Chemical Conjugation of a Purified DEC-205-Directed Antibody with Full-Length Protein for Targeting Mouse Dendritic Cells In Vitro and In Vivo
Summary
We describe a protocol for the chemical conjugation of the model antigen ovalbumin to an endocytosis receptor-specific antibody for in vivo dendritic cell targeting. The protocol includes purification of the antibody, chemical conjugation of the antigen, as well as purification of the conjugate and the verification of efficient conjugation.
Abstract
Targeted antigen delivery to cross-presenting dendritic cells (DC) in vivo efficiently induces T effector cell responses and displays a valuable approach in vaccine design. Antigen is delivered to DC via antibodies specific for endocytosis receptors such as DEC-205 that induce uptake, processing, and MHC class I- and II-presentation.
Efficient and reliable conjugation of the desired antigen to a suitable antibody is a critical step in DC targeting and among other factors depends on the format of the antigen. Chemical conjugation of full-length protein to purified antibodies is one possible strategy. In the past, we have successfully established cross-linking of the model antigen ovalbumin (OVA) and a DEC-205-specific IgG2a antibody (αDEC-205) for in vivo DC targeting studies in mice. The first step of the protocol is the purification of the antibody from the supernatant of the NLDC (non-lymphoid dendritic cells)-145 hybridoma by affinity chromatography. The purified antibody is activated for chemical conjugation by sulfo-SMCC (sulfosuccinimidyl 4-[N-maleimidomethyl] cyclohexane-1-carboxylate) while at the same time the sulfhydryl-groups of the OVA protein are exposed through incubation with TCEP-HCl (tris (2-carboxyethyl) phosphine hydrochloride). Excess TCEP-HCl and sulfo-SMCC are removed and the antigen is mixed with the activated antibody for overnight coupling. The resulting αDEC-205/OVA conjugate is concentrated and freed from unbound OVA. Successful conjugation of OVA to αDEC-205 is verified by western blot analysis and enzyme-linked immunosorbent assay (ELISA).
We have successfully used chemically crosslinked αDEC-205/OVA to induce cytotoxic T cell responses in the liver and to compare different adjuvants for their potential in inducing humoral and cellular immunity following in vivo targeting of DEC-205+ DC. Beyond that, such chemically coupled antibody/antigen conjugates offer valuable tools for the efficient induction of vaccine responses to tumor antigens and have been proven to be superior to classical immunization approaches regarding the prevention and therapy of various types of tumors.
Introduction
Dendritic cells (DC) are central players of the immune system. They are a diverse group of cells specialized in antigen-presentation and their major function is to bridge innate and adaptive immunity1,2. Importantly, DC not only play an important role in efficient and specific pathogen-directed responses but are also involved in many aspects of antitumor immunity1,3.
Due to their exclusive role in host immunity, DC came into focus as target cells for vaccination4. One approach is to target antigens to DC in vivo to induce antigen-specific immune responses and over the last years, a large number of studies have been dedicated to defining suitable receptors and targeting strategies1,4. One example is the C-type lectin receptor DEC-205, which can be targeted by DEC-205-specific antibodies to induce endocytosis. Importantly, DEC-205 targeting in the combination with suitable adjuvants has been shown to efficiently induce long-lived and protective CD4+ and CD8+ T cells, as well as antibody responses, also against tumor antigens3,5,6,7,8,9.
There are a number of studies showing conjugated antigens targeted to DC to be superior to free un-conjugated antigen3,5,10,11,12. This makes the conjugation of the antigen to the respective DC targeting moiety a central step in DC targeting approaches. In the case of DC targeting via antibodies or antibody fragments, antigens can be either chemically or genetically linked and either strategy provides its own (dis)advantages1. On the one hand, in genetically engineered antibody-antigen constructs there is a control over the antigen dose as well as the location providing superior comparability between lots1. At the same time however, chemical conjugation needs less preparation and provides more flexibility especially when attempting to test and compare different antigens and/or vaccination strategies in experimental and pre-clinical models.
Here, we present a protocol for the efficient and reliable chemical conjugation of ovalbumin (OVA) as a model protein antigen to a DEC-205-specific IgG2a antibody (αDEC-205) suitable for in vivo DC targeting in mice. First, αDEC-205 is purified from NLDC-145 hybridoma cells13. For chemical conjugation, the heterobifunctional crosslinker sulfosuccinimidyl 4-[N-maleimidomethyl] cyclohexane-1-carboxylate (sulfo-SMCC), which contains NHS (N-hydroxysuccinimide) ester and maleimide groups, is used, allowing covalent conjugation of amine- and sulfhydryl-containing molecules. Specifically, the primary amines of the antibody initially react with sulfo-SMCC and the resulting maleimide-activated αDEC-205 then reacts with the sulfhydryl-containing OVA protein reduced through TCEP-HCl (Tris(2-carboxyethyl) phosphine hydrochloride). The final product is chemically conjugated αDEC-205/OVA (Figure 1). Beyond chemical conjugation itself, our protocol describes removal of excess OVA from the conjugate as well as the verification of successful conjugation through western blot analysis and a specific enzyme-linked immunosorbent assay. We have successfully employed this approach in the past to chemically conjugate OVA and other proteins or immunogenic peptides to αDEC-205. We demonstrate efficient binding to CD11c+ cells in vitro as well as the efficient induction of cellular and humoral immunity in vivo.
Certainly, there are drawbacks to this method such as in lot-to-lot comparability and in the exact dosing of the antigen within the final conjugate. Nevertheless, chemical conjugation provides experimental flexibility in the choice of the antibody and the protein antigen as compared to genetically engineered constructs. Therefore, we believe this approach is especially valuable in evaluating different antigens for their efficiency in DC targeting in pre-clinical mouse models, importantly also in the context of specific antitumor immune responses.
Protocol
All of the described animal experiments were approved by the local government agency (Niedersächsisches Landesamt für Verbraucherschutz und Lebensmittelsicherheit; file number 33.12-42502-04-10/0108) and were performed according to the national and institutional guidelines.
1. Production of αDEC-205 from the hybridoma cell line NLDC-145
- For antibody production, thaw cryopreserved NLDC-145 cells producing αDEC-205 at 37 °C in a water bath. Expand the cells at 37 °C and 5% CO2. Two 75 cm2 bottles will be needed to proceed to antibody production. Cell culture procedures should be performed in a safety cabinet to ensure safe working conditions and prevent contamination of the cultures.
- Resuspend 1 mL of thawed cells (1 x 106 – 5 x 106 cells/mL) in 9 mL of ISF-1 medium supplemented with 1% penicillin/streptomycin (pre-warmed to 37 °C) into a cell culture flask (25 cm2). Place the flask horizontally in a cell culture incubator at 37 °C, 5% CO2.
- Culture the cells at 37 °C and 5% CO2, until 70% confluence. This should normally be achieved after 24 – 48 h.
- Once the cells are 70% confluent, transfer the complete NLDC-145 cell suspension (10 mL) into a 15 mL conical centrifuge tube using a pipette controller with a pipette in a volume range from 1-10 mL. Pellet the cells by centrifugation at 250 x g for 10 min at room temperature.
- Pre-warm ISF-1 medium to 37 °C in a water bath and resuspend the pellet in 12 mL of ISF-1 medium supplemented with 1% penicillin/streptomycin. Transfer the resuspended cell suspension into a fresh cell culture flask (75 cm2).
- Culture and expand the cells in ISF-1 medium supplemented with 1% penicillin/streptomycin at 37 °C and 5% CO2, until 70% confluent and 99% viable. This should normally be achieved after 48 – 72 h.
- Split the cells to two 75 cm2 flasks. To do this first flush the cell culture flask bottom/culture surface with the cell suspension to remove all NLDC-145 cells from the surface. Transfer 6 mL of the NLDC-145 cell suspension each into one of two fresh 75 cm2 bottles and add pre-warmed ISF-1 medium supplemented with 1% penicillin/streptomycin up to 12 mL.
NOTE: Do not renew the cell culture medium as the NLDC-145 cells will have conditioned the medium. Transfer the cells together with their medium and fill the culture up to the desired volume with fresh medium. This is crucial for viability and maximum antibody production by the NLDC-145 cells. - Expand the cell cultures at 37 °C and 5% CO2, until about 70% confluent which is generally achieved after 48 – 72 h.
- Once the cells are 70% confluent, transfer 10 mL of the expanded NLDC-145 cell suspension from each of the 75 cm2 bottles into one PETG (polyethylene terephthalate glycol) roller bottle (1,050 cm2). For this, flush the cell culture flask bottom/culture surface with the cell suspension to remove all cells from the surface using a pipette controller and 10 mL pipette.
- Add 140 mL of ISF-1 medium supplemented with 1% penicillin/streptomycin (pre-warmed to 37 °C) directly out of the medium bottle to each of the NLDC-145 containing roller bottles filling them up to the 150 mL mark.
NOTE: See note in step 1.1.6. - Culture the roller bottles at 37 °C, 5% CO2 and 25 rounds/min for three days.
- Add 150 mL of ISF-1 medium supplemented with 1% penicillin/streptomycin (pre-warmed to 37 °C) to each of the NLDC-145 containing roller bottles filling them up to the 300 mL mark.
- Culture the roller bottles now containing 300 mL culture each at 37 °C, 5% CO2 and 25 rounds/min for another three days.
- Add another 100 mL of ISF-1 medium supplemented with 1% penicillin/streptomycin (pre-warmed to 37 °C) to each of the NLDC-145 containing roller bottles, thereby filling them up to the 400 mL mark.
- Culture the roller bottles now containing 400 mL culture each at 37 °C, 5% CO2 and 25 rounds/min for another seven days.
NOTE: During this week, the culture gets very dense (up to 95% density) and viability decreases (down to as little as 50%), enabling maximum antibody release.
- For the purification of αDEC-205 from the culture supernatant pour the NLDC-145 cell suspension (from both roller bottles) directly to 500 mL autoclaved centrifugation bottles.
NOTE: A total volume of 800 mL of culture should be collected.- Centrifuge the culture for 30 min at 8,600 x g and 4 °C to remove cells and debris.
- Collect the supernatants, discarding the pellets and pooling the supernatants in a sterile reagent bottle.
NOTE: Purification of αDEC-205 can be commenced immediately (step 2.) or the supernatant can be stored short-term at 4 °C.
2. Purification of the αDEC-205 antibody from the NLDC-145 cell supernatant
NOTE: From the NLDC-145 cell supernatant, αDEC-205 is purified using a protein G Sepharose column (reusable). The column dimensions are 15 mm x 74 mm and 5 mL protein G Sepharose are packed per column.
- For preparation and washing of the protein G Sepharose column, put an airtight rubber plug on the upper opening of the protein G Sepharose column. Puncture the rubber plug with two sterile cannulas (20 G x 1 1/2", 0.90 x 40 mm).
- Connect a 10 mL syringe to one of the two cannulas and a flexible silicon tube (approximately 100 cm long, 2.5 – 3 mm diameter) with a tubing connector to the second cannula.
NOTE: The syringe/rubber plug construction is reusable and provides a vacuum resulting in a continuous flow of the large volume of culture supernatant to the protein G Sepharose column. To this end, slightly pull back the plunger of the syringe to ensure continuous fluid flow in the following steps. - Wash the column with 50 mL of 0.1 M glacial acetic acid (pH 2) to remove potentially remaining antibody from any previous antibody purification. Put the end of the silicon tube in the 0.1 M glacial acetic acid (pH 2) filled reagent bottle. As a result of the induced vacuum, 50 mL of the 0.1 M glacial acetic acid (pH 2) dropwise run through the protein G Sepharose column.
NOTE: 0.1 M glacial acetic acid (pH 2) should be stored in a reagent bottle or freshly filled in a beaker. - Wash the column with 100-200 mL phosphate-buffered saline (PBS). Put the end of the silicon tube into a PBS filled reagent bottle or beaker. Let 100-200 mL PBS run dropwise through the protein G Sepharose column.
- Connect a 10 mL syringe to one of the two cannulas and a flexible silicon tube (approximately 100 cm long, 2.5 – 3 mm diameter) with a tubing connector to the second cannula.
- For antibody purification from the NLDC-145 supernatant, load 800 mL of the NLDC-145 supernatant (obtained from step 1.2.2.) onto the column. Put the end of the silicon tube into the NLDC-145 supernatant filled reagent bottle. Let 800 mL NLDC-145 supernatant run dropwise through the column.
- Wash the column with 500 mL of PBS. Put the end of the silicon tube in a PBS filled reagent bottle or beaker. Let 500 mL PBS run dropwise through the column.
- For elution, use 20 1.5 mL tubes and pipette 100 µL of 1.5 M Tris-HCl (pH 8.8) into each 1.5 mL tube. Remove the rubber plug from the column and pipette 1 mL of 0.1 M glycine (pH 3) to the upper chamber of the protein G Sepharose column to elute the antibody from the column. Collect the flow-through directly as eluate in one of the prepared 1.5 mL tubes.
- Repeat the elution step (2.2.2.) for all 20 tubes (1.5 mL).
- Determine the optical density of all elution fractions at 280 nm (OD280) using a spectrophotometer in order to identify the antibody-containing fractions.
NOTE: Use the first elution fraction as blank. - Pool all fractions with an OD280 greater than 0.5 (approximately 10 fractions).
- Store the protein G Sepharose column filled with 20% ethanol at 4 °C.
- Dialyze the pooled elutions against 1000 mL PBS (in a 2000 mL beaker) at 4 °C overnight using dialysis tubing with a molecular weight cut off (MWCO) of 12 – 14 kDa.
- Cut the dialysis tubing into pieces of 20 cm. Boil the dialysis tubing in 500-800 mL of 10 mM EDTA (pH 7.5) for 30 min in a beaker using a hot plate to remove contamination. Discard the 10 mM EDTA (pH 7.5) solution and boil the dialysis tubing in deionized water for 10 min.
NOTE: The dialysis tubing can be used directly or stored in 0.01% sodium azide (NaN3)/H2O solution at 4 °C until next usage. - Close the bottom of the dialysis tubing with an appropriate dialysis tubing closure/single-piece, hinged clamp and carefully pipette the antibody elution into the dialysis tubing. Close the top of the dialysis tubing with a second clamp.
- Fix the upper clamp of the dialysis tubing to a floating stand, put it together with a magnetic stir bar into the PBS filled beaker and place the beaker on a magnetic stirrer.
- Dialyze overnight stirring at 4 °C.
- Cut the dialysis tubing into pieces of 20 cm. Boil the dialysis tubing in 500-800 mL of 10 mM EDTA (pH 7.5) for 30 min in a beaker using a hot plate to remove contamination. Discard the 10 mM EDTA (pH 7.5) solution and boil the dialysis tubing in deionized water for 10 min.
- To increase the concentration of αDEC-205, load the complete dialysate to a centrifugal concentrator with 10 kDa MWCO. Open one clamp of the tubing and carefully pipette the complete dialysate out of the dialysis tubing into the centrifugal concentrator (10 kDa MWCO).
NOTE: Do not touch the concentrator bottom with the pipette tip.- Centrifuge for 30 min at 693 x g (2,000 rpm) and 4 °C.
- Load the centrifugal concentrator with 10 mL of PBS and centrifuge at 693 x g (2,000 rpm) and 4 °C until the final volume of antibody solution left is 1-1.5 mL.
NOTE: If necessary, repeat the centrifugation step of 2.4.1. to adjust to the desired amount. - Using a spectrophotometer, determine the optical density of the concentrated αDEC-205 solution at 280 nm (OD280). Use PBS as blank.
- Calculate the concentration of αDEC-205 using the following formula:
concentration [mg/mL] = OD280/1.4. - Filter the αDEC-205 solution using a 0.22 µm syringe filter unit.
NOTE: Purification of αDEC-205 from the NLDC-145 hybridoma cell supernatant can also be achieved by FPLC (fast protein liquid chromatography). Purified αDEC-205 can be stored at 4 °C or at -18 °C for long-term storage.
3. Chemical conjugation of OVA to αDEC-205
NOTE: A ratio of 0.5 mg OVA protein to 2.5 mg αDEC-205 (1:5) is required for optimal chemical conjugation. However, this ratio can vary for other proteins and antibodies and needs to be optimized for alternative conjugates. Reduction of the disulfide bonds of the OVA protein is performed through incubation with 30 mM TCEP-HCl, which exposes the sulfhydryl-groups for chemical conjugation to αDEC-205 and 240 µl of TCEP-HCl are needed in step 3.2. Both steps, TCEP-induced reduction of OVA (step 3.1.) and sulfo-SMCC activation of αDEC-205 (step 3.2.), should preferably be performed in parallel.
- Freshly prepare a 125 mM TCEP-HCl solution (pH 7.0). Weigh out the desired amount of TCEP-HCl and dissolve the TCEP-HCl in 0.9 M Tris base (pH 8.8). Use pH indicator strips to test the pH of the 125 mM TCEP-HCl solution (which should be neutral) and adjust the pH with Tris base (pH 8.8).
- Pipette 200 µL OVA protein solution (containing 0.5 mg OVA) into a 1.5 mL sterile tube. Add 240 µl 125 mM TCEP-HCl and 560 µL of sterile ultrapure water to the OVA protein using a pipette to a final concentration of 0.5 mg/mL OVA protein and 30 mM TCEP-HCl (OVA/TCEP-HCl).
NOTE: 2.5 mg EndoGrade OVA (lyophilized) is dissolved in 1 mL PBS resulting in a 2.5 mg/mL OVA solution. - Incubate the resulting OVA/TCEP-HCl at room temperature for 1.5 h.
NOTE: Do not extend this incubation step.
- Pipette 200 µL OVA protein solution (containing 0.5 mg OVA) into a 1.5 mL sterile tube. Add 240 µl 125 mM TCEP-HCl and 560 µL of sterile ultrapure water to the OVA protein using a pipette to a final concentration of 0.5 mg/mL OVA protein and 30 mM TCEP-HCl (OVA/TCEP-HCl).
- To activate αDEC-205 for conjugation, dissolve 2 mg sulfo-SMCC in 100 µL of ultrapure water.
NOTE: Sulfo-SMCC is susceptible to hydrolysis. Therefore, larger amounts of undissolved sulfo-SMCC should be handled rapidly or available 2 mg aliquots should be used.- Dilute αDEC-205 in PBS so that 2.5 mg are contained in 900 µL.
- Mix 2.5 mg of αDEC-205 (900 µL volume; obtained from step 3.2.1.) and 100 µL of sulfo-SMCC (obtained from step 3.2.) in a 1.5 mL tube, resulting in a total volume of 1 mL.
- Incubate the αDEC-205/sulfo-SMCC solution for 30 min at 37 °C and 550 rpm in a heating block.
- Following these incubations, excess sulfo-SMCC and TCEP-HCl are immediately removed from the solutions using desalting columns (MWCO 7 kDa; 5 mL column volume).
- Twist off the columns (MWCO 7 kDa) bottom closure, loosen the cap and place the column into a 15 mL conical tube.
- Centrifuge for 2 min at 1,000 x g at room temperature to remove the liquid.
- Place the column in a fresh tube and remove the cap. Slowly load the antibody/sulfo-SMCC and the OVA/TCEP-HCl, respectively, to the center of the compact resin bed of one column each.
- Centrifuge for 2 min at 1,000 x g at room temperature.
- Discard the columns after use. The solutions containing antibody and OVA primed for conjugation are in the tubes.
- Immediately mix both solutions by pipetting for conjugation of αDEC-205 and OVA.
- Incubate the resulting αDEC-205 and OVA mixture overnight at 4°C.
- Following conjugation, excess unbound OVA is removed from the solution and the coupled αDEC-205/OVA is concentrated using a centrifugal protein concentrator (MWCO 150 kDa).
- Pre-rinse the centrifugal protein concentrator (MWCO 150 kDa) by pipetting 12 mL of PBS on the column and centrifuging for 2 min at 2,000 x g at room temperature.
NOTE: If necessary, repeat the centrifugation step (3.4.1.) until a volume of about 5 mL has passed through the column. - Before loading the αDEC-205/OVA onto the centrifugal protein concentrator, save a 20 µL sample of the un-concentrated αDEC-205/OVA for western blot analysis. Store this aliquot at 4 °C until analysis.
- Load the αDEC-205/OVA onto the centrifugal protein concentrator by pipetting.
NOTE: Avoid any contact with the bed of the upper chamber of the centrifugal concentrator. - Fill the concentrator to 15 mL with PBS and centrifuge the concentrator for 5 min at 2,000 x g at room temperature.
- Save a sample of the flow-through (flow-through I) for western blot analysis and discard remaining flow-through.
- Fill the concentrator to 10 mL with PBS and centrifuge the concentrator for at least 8 min at 2,000 x g at room temperature.
- Save a sample for the second flow-through (flow-through II) for western blot analysis and discard the remaining flow-through.
- Once the desired enrichment is achieved (around 1.5 mL of the αDEC-205/OVA solution should be left in the upper chamber) gently aspirate the concentrated sample.
NOTE: If too much fluid is left in the upper chamber, centrifugation can be repeated but should be kept as short as possible.
- Pre-rinse the centrifugal protein concentrator (MWCO 150 kDa) by pipetting 12 mL of PBS on the column and centrifuging for 2 min at 2,000 x g at room temperature.
- Determine the protein concentration of the resulting αDEC-205/OVA using a microvolume spectrophotometer. Use PBS as blank.
- Filter the αDEC-205/OVA using a 0.22 µm syringe filter unit.
NOTE: For later analysis and in vivo experiments, αDEC-205/OVA can be stored at 4 °C or -18 °C.
4. Verification of the chemical conjugation by western blot
NOTE: For verification of successful chemical conjugation, western blot analysis detecting either OVA (4.2) or αDEC-205 (4.10) is performed. Detection of OVA (4.2.) or αDEC-205 (4.10.) should be performed in parallel. An orbital platform shaker should preferably be used for all incubation steps of the western blot membranes to allow uniform distribution of the respective solutions.
- Prepare standard 10% SDS (sodium dodecyl sulfate) gels for SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis).
- Prepare the samples for SDS-PAGE to detect conjugated and un-conjugated OVA and run gel electrophoresis.
- Dilute different amounts of coupled αDEC-205/OVA (e.g., 200, 100 and 50 ng), of pure OVA (e.g., 80, 70, 60, 50, 40 and 30 ng), an aliquot of un-conjugated αDEC-205 (e.g., 100 and 50 ng), a sample of un-concentrated αDEC-205/OVA conjugate from step 3.4.2. (e.g., 125 ng) and an aliquot of flow-through I and II (steps 3.4.5. and 3.4.7., respectively; optional) in 4 x non-reducing SDS sample buffer.
NOTE: Different concentrations of conjugate and protein are loaded to ensure that both free OVA as well the conjugate can be detected on the same blot. - Denature the samples at 65 °C for 10 min and 550 rpm in a heating block.
- Dilute different amounts of coupled αDEC-205/OVA (e.g., 200, 100 and 50 ng), of pure OVA (e.g., 80, 70, 60, 50, 40 and 30 ng), an aliquot of un-conjugated αDEC-205 (e.g., 100 and 50 ng), a sample of un-concentrated αDEC-205/OVA conjugate from step 3.4.2. (e.g., 125 ng) and an aliquot of flow-through I and II (steps 3.4.5. and 3.4.7., respectively; optional) in 4 x non-reducing SDS sample buffer.
- Load the samples and a protein standard to an SDS gel and run SDS-PAGE.
- Perform standard blotting of the proteins from the SDS gel to a methanol-activated PVDF (polyvinylidene difluoride) membrane (75 min, 125 mA).
- Following blotting, put the membrane in an appropriate dish and block the membrane by pipetting 25 mL of 4% meal replacement shake powder/TBS-T (Tris-buffered saline/0.1% Tween 20) blocking buffer into the dish containing the membrane.
- Incubate the membrane in the blocking solution for 60 min at room temperature or overnight at 4 °C.
- Discard the blocking solution and stain the membrane with the rabbit αOVA primary antibody in 2% meal replacement shake powder/TBS-T antibody buffer (dilution 1:3,000) by pipetting 15 mL of primary antibody solution to the membrane in the dish.
- Incubate the membrane for 45 min at room temperature or overnight at 4 °C (use the platform shaker).
- Discard the antibody solution and wash the membrane in the dish (use the platform shaker).
- Add 25 mL of TBS-T to the membrane and incubate for 5 min. Discard the solution.
- Add 25 mL of TBS-T/0.5 M NaCl to the membrane and incubate for 5 min. Discard the solution.
- Add 25 mL of TBS-T/0.5% Triton X-100 to the membrane and incubate for 5 min. Discard the solution.
- Add 25 mL of TBS-T to the membrane and incubate for 5 min. Discard the solution.
- Stain the membrane with the goat αrabbit-IgG-HRPO (horse radish peroxidase) secondary antibody in 2% meal replacement shake powder/TBS-T antibody buffer (dilution 1:2,000) by pipetting 15 mL of secondary antibody solution to the membrane in the dish.
- Incubate the membrane for 45 min at room temperature or overnight at 4 °C (on the platform shaker).
- Wash the membrane again as described in step 4.7.1.- 4.7.4. using the platform shaker. For this membrane, next carry on with step 4.16. (the following steps 4.10.- 4.15. (detection of αDEC-205) can be performed parallel to steps 4.2.- 4.9.).
- Prepare the samples for the detection of αDEC-205 for SDS-PAGE and run the gel electrophoresis.
- Dilute different amounts of αDEC-205/OVA (e.g., 200, 100 and 50 ng), of the un-conjugated αDEC-205 (e.g., 250, 125, 62.5 ng), of pure OVA (e.g., 80 and 70 ng), a sample of un-concentrated αDEC-205/OVA from step 3.4.2. (e.g., 125 ng) and an aliquot of flow-through I and II (steps 3.4.5. and 3.4.7., respectively; optional) in 4 x non-reducing SDS sample buffer.
- Denature the samples at 65 °C for 10 min and 550 rpm in a heating block.
- Load the samples and a protein standard to an SDS gel and run gel electrophoresis.
- Perform standard blotting of the proteins from the SDS gel to a methanol activated PVDF (polyvinylidene difluoride) membrane (75 min, 125 mA).
- Following blotting, put the membrane in an appropriate dish and block the membrane by pipetting 25 mL of 10% blocking buffer (milk powder/TBS-T) into the dish containing the membrane.
NOTE: The blocking solutions differ between αDEC-205 and OVA detection (step 4.5.).- Incubate the membrane in the blocking solution for 60 min at room temperature or overnight at 4 °C (using the platform shaker).
- Discard the blocking solution and stain the membrane with the goat αrat-IgG(H+L)-HRPO antibody in 5% milk powder/TBS-T antibody buffer (dilution 1:5,000) by pipetting 15 mL of antibody solution to the membrane in the dish.
NOTE: The antibody buffers differ between αDEC-205 detection and OVA detection (steps 4.6./ 4.8.).- Incubate the membrane for 45 min at room temperature or overnight at 4 °C (using the platform shaker).
- Wash the membrane thoroughly in the dish as described in steps 4.7.1.- 4.7.4. (use the platform shaker).
- Use an appropriate detection reagent to develop the HRP signal and detect the chemiluminescence in a dark room using x-ray film or via an imaging system.
5. Verification of the chemical conjugation by ELISA
- Perform ELISA for further verification of successful chemical conjugation resulting in αDEC-205/OVA.
- Coat an appropriate 96-well ELISA plate with 100 µL/well of 3 ng/µL rabbit αOVA antibody in coating buffer (0.1 M sodium bicarbonate (NaHCO3) pH 9.6 diluted in H2O).
- Incubate the plate overnight at 4 °C.
- Following coating, wash the plate three times with PBS, e.g., using an ELISA washer.
- Block the plate by pipetting 200 µL of blocking buffer (10% FCS in PBS) in each well of the plate and incubate the plate for 30 min at room temperature.
- Serially dilute αDEC-205/OVA (obtained from step 3.6.) 1:2 in blocking buffer (10% FCS in PBS) to obtain dilutions ranging from 6 µg/mL down to 93.8 ng/mL αDEC-205/OVA and add 100 µL/well of these decreasing amounts of αDEC-205/OVA to the wells.
- Incubate the plate for 1 h at room temperature.
- Wash the plate three times with PBS, e.g., using an ELISA washer.
- Add 100 µL of the goat αrat-IgG+IgM(H+L)-HRPO antibody (diluted to 1:2,000 in blocking buffer (10% FCS in PBS)) to each well of the plate.
- Incubate for 1 h at room temperature.
- Wash the plate three times using PBS, e.g. using an ELISA washer.
- Add 50 µL of HRPO-substrate to the wells. When observing a clear color reaction, stop the reaction through addition of 150 µl stopping solution (1M H2SO4) per well.
- After 5 min, read absorption at 450 nm using an ELISA reader.
Representative Results
Chemical conjugation of αDEC-205 to OVA protein using this protocol will typically allow efficient generation of αDEC-205/OVA for in vivo DC targeting approaches. There are different strategies to verify the technique itself and to test the functionality of the yielded conjugate. Western blot analysis and ELISA are used to verify successful conjugation and at the same time detect potentially left free OVA (Figure 2). In vitro binding studies (Figure 3) and in vivo immunizations (Figure 4) confirm binding of the conjugate to DEC-205 and targeting of DC.
Parallel western blot analysis is used to detect both the conjugated OVA (Figure 2A) as well as conjugated αDEC-205 (Figure 2B). Specifically, the positive signal for OVA at the level of the antibody's molecular weight in the blot confirms association of OVA and the antibody (Figure 2A). Furthermore, staining for OVA in the western blot analysis allows the detection of excess free OVA potentially still present next to the αDEC-205/OVA yielded in step 3.6., which is not the case for the blot shown (Figure 2A). In case large amounts of free OVA are detected, steps 3.4.1. to 3.4.8. of the protocol should be repeated. Complementary to the staining for OVA (Figure 2A), staining for αDEC-205 in western blot analysis verifies successful conjugation through an increase in the molecular weight between "naked" αDEC-205 and the conjugate as shown in Figure 2B.
Next to western blotting, also a specific ELISA allows verification of the successful conjugation of αDEC-205 to OVA. In contrast to the western blot analyses however, this ELISA does not allow the detection of free and un-conjugated αDEC-205 or OVA. Due to the assay setup (Figure 2C), a positive signal is only produced if conjugation was efficient. The positive association between the detected signal (absorption at 450 nm) and the analyzed amount of protein verifies the successful generation of αDEC-205/OVA through chemical conjugation as shown in Figure 2D. At the same time, the positive signal already yielded from 9.38 ng of the αDEC-205/OVA conjugate demonstrates the strong sensitivity of this method (Figure 2D). In case there is no increase in adsorption for increasing amounts of the conjugate, the conjugation was presumably not successful. In this case, also the western blot analyses would yield negative results, i.e., no detection of the conjugate in the blot stained for OVA and no increase in the molecular weight in the blot stained for αDEC-205.
While the western blot and ELISA assays are used to evaluate the conjugation and the removal of free antigen as such, subsequent functional analyses are needed to confirm binding to DEC-205 and targeting of DC. To this end, we have performed in vitro binding studies (Figure 3) and in vivo immunizations (Figure 4). For these experiments, female 6-8 week C57BL/6 and 8-12 week Balb/c mice were obtained from commercial sources or bred at the animal facility of the Helmholtz Centre for Infection Research (HZI) and were housed under specific pathogen-free conditions. Figure 3 demonstrates functional assays for the binding of a conjugate of αDEC-205 and the HCV Core protein (αDEC-205/Core) to CD11c+ cells in vitro. Flow-cytometry clearly showed αDEC-205/Core to efficiently bind bone-marrow derived CD11c+ cells (Figure 3A,B) as well as freshly isolated mouse CD11c+ splenocytes (data not shown). These assays demonstrate the chemical conjugation not to interfere with the binding capacity of αDEC-205. This is further confirmed by immunofluorescence analyses showing binding of αDEC-205/Core to MHC-II+ CD11c+ cells sorted from in vitro generated bone-marrow derived dendritic cells (BMDC) (Figure 3C).
In the past, we have shown the αDEC-205/OVA conjugates produced by the demonstrated protocol to efficiently induce OVA-specific immune responses in vivo in mice, confirming successful generation of the conjugate as well as functional targeting of DC (Figure 4)12,14. Specifically, subcutaneous vaccination with αDEC-205/OVA efficiently induced humoral and cellular OVA-specific immune responses. Importantly, in a recombinant adenovirus challenge model we detected antiviral CD8+ T cells capable of eliminating virus-infected hepatocytes, which has strong implications for vaccines directed at hepatotropic viruses12. Moreover, the highly effective induction of antigen-specific cytotoxic T cells underlines the potential of this approach for the in vivo priming of antitumor immunity. Also, we have successfully used αDEC-205/OVA to test and compare different adjuvants in the context of in vivo DC targeting14. In vaccination with αDEC-205/OVA together with the adjuvant combination Poly(I:C) (polyinosinic-polycytidylic acid) and CpG (synthetic oligodeoxynucleotides containing unmethylated CpG motifs) we observed generally (Figure 4A) and for some time-points significantly higher OVA-specific IgG levels as compared to vaccination with OVA alone (Figure 4B). Furthermore, αDEC-205/OVA efficiently induced OVA-specific CD4+ as well as CD8+ T cell responses (Figure 4C,D) and the αDEC-205/OVA-induced CD8+ T cell response significantly exceeded that induced by OVA alone (Figure 4D).
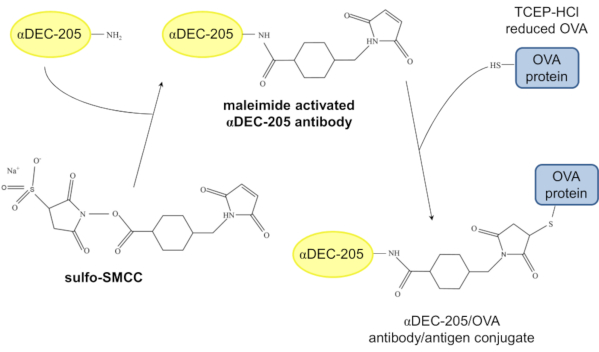
Figure 1: Model of the chemical conjugation of αDEC-205 and OVA. In a first step, the primary amine of αDEC-205 reacts with the NHS ester of the crosslinker sulfo-SMCC resulting in maleimide activated αDEC-205. Following reduction of the disulfide bonds of the OVA protein through incubation with TCEP-HCl, the maleimide activated αDEC-205 reacts with the TCEP-HCl-reduced OVA protein to form the αDEC-205/OVA antibody/antigen conjugate. Abbreviations: N-hydroxysuccinimide ester (NHS ester); sulfosuccinimidyl 4-[N-maleimidomethyl]cyclohexane-1-carboxylate (sulfo-SMCC); Tris(2-carboxysethyl)phosphine hydrochloride (TCEP-HCl). Please click here to view a larger version of this figure.
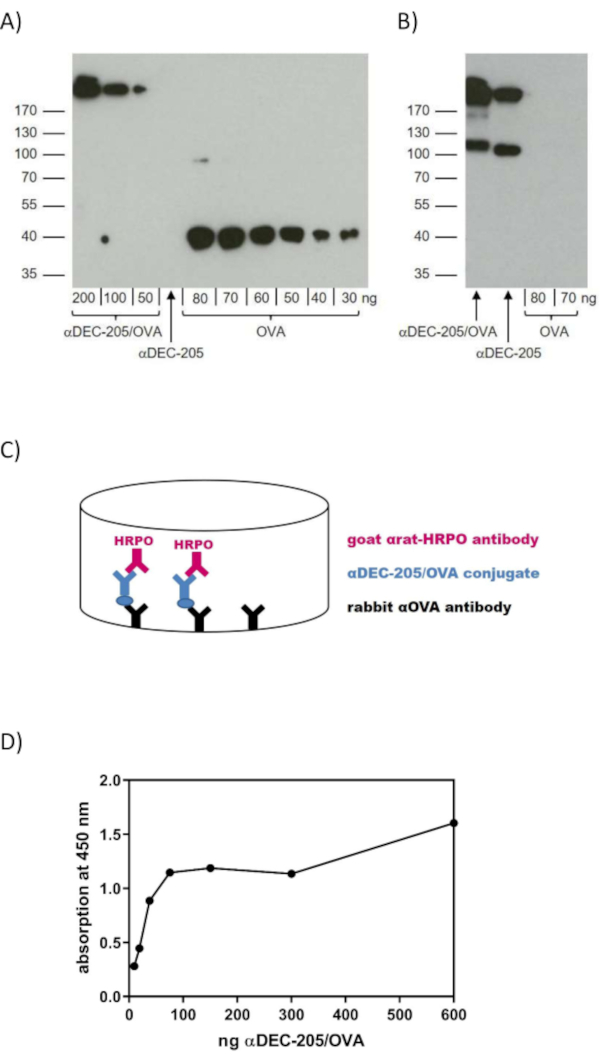
Figure 2: Verification of the chemically conjugated αDEC-205/OVA. To verify effective chemical conjugation of αDEC-205 and the OVA protein, western blot analysis (A,B) and ELISA (C,D) are performed. Samples of αDEC-205/OVA, αDEC-205 and different concentrations of OVA protein were subjected to SDS-PAGE (10%) and subsequent western blot analysis utilizing a rabbit αOVA primary antibody and a goat αrabbit-IgG-HRPO secondary antibody to detect OVA protein (A) or a goat αrat-IgG(H+L)-HRPO antibody to detect αDEC-205 (B). (C) Schematic representation of the ELISA for the verification of the αDEC-205/OVA conjugate. The rabbit αOVA coating antibody binds αDEC-205/OVA via the conjugated OVA. Goat αrat-IgG(H+L)-HRPO recognizes the αDEC-205 fraction of the bound conjugate and a positive signal thus confirms effective conjugation. (D) ELISA was performed as described in (C). Serially diluted amounts (1:2) of αDEC-205/OVA (600 ng to 9.38 ng) were analyzed. Data are shown as the mean of triplicates of a representative assay. Abbreviations: enzyme-linked immunosorbent assay (ELISA); horse radish peroxidase (HRPO); ovalbumin (OVA); sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Panels A and B have been modified from Volckmar et al.12. http://creativecommons.org/licenses/by/4.0/. Please click here to view a larger version of this figure.
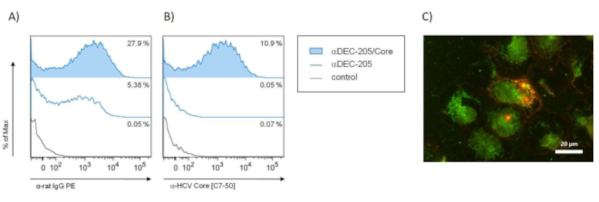
Figure 3: Binding of αDEC-205/Core to bone-marrow derived dendritic cells (BMDC) by flow-cytometry and immunofluorescence microscopy. To analyze the capacity of αDEC-205/Core to bind its target molecule DEC-205 on BMDCs in vitro, fluorescence-activated cell sorting (FACS) analysis (A,B) and immunofluorescence microscopy (C) were performed. In brief, bone marrow cells were isolated from the hind legs of female Balb/c mice (n=3) 8-12 weeks of age and cultured in RPMI medium supplemented with 1% penicillin/streptomycin, 1% glutamine, 0.25 mM mercaptoethanol and 5 ng/mL GM-CSF (granulocyte-macrophage colony stimulating factor). On day 6, the non-adherent BMDCs were carefully harvested and used for binding analysss. (A,B) In vitro generated BMDCs were incubated with 10 µg/mL αDEC-205/Core, αDEC-205 or medium (control) for 1 h at 4 °C followed by staining with APC-labeled αCD11c [clone HL3]. Detection of bound αDEC-205/Core on the surface of BMDCs was performed by additionally staining of the cells with either PE-labeled goat αrat (A) or mouse αHCV Core [clone C7-50] followed by secondary αmouse-IgG1-PE staining (B). Representative histograms show the PE signal and % PE-positive cells of the gated CD11c+ cells. (C) BMDCs generated in vitro from naïve Balb/c mice were sorted for MHC-II+ and CD11c+ cells and were incubated with 10 µg/mL αDEC-205/Core for 1 h at 4 °C. Cell-bound αDEC-205/Core was stained with Alexa 594-coupled αrat-IgG or with mouse αHCV Core [clone C7-50] and Alexa 488-coupled αmouse-IgG for 30 min at 4 °C after washing. The cells were visualized by immunofluorescence microscopy (scale bar = 20 µm). The binding capacity of αDEC-205/Core to BMDCs was confirmed by an overlay of both stainings (double positive = orange). Abbreviations: bone-marrow derived dendritic cells (BMDC); fluorescence-activated cell sorting (FACS); granulocyte-macrophage colony stimulating factor (GM-CSF); hepatitis C virus (HCV). Please click here to view a larger version of this figure.
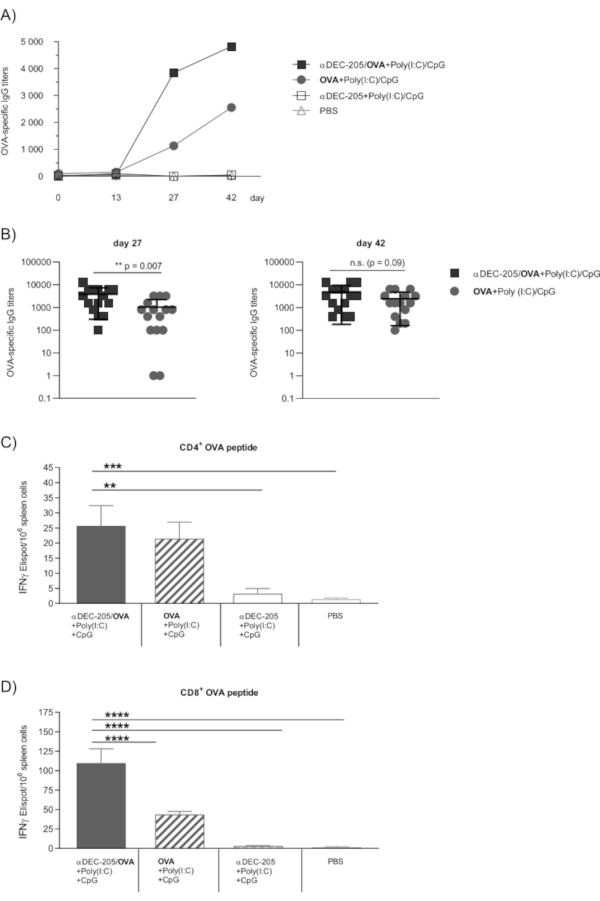
Figure 4: OVA-specific humoral and cellular immune responses following immunization with αDEC-205/OVA. The functionality of αDEC-205/OVA to target DC in vivo was proven through immunization experiments as previously published in Volckmar et al.12. Briefly, female 6-8 week old C57BL/6 mice (n=5) were subcutaneously immunized on days 0, 14 and 28 with 30 µg αDEC-205/OVA conjugate together with the adjuvants 50 µg Poly(I:C)/50 µg CpG, 30 µg αDEC-205 alone or 7 µg OVA protein alone in a total volume of 50 µL PBS per animal. Further controls were treated with PBS alone. (A,B) To monitor the humoral immune response, vaccinated mice were lightly anesthetized through isoflurane inhalation and blood samples were collected from the retro-orbital sinus on day 0, 13 and 27 and by cardiac puncture on day 42. Sera were prepared as described and assayed for the presence of OVA-specific IgG by ELISA12. Endpoint titers were expressed as the reciprocal value of the last serum dilution that yielded an absorbance two times above the values of negative controls. Results are compiled from three independent experiments. (A) Kinetic of OVA-specific total serum IgG titers shown as the group mean. (B) OVA-specific IgG titer on day 27 and day 42 shown for individual mice together with the group mean. Statistics: unpaired two-sided t-test. (C,D) The induction of cellular immune responses was analyzed by enzyme-linked immunosorbent spot (ELISPOT) assays using the murine IFNγ detection kit on day 42 as previously published12. Isolated splenocytes from immunized mice were pooled for the experimental groups and the number of IFNγ spot forming units/106 cells following stimulation with 5 mg/mL CD4+ (C) or CD8+ OVA peptide (D) was analyzed (OVA peptides: CD4323–339 (ISQAVHAAHAEINEAGR) and CD8257–264 (SIINFEKL)). Bars represent the mean ± SEM (n=5, triplicates from pooled samples). Statistics: one-way ANOVA with Dunnett's multiple comparisons test (**p<0.01, ***p<0.001, ****p<0.0001). Abbreviations: cytosine-phosphate-guanine oligonucleotide sequences (CpG), enzyme-linked immunosorbent assay (ELISA), enzyme-linked immunosorbent spot (ELISPOT), ovalbumin (OVA), phosphate-buffered saline (PBS), polyinosinic-polycytidylic acid (Poly(I:C)). This figure has been modified from Volckmar et al.12 http://creativecommons.org/licenses/by/4.0/. Please click here to view a larger version of this figure.
Discussion
Chemical conjugation of an endocytosis receptor-specific antibody and a protein antigen provides an efficient and, importantly, also flexible approach for in vivo DC targeting in pre-clinical mouse models. With our protocol we provide an efficient approach for the successful conjugation of the model antigen OVA to a DEC-205-specific IgG antibody.
In our protocol, αDEC-205 is purified from a hybridoma cellline and in the past, we have purified the antibody using protein G sepharose as described. Of note, we have also used FPLC to purify αDEC-205 in later studies and subsequent chemical conjugation was likewise efficient. The critical steps for efficient chemical conjugation are the priming of both the antibody and the protein. We have optimized these steps from different available protocols to finally perform incubation of the antibody with the crosslinker sulfo-SMCC and reduction of the protein through TCEP-HCl. Specifically, at this point the critical steps are the fresh preparation of TCEP-HCl (step 3.1.) and the duration of the incubation of TCEP-HCl with the protein, which should not be extended (step 3.1.2.). Moreover, the protein buffer used for the reduction of the disulfide bonds through TCEP-HCl is critical, as it can negatively affect reduction by TCEP-HCl. Also, it is important to immediately mix the activated antibody and the reduced protein (step 3.3.6.) to ensure optimal reaction conditions for the conjugation. In our hands, this approach yielded the most efficient and reliable results regarding conjugation. A further critical step for subsequent in vivo approaches is the removal of unbound OVA from the αDEC-205/OVA conjugate. To verify this step, we have optimized western blot analyses and recommend detection of both the conjugated αDEC-205 as well as the conjugated OVA protein (Figure 2A and B). In case unbound OVA is present in the final conjugate solution, blotting and detecting known concentrations of OVA (as in Figure 2A) will help estimate the amount of unbound OVA in relation to the total amount of protein loaded for the SDS-PAGE. If conjugation was inefficient, troubleshooting should address the antigen/antibody ratio used for the conjugation. In case conjugation was effective as detected by ELISA, but cannot be detected by western blot analysis, we have experienced the antibody concentrations in the western blot analyses (steps 4.6., 4.8., 4.14.) the most critical factor.
The main limitation of our approach is an at times varying conjugation efficiency in which there is no fixed correlation between the number of antibody molecules and the amount of coupled protein. Nevertheless, we believe and have experienced that the demonstrated protocol reliably allows subsequent in vivo studies of DC targeting that yield reproducible results. Furthermore, while in principle in our protocol both αDEC-205 as well as the OVA protein can be exchanged for alternative antibodies and antigens for in vivo studies of various interests, individual steps of the chemical conjugation will need to be newly optimized for the new components. This applies mainly to the optimal antibody/antigen ratio and can depend e.g. on the accessibility of reducible Cys residues. In our approaches, the optimal ratios resulting in successful conjugation reactions were 1:5 for OVA to αDEC-205 and 1.35:1 for the HCV proteins NS3 (nonstructural protein 3) and Core to αDEC-205. For each conjugate, negative effects of the crosslinker or the conjugation as such on the antigen binding capacity of the antibody need to be excluded. Of note, conjugation of immunogenic peptides instead of full-length proteins is less problematic in this regard. However, proteins provide higher antigen diversity in subsequent immunization. Genetic fusion approaches display an alternative to chemical conjugation and also have clear advantages1. Ultimately, the choice of the strategy to link antibodies and antigens for DC targeting will depend on resources, the research focus and anticipated applications of the conjugates. We believe the relative flexibility regarding antibodies and antigens displays a major advantage of chemical conjugation and we have indeed used the protocol described here for the efficient chemical conjugation of αDEC-205 to different proteins of the hepatitis C virus (HCV) (Figure 3 and data not shown).
Overall, we believe that chemical conjugation of endocytosis-receptor specific antibodies to protein antigens such as in αDEC-205/OVA displays a flexible and reliable tool to generate antibody/antigen conjugates of exceptional value in studying DC targeting approaches, also including those aiming as inducing antitumor immunity, especially in preclinical animal models.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors thank S. Prettin for expert technical assistance. This work was supported by a grant of the Helmholtz Association of German Research Centers (HGF) that was provided as part of the Helmholtz Alliance ''Immunotherapy of Cancers" (HCC_WP2b).
Materials
| antibody buffer 2 % | 2 % (w/v) Slim-Fast Chocolate powder in TBS-T | ||
| antibody buffer 5 % | 5 % milk powder (w/v) in TBS-T | ||
| blocking buffer (ELISA) | 10 % FBS in PBS | ||
| blocking buffer 4 % | 4 % (w/v) Slim-Fast Chocolate powder in TBS-T | ||
| blocking buffer 10 % | 10 % milk powder (w/v) in TBS-T | ||
| cell culture flask T25 | Greiner Bio-One | 690175 | we use standard CELLSTAR filter cap cell culture flasks; alternatively use suspension culture flask (690195 ) |
| cell culture flask T75 | Greiner Bio-One | 658175 | we use standard CELLSTAR filter cap cell culture flasks; alternatively use suspension culture flask (658195) |
| cell culture flask T175 | Greiner Bio-One | 661175 | we use standard CELLSTAR filter cap cell culture flasks; alternatively use suspension culture flask (661195) |
| centrifugal concentrator MWCO 10 kDa | Sartorius | VS2001 | Vivaspin 20 centrifugal concentrator |
| centrifugal protein concentrator MWCO 100 kDa, 5 – 20 ml | Thermo Fisher Scientific | 88532 | Pierce Protein Concentrator, PES 5 -20 ml; we use the Pierce Concentrator 150K MWCO 20mL (catalog number 89921), which is however no longer available |
| centrifuge bottles | Nalgene | 525-2314 | PPCO (polypropylene copolymer) with PP (polypropylene) screw closure, 500 ml; obtained from VWR, Germany |
| coating buffer (ELISA) | 0.1 M sodium bicarbonate (NaHCO3) in H2O (pH 9.6) | ||
| desalting columns MWCO 7 kDa | Thermo Fisher Scientific | 89891 | Thermo Scientific Zeba Spin Desalting Columns, 7K MWCO, 5 mL |
| detection reagent ELISA (HRPO substrate) | Sigma-Aldrich/Merck | T8665-100ML | 3,3′,5,5′-Tetramethylbenzidine (TMB) liquid substrate system |
| detection reagent western blot (HRPO substrate) | Roche/Merck | 12 015 200 01 | Lumi-Light Western Blotting Substrate (Roche) |
| dialysis tubing MWCO 12 – 14 kDa | SERVA Electrophoresis | 44110 | Visking dialysis tubing, 16 mm diameter |
| ELISA 96-well plate | Thermo Fisher Scientific | 442404 | MaxiSorp Nunc-Immuno Plate |
| fetal calf serum | PAN-BIOtech | P40-47500 | FBS Good forte |
| ISF-1 medium | Biochrom/bioswisstec | F 9061-01 | |
| milk powder | Carl Roth | T145.2 | powdered milk, blotting grade, low in fat; alternatively we have also used conventional skimmed milk powder from the supermarket |
| NLDC-145 hybridoma | ATCC | HB-290 | if not already at hand, the hybridoma cells can be acquired from ATCC |
| non-reducing SDS sample buffer 4 x | for 12 ml: 4 ml of 10 % SDS, 600 µl 0.5 M Tris-HCl (ph 6.8), 3.3 ml sterile H2O, 4 ml glycerine, 100 µl of 5 % Bromphenol Blue | ||
| ovalbumin | Hyglos (via BioVendor) | 321000 | EndoGrade OVA ultrapure with <0.1 EU/mg |
| Penicillin/Streptomycin | Thermo Fisher Scientific | 15140122 | Gibco Penicillin/Streptomycin 10.000 U/ml; alternatively Gibco Penicillin/Streptomycin 5.000 U/ml (15070-063) can be used |
| PETG polyethylene terephthalate glycol cell culture roller bottles | Nunc In Vitro | 734-2394 | standard PDL-coated, vented (1.2X), 1050 cm², 100 – 500 ml volume; obtained from VWR, Germany |
| pH indicator strips | Merck | 109535 | pH indicator strips 0-14 |
| polyclonal goat αrat-IgG(H+L)-HRPO (western blot) | Jackson ImmunoResearch | 112-035-062 | obtained from Dianova, Germany; used at 1:5000 for western blot |
| polyclonal goat αrat-IgG+IgM-HRPO antibody (ELISA) | Jackson ImmunoResearch | 112-035-068 | obtained from Dianova, Germany; used at 1:2000 for ELISA |
| polyclonal goat αrabbit-IgG-HRPO (western blot) | Jackson ImmunoResearch | 111-035-045 | obtained from Dianova, Germany; used at 1:2000 for western blot |
| polyclonal rabbit αOVA (ELISA) | Abcam | ab181688 | used at 3 ng/µl |
| polyclonal rabbit αOVA antibody (western blot) | OriGene | R1101 | used at 1:3,000 for western blot |
| Protein G Sepharose column | Merck/Millipore | P3296 | 5 ml Protein G Sepharose, Fast Flow are packed onto an empty column PD-10 (Merck, GE 17-0435-01) |
| protein standard | Thermo Fisher Scientific | 26616 | PageRuler Prestained Protein ladder 10 – 180 kDa |
| PVDF (polyvinylidene difluoride) membrane | Merck/Millipore | IPVH00010 | immobilon-P PVDF (polyvinylidene difluoride) membrane |
| rubber plug | Omnilab | 5230217 | DEUTSCH & NEUMANN rubber stoppers (lower Φ 17 mm; upper Φ 22 mm) |
| silicone tube | Omnilab | 5430925 | DEUTSCH & NEUMANN (inside Φ 1 mm; outer Φ 3 mm) |
| Slim-Fast | we have used regular Slim-Fast Chocolate freely available at the pharmacy as in this western blot approach it yielded better results than milk powder | ||
| stopping solution (ELISA) | 1M H2SO4 | ||
| sulfo-SMCC | Thermo Fisher Scientific | 22322 | Pierce Sulfo-SMCC Cross-Linker; alternatively use catalog number A39268 (10 x 2 mg) |
| syringe filter unit 0.22 µm | Merck/Millipore | SLGV033RS | Millex-GV Syringe Filter Unit, 0.22 µm, PVDF, 33 mm, gamma sterilized |
| syringe 10 ml | Omnilab | Disposable syringes Injekt® Solo B.Braun | |
| Sterican® cannulas | B. Braun | Sterican® G 20 x 1 1/2""; 0.90 x 40 mm; yellow | |
| TBS-T | Tris-buffered saline containing 0.1 % (v/v) Tween 20 | ||
| TCEP-HCl | Thermo Fisher Scientific | A35349 | |
| tubing connector | Omnilab | Kleinfeld miniature tubing connectors for silicone tube |
References
- Amon, L., Hatscher, L., Heger, L., Dudziak, D., Lehmann, C. H. K. Harnessing the complete repertoire of conventional dendritic cell functions for cancer immunotherapy. Pharmaceutics. 12 (7), 12070663 (2020).
- Banchereau, J., et al. Immunobiology of dendritic cells. Annual Review of Immunology. 18, 767-811 (2000).
- Wculek, S. K., et al. Dendritic cells in cancer immunology and immunotherapy. Nature Reviews: Immunology. 20 (1), 7-24 (2020).
- Kastenmuller, W., Kastenmuller, K., Kurts, C., Seder, R. A. Dendritic cell-targeted vaccines–hope or hype. Nature Reviews: Immunology. 14 (10), 705-711 (2014).
- Bonifaz, L. C., et al. In vivo targeting of antigens to maturing dendritic cells via the DEC-205 receptor improves T cell vaccination. Journal of Experimental Medicine. 199 (6), 815-824 (2004).
- Boscardin, S. B., et al. Antigen targeting to dendritic cells elicits long-lived T cell help for antibody responses. Journal of Experimental Medicine. 203 (3), 599-606 (2006).
- Hossain, M. K., Wall, K. A. Use of Dendritic Cell Receptors as Targets for Enhancing Anti-Cancer Immune Responses. Cancers. 11 (3), 11030418 (2019).
- Johnson, T. S., et al. Inhibition of melanoma growth by targeting of antigen to dendritic cells via an anti-DEC-205 single-chain fragment variable molecule. Clinical Cancer Research. 14 (24), 8169-8177 (2008).
- Trumpfheller, C., et al. The microbial mimic poly IC induces durable and protective CD4+ T cell immunity together with a dendritic cell targeted vaccine. Proceedings of the National Academy of Sciences of the United States of America. 105 (7), 2574-2579 (2008).
- Idoyaga, J., et al. Comparable T helper 1 (Th1) and CD8 T-cell immunity by targeting HIV gag p24 to CD8 dendritic cells within antibodies to Langerin, DEC205, and Clec9A. Proceedings of the National Academy of Sciences of the United States of America. 108 (6), 2384-2389 (2011).
- Sancho, D., et al. Tumor therapy in mice via antigen targeting to a novel, DC-restricted C-type lectin. Journal of Clinical Investigation. 118 (6), 2098-2110 (2008).
- Volckmar, J., et al. Targeted antigen delivery to dendritic cells elicits robust antiviral T cell-mediated immunity in the liver. Scientific Reports. 7, 43985 (2017).
- Kraal, G., Breel, M., Janse, M., Bruin, G. Langerhans’ cells, veiled cells, and interdigitating cells in the mouse recognized by a monoclonal antibody. Journal of Experimental Medicine. 163 (4), 981-997 (1986).
- Volckmar, J., et al. The STING activator c-di-AMP exerts superior adjuvant properties than the formulation poly(I:C)/CpG after subcutaneous vaccination with soluble protein antigen or DEC-205-mediated antigen targeting to dendritic cells. Vaccine. 37 (35), 4963-4974 (2019).

