predict lipoprotein structure. Indeed, we recently identified a novel class of lipoprotein N-acyl transferases, called Lit, from Enterococcus faecalis and Bacillus cereus that makes lyso-form lipoproteins9. This indicates the need to experimentally verify lipoprotein structure, which can be challenging due to their extremely hydrophobic nature and limited methods available to characterize their molecular structure.
To facilitate studies on lipoprotein induction of the host immune response, as well as N-terminal structural determination, we have adapted several previously-described protocols in order to purify bacterial lipoproteins and prepare the N-terminal tryptic lipopeptides for analysis by MALDI-TOF MS6,10,11,12. Lipoproteins are enriched using an established Triton X-114 (henceforth referred to as surfactant or TX-114) phase partitioning method, with optimization to remove contaminating non-lipoproteins and increase lipoprotein yield. These lipoproteins are suitable for direct use in TLR assays or for further purification by SDS-PAGE. For MALDI-TOF MS, transfer of the lipoproteins to nitrocellulose membrane provides a scaffold for efficient in situ trypsin digestion, washing, and subsequent elution from the membrane surface, resulting in highly purified N-terminal lipopeptides. Nitrocellulose has been shown to facilitate sample handling and improve sequence coverage for highly hydrophobic peptides from integral membrane proteins13,14, as well as lipoproteins9,10. The method has the additional advantage of fractionating peptides based on polarity, so that intermediate wash solutions can be analyzed for high confidence protein identification simultaneously with N-terminal structural determination in a single experiment. This protocol uniquely features intentional sodium adduct formation to promote parent ion fragmentation towards dehydroalanyl ions during MS/MS, aiding in structural assignment of the N-acylation state. The N-terminus is both the most variable and key feature related to TLR recognition of lipoproteins. Taken together, this protocol has allowed intensive and reproducible studies on lipoproteins, with the individual stages of purification and structural determination by MALDI-TOF MS easily adapted depending on the overall goal of the experiment.
Protocol
1. Cell Growth and Lysis
- Grow bacteria in 15 mL tryptic soy broth (TSB) or similar rich media to late exponential phase (OD600 of 1.0 – 1.5). Harvest cells by centrifugation, wash once with Tris-buffered saline/EDTA (TBSE) and continue with protocol or freeze until use.
NOTE: TBSE: 20 mM Tris-hydrochloride (HCl), pH 8.0, 130 mM sodium chloride (NaCl), and 5 mM ethylenediaminetetraacetic acid (EDTA)). Lipoprotein expression and modification can be influenced by growth conditions (e.g. acidity, salinity, growth media) and growth phase15. Cell growth can be scaled as desired, but 15-mL of cells is recommended for a single preparation of lipoproteins as excess biomass can decrease lipoprotein yield. One 15-mL preparation generally yields enough sample for optimal loading of ~2-4 lanes on a standard SDS-PAGE mini gel. - Resuspend cells in 800 μL TBSE with 1 mM phenylmethyl sulfonyl fluoride (PMSF) and 0.5 mg/mL lysozyme. Transfer solution to a 2.0-mL threaded microcentrifuge tube with screw cap and O-ring and incubate for 20 min at 37 °C.
NOTE: Certain species are not susceptible to lysis by lysozyme. An appropriate lytic enzyme should be substituted to aid in lysis. - Add ~800 μL 0.1 mm zirconia/silica beads to the tube and disrupt cells by shaking at maximum speed (7,000 rpm) on a homogenizer for 5 cycles of 30 s each, with 2 min rest on ice between each cycle.
- Centrifuge sample at 3000 x g for 5 min at 4 °C (to pellet beads and unbroken cells). Transfer the supernatant to a new 2.0-mL microcentrifuge tube and keep on ice.
- Add 200 μL TBSE to the remaining pellet and return to the homogenizer for an additional cycle. Centrifuge (as above) and combine supernatant with the previous supernatant (should be ~800-1000 μL total volume).
2. Enrichment of Lipoproteins by TX-114 Phase Partitioning
- Supplement the supernatant with TX-114 surfactant to a final concentration of 2% (vol/vol)by adding an equal volume of 4% (vol/vol) the surfactant in ice-cold TBSE and incubate on ice for 1 h, mixing by inversion every ~15 min.
NOTE: When chilled, the supernatant and surfactant will be miscible. - Transfer the tube to a 37 °C water bath and incubate for 10 min to induce phase separation. Centrifuge the sample at 10,000 x g for 10 min at room temperature to maintain bi-phasic separation.
- Gently pipette off the upper aqueous phase and discard. Add ice-cold TBSE to the lower surfactant phase to refill the tube to its original volume and invert to mix. Incubate on ice for 10 min.
- Transfer the tube to a 37 °C water bath and incubate for 10 min to induce phase separation, then centrifuge at 10,000 x g for 10 min at room temperature.
- Repeat steps 2.3-2.4 once more for a total of 3 separations. Remove the upper aqueous phase and discard.
- Remove pellet of precipitated proteins that formed during the course of extractions (visible at the bottom of the tube), by adding 1 volume of ice-cold TBSE to the surfactant phase. Centrifuge at 4 °C at 16,000 x g for 2 min to pellet insoluble protein.
NOTE: Sample should remain a single phase. If phase separation occurs, re-chill sample and centrifuge again. The pellet is generally composed of non-lipoprotein contaminants, but can be retained for further analysis. - Immediately transfer the supernatant to a fresh 2.0-mL microcentrifuge tube containing 1250 μL of 100% acetone. Pipette up and down thoroughly to wash the sample from the tip as it will be viscous. Mix by inversion and incubate overnight at -20 °C to precipitate protein.
- Centrifuge the sample at 16,000 x g for 20 min at room temperature to pellet lipoproteins with attention to the orientation of the tube. Lipoproteins will form a thin, white film along the outside wall of the tube.
- Wash pellet twice with 100% acetone. Decant acetone and allow sample to air dry.
- Add 20-40 μL of 10 mM Tris-HCl, pH 8.0 or a standard Laemmli SDS-PAGE sample buffer16 and thoroughly resuspend by pipetting up and down against the wall with the precipitated lipoproteins. Scrape the side of the tube with the pipette tip to dislodge lipoproteins.
NOTE: Lipoproteins will not dissolve in Tris buffer, but rather form a suspension.- Store lipoproteins at -20 °C until use.
3. SDS-PAGE, Electroblotting, and Staining with Ponceau S
- Separate lipoproteins by SDS-PAGE using standard methods16.
NOTE: The appropriate gel acrylamide percentage will vary depending on its intended use and size of lipoproteins. Here, E. faecalis lipoproteins were separated over a 10% Tris-glycine gel, while a 16.5% Tris-tricine gel was used for the smaller E. coli lipoprotein Lpp17. - Transfer the lipoproteins to a nitrocellulose transfer membrane using a standard electroblotting procedure.
NOTE: A semi-dry transfer using Bjerrum Schafer-Nielsen buffer plus 0.1% SDS was performed at 25 V, 1.3 mA for 15 min18. Alternately, polyvinylidene difluoride (PVDF) membrane could be used, however as it is more hydrophobic than nitrocellulose, it may reduce efficient elution of hydrophobic lipopeptides from the membrane at later steps. - Transfer the nitrocellulose membrane to a container and cover with Ponceau S solution (0.2% (w/v) Ponceau S in 5% acetic acid). Rock gently for 5 min or until red-pink bands are visible.
- Pour off Ponceau S solution and carefully rinse the nitrocellulose membrane with dH2O to remove excess stain.
NOTE: Ponceau-stained bands will destain rapidly. If bands disappear, repeat staining process. - With a clean razor blade, excise the desired band and transfer to a microcentrifuge tube. Wash three times with 1 mL of dH2O to completely destain the band.
NOTE: The protocol can be paused here. Store the excised section covered with water at -20 °C until use. - Transfer the section to a clean surface and with a clean razor blade, dice the nitrocellulose strip into small pieces of approximately 1 mm x 1 mm. Collect the pieces into a 0.5-mL low protein binding microcentrifuge tube.
- Wash the pieces twice with 0.5 mL of freshly-prepared 50 mM ammonium bicarbonate (NH4HCO3), pH 7.8 in high-performance liquid chromatography (HPLC) grade water.
4. Tryptic Digestion, Lipopeptide Extraction from Nitrocellulose Membrane, Deposition onto MALDI Target, and Data Acquisition
- Resuspend nitrocellulose pieces in 20 μL of a 20 μg/mL solution of trypsin in 50 mM NH4HCO3, pH 7.8 in HPLC grade water. Vortex to mix, then spin briefly to ensure that all pieces are fully covered by the trypsin solution. Cover the tube lid using paraffin film to prevent evaporation and incubate digest overnight at 37 °C.
- Spin the sample at 16,000 x g for 30 s and remove the liquid by pipetting.
NOTE: This removes trypsinized hydrophilic peptides that can be examined later by MALDI-TOF MS for protein identification. - Add 50 μL of 0.5% trifluoroacetic acid (TFA) in HPLC grade water. Vortex to mix, then spin the sample briefly to ensure all pieces are covered by the solution. Incubate for 10 min at room temperature. Remove the liquid by pipetting.
CAUTION: TFA is harmful if inhaled. Handle with caution and use in the chemical fume hood. Avoid contact with skin. - Repeat step 4.3 with 50 μL of 10% acetonitrile in HPLC grade water.
CAUTION: Acetonitrile is harmful if inhaled. Handle with caution and use in the chemical fume hood. Avoid contact with skin. - Repeat step 4.3 with 50 μL of 20% acetonitrile in HPLC grade water.
NOTE: This removes any moderately hydrophobic peptides loosely bound to the nitrocellulose. - To elute the tightly-bound lipopeptides, add 15 μL of freshly-made 10 mg/mL α-cyano-4-hydroxycinnamic acid (CHCA) matrix dissolved in chloroform-methanol (2:1, v/v) and incubate for 10 min at room temperature with intermittent vortexing.
- Alternately, add 15 μL chloroform-methanol to elute lipopeptides and mix with matrix at a later time. Other matrices, such as 2,5-dihydroxybenzoic acid (DHB), should be tested as alternatives to CHCA if unsatisfactory results are obtained for a particular lipopeptide composition.
CAUTION: Both chloroform and methanol are harmful if inhaled. Handle with caution and use in the chemical fume hood. Avoid contact with skin. - Optional: To promote sodium adduct formation, supplement the solution of CHCA in chloroform-methanol with aqueous sodium bicarbonate (NaHCO3) to a final concentration of 1 mM.
- Alternately, add 15 μL chloroform-methanol to elute lipopeptides and mix with matrix at a later time. Other matrices, such as 2,5-dihydroxybenzoic acid (DHB), should be tested as alternatives to CHCA if unsatisfactory results are obtained for a particular lipopeptide composition.
- Spin the sample briefly. With a pipette, carefully transfer the liquid to a new low protein binding microcentrifuge tube. This solution will contain the bulk of the N-terminal lipopeptides.
NOTE: The nitrocellulose may partially or fully dissolve in the chloroform-methanol solution. This will not negatively affect MS results, and may be used as an alternate method to increase lipopeptide return. PVDF membrane will not dissolve in the same manner. - Deposit 1 μL of the eluted lipopeptides with CHCA onto a polished steel MALDI target.
NOTE: The chloroform-methanol will evaporate quickly, leaving behind crystallized CHCA and lipopeptides. An optional second 1 μL aliquot can be deposited onto the same spot to increase sample concentration. Chloroform-methanol may spread significantly after deposition onto the target and as such, care should be taken to avoid sample mixing on the target. It is recommended to deposit and shoot multiple spots of each sample. - Immediately proceed to mass spectrometry. Scan all areas of the spot for lipopeptide signal, especially if the spot contains two layers of sample, as the second layer may push the lipopeptides to the spot's outer rim.
NOTE: Recommended starting laser intensity is 25%, though it may be necessary to increase intensity to acquire signal and sum multiple spectra (20 or more scans) to achieve adequate signal-to-noise ratio. Here, spectra were acquired on a MALDI-TOF-TOF instrument (see the Table of Materials) using a factory-configured instrument method for the reflector positive-ion detection over the 700-3500 m/z range. The instrument was calibrated with a bovine serum albumin (BSA) tryptic peptide mixture.
Representative Results
A schematic of the protocol is provided in Figure 1. The lipoprotein-enriched fraction extracted from Enterococcus faecalis ATCC 19433 by TX-114 is shown in Figure 2. For comparison, the banding pattern of the precipitated protein fraction is also shown. Proteins from this fraction were confirmed by MALDI-MS to be highly abundant contaminating proteins other than lipoproteins (Table 1). The mass spectra in Figure 3 demonstrate the tryptic peptide ion profile of the E. faecalis lipoprotein PnrA that occurs with subsequent washes of nitrocellulose-bound PnrA with solvents of increasing polarity (peak assignments listed in Table 2). Figure 4 shows the N-terminal structural characterization of PnrA as determined by MALDI-TOF MS/MS, revealing diagnostic N-acylated dehydroalanyl peaks consistent with the lyso-lipoprotein form. Figure 5 illustrates the effect of sodium adduct formation on fragmentation, with the sodiated parent ion preferentially fragmenting in favor of the N-acylated dehydroalanyl ion.
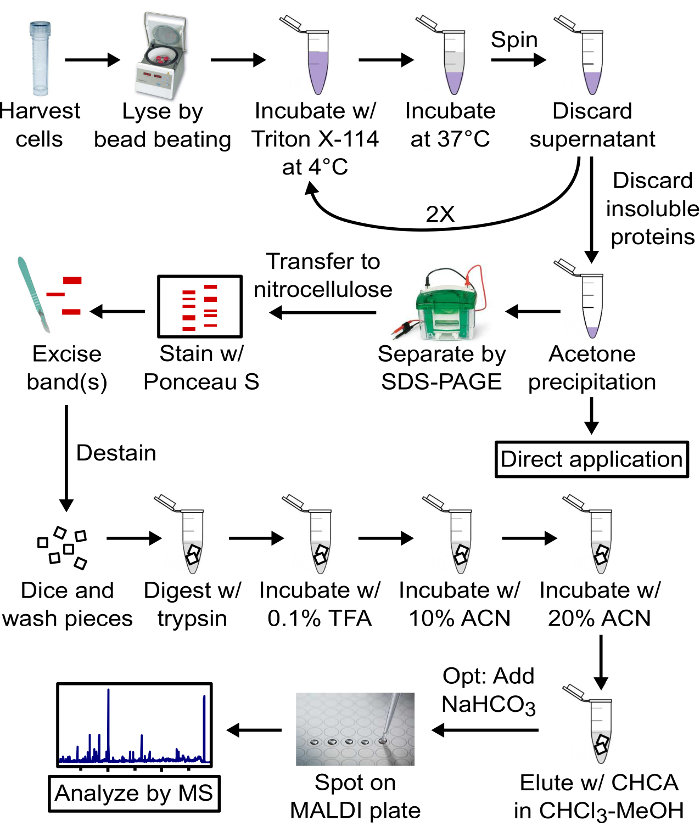
Figure 1: Schematic of the protocol. Lipoproteins are enriched by TX-114 phase partitioning and can be used directly, most commonly in TLR assays, or further purified by SDS-PAGE for structural determination. Lipoproteins are transferred to nitrocellulose, digested with trypsin, washed stepwise, and the resulting lipopeptides eluted with chloroform-methanol for structural analysis by MALDI-MS. The trypsin and nitrocellulose wash solutions may be saved for protein identification and MS analysis. w/: with; TFA: trifluoroacetic acid; ACN: acetonitrile; CHCA: α-cyano-4-hydroxycinnamic acid; opt: optional; MALDI: matrix-assisted laser desorption ionization; MS: mass spectrometry. Please click here to view a larger version of this figure.
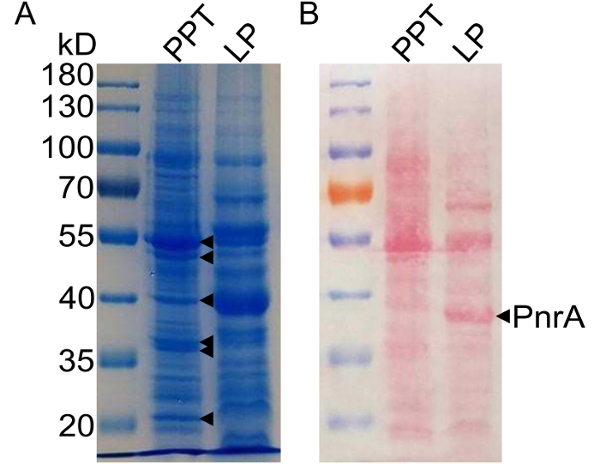
Figure 2: Profile of TX-114 enriched proteins from E. faecalis. A 10% Tris-glycine SDS-PAGE gel stained with Coomassie blue (A) and the corresponding nitrocellulose membrane stained with Ponceau S (B) reveal a different banding pattern between the precipitated protein fraction ("PPT") and the purified lipoproteins ("LP"). The indicated Coomassie-stained bands were excised and identified as non-lipoproteins by MALDI-MS of tryptic peptides (see Table 1). PnrA was identified and analyzed by the protocol described herein. Please click here to view a larger version of this figure.
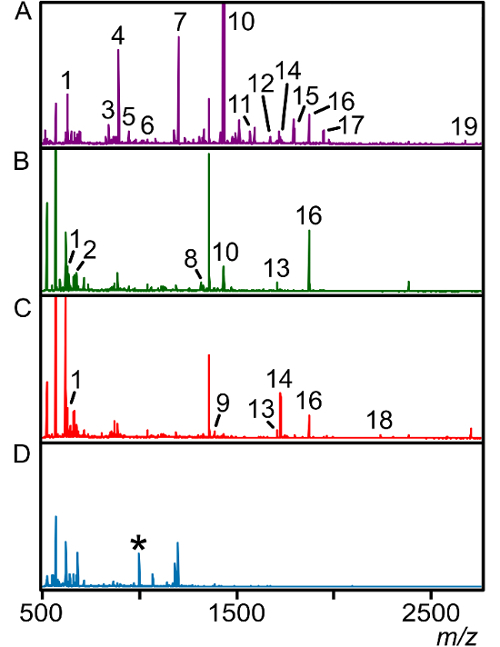
Figure 3: Ion profile and abundance changes with polarity of nitrocellulose wash solutions. (A) The trypsin fraction shows several internal peptide fragments corresponding to the E. faecalis lipoprotein PnrA indicated in Figure 2. The 10% (B) and 20% (C) acetonitrile wash fractions show changes in signal intensity and a decrease in the overall number of individual peaks. (D) The final elution fraction is highly enriched with the N-terminal lipopeptide at m/z 997, indicated by an asterisk (*). Intensity of each spectra is normalized to (A) for comparison of signal intensity. Peaks masses and assigned sequences are listed in Table 2. Please click here to view a larger version of this figure.
| Band | Protein ID | Accession No. | Est. Molecular Weight | Peptide Count | |
| 1 | elongation factor Tu | gb|EOL37301.1| | 43,387 | 15 | |
| 2 | NADH peroxidase | gb|EOL34572.1| | 49,520 | 9 | |
| 3 | pyruvate dehydrogenase E1component, alpha subunit | gb|EOL34709.1| | 41,358 | 12 | |
| 4 | pyruvate dehydrogenase E1 component, subunit beta | gb|EOL34710.1| | 35,373 | 19 | |
| 5 | 30S ribosomal protein S2 | gb|EOL33066.1| | 29,444 | 16 | |
| 6 | 30S ribosomal protein S3 | gb|EOL37312.1| | 24,355 | 14 | |
Table 1: Precipitated proteins are non-lipoprotein contaminants. Proteins were identified by tryptic digest and MALDI-MS using standard in gel digestion protocols19 and are listed from top to bottom in the same order they appear on the Coomassie gel in Figure 2. Each protein was identified with a Confidence Interval (C.I.) greater than 95%.
| Peak Number | Theoretical Mass | Protein Position | No. of Missed Cut Sites | Peptide Sequence | |
| 1 | 631.3409 | 170-176 | 0 | GVADAAK | |
| 2 | 675.4035 | 316-321 | 1 | TKEAVK | |
| 3 | 826.4093 | 163-169 | 0 | FQAGFEK | |
| 4 | 893.4839 | 121-128 | 0 | NVVSATFR | |
| 5 | 943.5359 | 240-247 | 0 | VWVIGVDR | |
| 6 | 1040.5047 | 188-197 | 0 | YAASFADPAK | |
| 7 | 1200.6331 | 273-284 | 0 | GVGTAVQDIANR | |
| 8 | 1316.6997 | 290-301 | 0 | FPGGEHLVYGLK | |
| 9 | 1385.7827 | 83-94 | 0 | FNTIFGIGYLLK | |
| 10 | 1432.743 | 149-162 | 0 | VGFVGGEEGVVIDR | |
| 11 | 1568.7802 | 259-272 | 1 | DGKEDNFTLTSTLK | |
| 12 | 1672.8289 | 240-254 | 1 | VWVIGVDRDQDADGK | |
| 13 | 1707.8184 | 129-145 | 0 | DNEAAYLAGVAAANETK | |
| 14 | 1724.8027 | 35-49 | 0 | SFNQSSWEGLQAWGK | |
| 15 | 1797.9228 | 257-272 | 2 | TKDGKEDNFTLTSTLK | |
| 16 | 1872.9854 | 285-301 | 1 | ALEDKFPGGEHLVYGLK | |
| 17 | 1945.9613 | 230-247 | 1 | DLNESGSGDKVWVIGVDR | |
| 18 | 2240.1345 | 149-169 | 1 | VGFVGGEEGVVIDRFQAGFEK | |
| 19 | 2675.2543 | 230-254 | 2 | DLNESGSGDKVWVIGVDRDQDADGK | |
Table 2: Observed masses and the corresponding tryptic peptides. In silico trypsin digestion of E. faecalis PnrA (GenBank: EOL37280.1) was performed using PeptideMass20,21, allowing up to two missed cut sites. The theoretical masses of the peaks observed on the spectra in Figure 3 are listed, along with the corresponding peptide sequences.
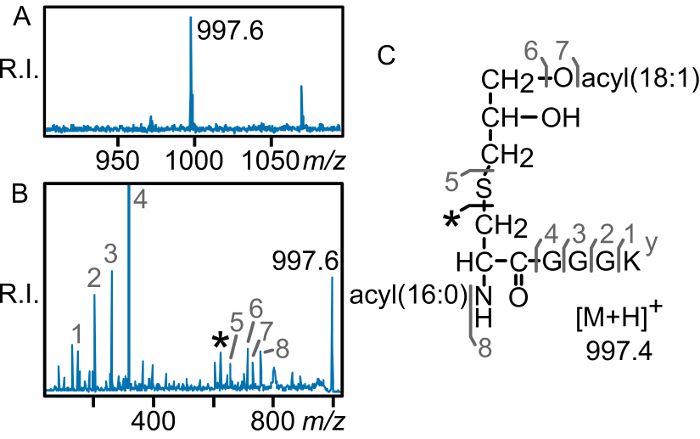
Figure 4: MALDI-TOF MS of the E. faecalis lipoprotein PnrA. (A) Parent MS spectrum of the m/z 997 region corresponding to the N-terminal lipopeptide of E. faecalis PnrA. (B) MS/MS of the lipopeptide peak reveals that it is the lyso form, with the diagnostic N-acylated dehydroalanyl peptide fragment ion peak indicated by an asterisk (*). The elucidated structure is shown (C). This figure has been modified from a previous publication9. R.I.: relative intensity Please click here to view a larger version of this figure.
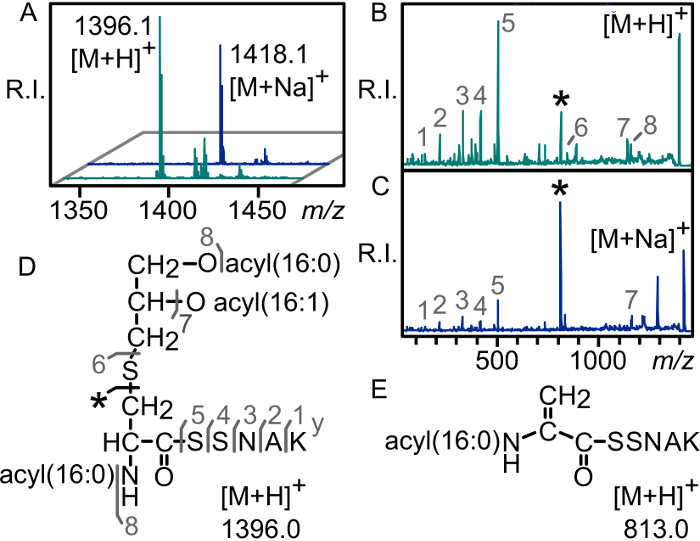
Figure 5: Sodium adduct formation promotes parent ion fragmentation towards dehydroalanyl lipopeptide ions. (A) MALDI-TOF spectra of the E. coli lipoprotein Lpp N-terminal peptide without (turquoise traces) and with (blue traces) the addition of sodium bicarbonate. Addition of sodium bicarbonate to the final eluted fraction results in a 22 Da increase from the calculated mass of the parent lipopeptide. When compared with the MS/MS spectrum of the protonated ion (B), the MS/MS spectrum of the corresponding sodiated ion (C) shows significant preferential fragmentation toward the N-acyl-dehydroalanyl ion through neutral elimination of the diacylthioglyceryl moiety, indicated by an asterisk (*). The structure of the parent triacylated Lpp N-terminal tryptic peptide (D) and the N-acyl-dehydroalanyl peptide fragment ion (E) are depicted. This figure has been modified from a previous publication9. R.I.: relative intensity Please click here to view a larger version of this figure.
Discussion
The protocol herein describes two distinct stages of lipoprotein characterization: enrichment by TX-114 phase partitioning and structural determination by MALDI-TOF MS. During TX-114 extraction, additional centrifugation removes contaminating proteins that precipitate during this process, followed by acetone precipitation to yield highly enriched lipoproteins. By limiting the scale of each preparation to 15-mL worth of cells, several samples can be easily processed in parallel, and if desired, pooled at the end of the protocol.
For structural analysis by MS, transfer of the lipoproteins to nitrocellulose facilitates trypsin digestion, washing, and subsequent elution from the membrane. In our hands, this approach has proven to be preferable to detergent-mediated extraction from traditional polyacrylamide gel plugs, as the lipoproteins are associated with the surface of the nitrocellulose membrane and not embedded in polyacrylamide. Lipoprotein association with the nitrocellulose surface also largely prevents the common problem of nonspecific adsorption to container surfaces by hydrophobic peptides. Stepwise elution of the nitrocellulose pieces removes more hydrophilic, internal tryptic peptides which can be analyzed by MALDI-TOF MS to achieve high confidence protein identification assignments, while the final elution step reproducibly yields concentrated lipopeptides in a small volume that is largely free of interfering salts and ion suppressing contaminants.
Successful MS analysis of a lipoprotein is subject to its abundance, and as such, the darkest bands from the Ponceau S-stained membrane are likely to give the best results. However, it is possible that several lipoproteins may migrate in a single band during PAGE separation, complicating the resulting spectra. Therefore, it is highly recommended to first identify the protein by peptide mass fingerprinting (PMF), which can be accomplished by collecting MS spectra on the total trypsin fraction or either acetonitrile wash fraction and inputting the resulting peaks into a software such as Mascot22. Once the protein sequence has been determined, calculating the mass of the tryptic N-terminal lipopeptide significantly aids in choosing which ion to fragment by MS/MS.
Additional sample heterogeneity can result from varying acyl chain lengths on lipoproteins within a population and their N-terminal modifications, both depending on the source bacteria6. While variation in chain length makes for distinctive peak clusters on parent spectra, characterized by 14 Da increments corresponding to methylene groups (-CH2-), it can split the overall lipopeptide signal and decrease sensitivity. It is also likely that the different lipoprotein forms influence enrichment and fragmentation, though we have successfully identified triacylated, diacylated, and lyso-form lipoproteins using the described protocol. Similarly, the varying peptide moieties of lipoproteins add another level of complexity to analyses, regardless of their N-terminal structure, as a very hydrophilic amino acid composition may prevent lipopeptide partitioning into the organic chloroform phase using other extraction methods8. Transfer of the lipoprotein to nitrocellulose would assist in studying such lipopeptides, as all elution fractions can be analyzed in this method.
Drawing from previous literature involving triacylglycerides and phospholipids23, intentional sodium adduct formation can be used to promote more informative fragmentation via formation of the (N-acyl)-dehydroalanyl peptide ion. These characteristic ions are key to assigning structure, since the acylation state of the N-terminus is isolated from the acyl substitutions on the glyceryl moiety. Sodium adduct formation provides a convenient alternate option to sulfur oxidation with hydrogen peroxide, which has been shown to also promote N-acyl-dehydroalanyl peptide fragmentation6,8. Although sodium adducts may show an overall suppression of the parent ion signal, the specific increase in fragmentation towards dehydroalanyl ions compensates for diminished parent ion intensity. It may be worth exploring other matrices and fragmentation of additional adducts to elicit the most informative spectra.
This protocol improves on the traditional TX-114 phase partitioning method for lipoprotein enrichment, while transfer of PAGE-separated lipoproteins to nitrocellulose membrane facilitates trypsin digestion and elution of lipopeptides. MALDI-TOF MS analysis on the total tryptic or wash fractions allows for protein identification in tandem with N-terminal structural determination by MS/MS in a single experiment. Optional sodium adduct formation highlights differences at the lipopeptide N-terminus by promoting more informative ion fragmentation. The ability to routinely purify lipoproteins and characterize their N-termini will enable extensive studies on how novel lipoprotein forms are made, their physiological role within the bacterial cell envelope, and how they are detected by the mammalian immune system.
Disclosures
The authors have nothing to disclose.
Acknowledgements
Research in the Meredith lab was supported by startup funds provided by the Eberly College of Science (Pennsylvania State University). We thank Dr. Tatiana Laremore for expert technical advice and access to equipment at the Penn State Proteomics and Mass Spectrometry Core Facility, University Park, PA, where mass spectrometric analyses were performed.
Materials
| Materials | |||
| 0.01 mm Zirconia/silica beads | BioSpec Products | 110791012 | |
| Acetic acid | EMD | AX0073-9 | |
| Acetone | EMD | AX0116-6 | |
| Acetonitrile | EMD | AX0142-6 | |
| Ammonium bicarbonate | Fluka Analytical | 09830 | |
| BioTrace NT Nitrocellulose | PALL Life Sciences | 66485 | |
| Bovine serum albumin (BSA) digest standard | Protea | PS-204-1 | |
| Chloroform | Acros Organics | 423550025 | |
| Ethylenediaminetetraacetic acid (EDTA) | Fisher Scientific | BP118 | |
| HPLC Grade water | EMD | WX0008-1 | |
| Lysozyme | Fisher Scientific | BP535-1 | |
| Methanol | Sigma-Aldrich | 34860 | |
| Phenylmethyl sulfonyl fluoride (PMSF) | Amresco | 0754 | |
| Pierce trypsin protease | Thermo Scientific | 90057 | |
| Ponceau S | Acros Organics | 161470100 | |
| Protein LoBind Tube 0.5mL | Eppendorf | 022431064 | |
| Sodium bicarbonate | Sigma-Aldrich | 792519 | |
| Sodium chloride | Macron Fine Chemicals | 7581-06 | |
| Trifluoroacetic acid (TFA) | Sigma-Aldrich | 299537 | |
| Tris-hydrochloride (HCl) | Fisher Scientific | BP152 | |
| Triton X-114 | Sigma-Aldrich | 93422 | |
| Tryptic soy broth (TSB) | BD | 211822 | |
| α-cyano-4-hydroxycinnamic acid (MS Grade) | Sigma-Aldrich | C8982 | |
| Name | Company | Catalog Number | Comments |
| Equipment | |||
| MagNALyser | Roche | ||
| Trans-Blot Turbo Transfer System | Bio-Rad | ||
| MALDI-TOF target | Bruker Daltonics | ||
| Ultraflextreme MALDI-TOF-TOF | Bruker Daltonics | Equipped with a 355 nm frequency-tripled Nd:YAG smartbeam-II laser |
References
- Babu, M. M., et al. A database of bacterial lipoproteins (DOLOP) with functional assignments to predicted lipoproteins. J Bacteriol. 188 (8), 2761-2773 (2006).
- Narita, S., Tokuda, H. Bacterial lipoproteins; biogenesis, sorting and quality control. Biochim Biophys Acta. 1862 (11), 1414-1423 (2016).
- Kovacs-Simon, A., Titball, R. W., Michell, S. L. Lipoproteins of bacterial pathogens. Infect Immun. 79 (2), 548-561 (2011).
- Nguyen, M. T., Götz, F. Lipoproteins of Gram-Positive Bacteria: Key Players in the Immune Response and Virulence. Microbiol Mol Biol Rev. 80 (3), 891-903 (2016).
- Nakayama, H., Kurokawa, K., Lee, B. L. Lipoproteins in bacteria: structures and biosynthetic pathways. FEBS J. 279 (23), 4247-4268 (2012).
- Kurokawa, K., et al. Novel bacterial lipoprotein structures conserved in low-GC content Gram-positive bacteria are recognized by Toll-like receptor 2. J Biol Chem. 287 (16), 13170-13181 (2012).
- Kurokawa, K., et al. The Triacylated ATP Binding Cluster Transporter Substrate-binding Lipoprotein of Staphylococcus aureus Functions as a Native Ligand for Toll-like Receptor 2. J Biol Chem. 284 (13), 8406-8411 (2009).
- Asanuma, M., et al. Structural evidence of α-aminoacylated lipoproteins of Staphylococcus aureus. FEBS J. 278 (5), 716-728 (2011).
- Armbruster, K. M., Meredith, T. C. Identification of the Lyso-Form N-Acyl Intramolecular Transferase in Low-GC Firmicutes. J Bacteriol. 199 (11), e00099-e00017 (2017).
- Serebryakova, M. V., Demina, I. A., Galyamina, M. A., Kondratov, I. G., Ladygina, V. G., Govorun, V. M. The acylation state of surface lipoproteins of Mollicute Acholeplasma laidlawii. J Biol Chem. 286 (26), 22769-22776 (2011).
- Feng, S. H., Lo, S. C. Induced mouse spleen B-cell proliferation and secretion of immunoglobulin by lipid-associated membrane proteins of Mycoplasma fermentans incognitus and Mycoplasma penetrans. Infect Immun. 62 (9), 3916-3921 (1994).
- Feng, S. H., Lo, S. C. Lipid extract of Mycoplasma penetrans proteinase K-digested lipid-associated membrane proteins rapidly activates NF-kappaB and activator protein 1. Infect Immun. 67 (6), 2951-2956 (1999).
- Luque-Garcia, J. L., Zhou, G., Spellman, D. S., Sun, T. -. T., Neubert, T. A. Analysis of electroblotted proteins by mass spectrometry: protein identification after Western blotting. Mol Cell Proteomics. 7 (2), 308-314 (2008).
- Luque-Garcia, J. L., Zhou, G., Sun, T. -. T., Neubert, T. A. Use of Nitrocellulose Membranes for Protein Characterization by Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry. Anal Chem. 78 (14), 5102-5108 (2006).
- Kurokawa, K., et al. Environment-mediated accumulation of diacyl lipoproteins over their triacyl counterparts in Staphylococcus aureus. J Bacteriol. 194 (13), 3299-3306 (2012).
- Laemmli, U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 227 (5259), 680-685 (1970).
- Schӓgger, H. Tricine-SDS-PAGE. Nat Protoc. 1, 16-22 (2006).
- Gravel, P. Protein Blotting by the Semidry Method. The Protein Protocols Handbook. , 321-334 (2002).
- Webster, J., Oxley, D. Protein Identification by Peptide Mass Fingerprinting using MALDI-TOF Mass Spectrometry. The Protein Protocols Handbook. , 1117-1129 (2009).
- Wilkins, M. R., et al. Detailed peptide characterization using PEPTIDEMASS – a World-Wide-Web-accessible tool. Electrophoresis. 18 (3-4), 403-408 (1997).
- Gasteiger, E., et al. Protein Identification and Analysis Tools on the ExPASy Server. The Proteomics Protocols Handbook. , 571-607 (2005).
- Perkins, D. N., Pappin, D. J. C., Creasy, D. M., Cottrell, J. S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis. 20 (18), 3551-3567 (1999).
- Al-Saad, K. A., Zabrouskov, V., Siems, W. F., Knowles, N. R., Hannan, R. M., Hill, H. H. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry of lipids: ionization and prompt fragmentation patterns. Rapid Commun Mass Spectrom. 17 (1), 87-96 (2003).

