Indel Detection following CRISPR/Cas9 Mutagenesis using High-resolution Melt Analysis in the Mosquito Aedes aegypti
Summary
This article details a protocol for rapid identification of indels induced by CRISPR/Cas9 and selection of mutant lines in the mosquito Aedes aegypti using high-resolution melt analysis.
Abstract
Mosquito gene editing has become routine in several laboratories with the establishment of systems such as transcription-activator-like effector nucleases (TALENs), zinc-finger nucleases (ZFNs), and homing endonucleases (HEs). More recently, clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) technology has offered an easier and cheaper alternative for precision genome engineering. Following nuclease action, DNA repair pathways will fix the broken DNA ends, often introducing indels. These out-of-frame mutations are then used for understanding gene function in the target organisms. A drawback, however, is that mutant individuals carry no dominant marker, making identification and tracking of mutant alleles challenging, especially at scales needed for many experiments.
High-resolution melt analysis (HRMA) is a simple method to identify variations in nucleic acid sequences and utilizes PCR melting curves to detect such variations. This post-PCR analysis method uses fluorescent double-stranded DNA-binding dyes with instrumentation that has temperature ramp control data capture capability and is easily scaled to 96-well plate formats. Described here is a simple workflow using HRMA for the rapid detection of CRISPR/Cas9-induced indels and the establishment of mutant lines in the mosquito Ae. aegypti. Critically, all steps can be performed with a small amount of leg tissue and do not require sacrificing the organism, allowing genetic crosses or phenotyping assays to be performed after genotyping.
Introduction
As vectors of pathogens such as dengue1, Zika2, and chikungunya3 viruses, as well as malarial parasites4, mosquitoes represent a significant public health threat to humans. For all these diseases, there is a substantial focus of transmission intervention on the control of mosquito vectors. Study of the genes important in, for example, permissiveness to pathogens, mosquito fitness, survivorship, reproduction, and resistance to insecticides is key for developing novel mosquito control strategies. For such purposes, genome editing in mosquitoes is becoming a common practice, especially with the development of technologies such as HEs, ZFNs, TALENs, and most recently, CRISPR with Cas9. The establishment of gene-edited strains typically involves backcrossing individuals carrying the desired mutations for a few generations to minimize off-target and founder (bottleneck) effects, followed by crossing heterozygous individuals to generate homozygous or trans-heterozygous lines. In the absence of a dominant marker, molecular genotyping is necessary in this process because, in many cases, no clear phenotypic traits can be detected for heterozygous mutants.
Although sequencing is the gold standard for genotypic characterization, performing this across hundreds, or possibly thousands of individuals, poses significant costs, labor, and time required to obtain results, which is especially critical for organisms with short lifespans such as mosquitoes. Commonly used alternatives are Surveyor nuclease assay5 (SNA), T7E1 assay6, and high resolution melt analysis (HRMA, reviewed in7). Both SNA and T7E1 use endonucleases that cleave only mismatched bases. When a mutated region of the heterozygous mutant genome is amplified, DNA fragments from mutant and wild-type alleles are annealed to make mismatched double-stranded DNA (dsDNA). SNA detects the presence of mismatches via digestion with a mismatch-specific endonuclease and simple agarose gel electrophoresis. Alternatively, HRMA uses the thermodynamic properties of dsDNA detected by dsDNA-binding fluorescent dyes, with the disassociation temperature of the dye varying based on the presence and type of mutation. HRMA has been used for the detection of single-nucleotide polymorphisms (SNPs)8, mutant genotyping of zebra fish9, microbiological applications10, and plant genetic research11, among others.
This paper describes HRMA, a simple method of molecular genotyping for mutant mosquitoes generated by CRISPR/Cas9 technology. The advantages of HRMA over alternative techniques include 1) flexibility, as it has been proven useful for various genes, a wide range of indel sizes, as well as the distinction between different indel sizes and heterozygous, homozygous, and trans-heterozygous differentiation12,13,14, 2) cost, as it is based on commonly used PCR reagents, and 3) time-saving, as it can be performed in just a few hours. In addition, the protocol uses a small body part (a leg) as a source of DNA, allowing the mosquito to survive the genotyping process, permitting the establishment and maintenance of mutant lines.
Protocol
1. Scanning for single nucleotide polymorphisms (SNPs), HRMA primer design, and primer validation
- SNP identification in wild-type laboratory colony mosquitoes
- Select the target exon to disrupt proper polypeptide translation.
NOTE: The target should be close to the start codon or amongst key residues required for protein function. The shorter the exon (e.g., ≤200 bases), the more difficult it is to target and analyze. Avoid editing close to the boundaries of an exon, as this forces one of the HRMA primers to either cross an intron or be in an intron. This is undesirable because SNP rates tend to be much greater in those regions. - Primer design
- Go to the NCBI Blast – Primer Blast website15, copy and paste the chosen exon in the box at the top of the page | choose the PCR product size (ensure it encompasses most of the exon) | choose the organism.
- Click on Advanced parameters | Opt (for PCR Product Tm) and add 60 (for an optimal temperature of 60 °C) | Get Primers. Keep the other parameters as default.
- Obtain genomic DNA (gDNA) from the wild-type laboratory colony. Set aside 10 mosquitoes, anesthetize them with CO2, and place them in a Petri dish on ice to keep them inactive. Set up 10 tubes containing 0.5 µL of the reagent for release of DNA from tissue and 20 µL of dilution buffer, both provided in the DNA release kit suggested in the Table of Materials.
- Remove one leg from a single mosquito using forceps and place it in a corresponding tube of a diluted solution of the DNA-release reagent (from step 1.1.3), completely submerging the leg in the solution. Repeat this step with the remaining mosquitoes, wiping the forceps with 70% ethanol before proceeding to the next one.
- Incubate the leg-containing solution at room temperature for 2-5 min, then at 98 °C for 2 min, and allow it to cool down while setting up the PCR reaction.
NOTE: Plates containing the released gDNA can be stored at -20 °C, and the protocol can be paused at this point. - Prepare 10 tubes containing 10 µL of 2x PCR Master Mix, primers to a 0.5 µM final concentration, and molecular grade water to a final volume of 20 µL, and transfer 1 µL of the diluted sample to each tube. Perform PCR following these cycling parameters: 98 °C 5 min, 40 cycles of 98 °C for 5 s, 60 °C 30 s, 72 °C 20 s per kb; final extension of 72 °C for 1 min.
- Purify the PCR products with an enzyme to degrade the residual PCR primers and dephosphorylate excess dNTPs or any column clean-up kit. Proceed with sequencing the samples.
- Analyze each electropherogram for the presence of double peaks/ambiguous bases, and adjust base calls manually in each sequence using the appropriate degenerate base code.
NOTE: This step must be performed before multiple sequence alignment as it is common for the base-calling software to select the more prominent peak as the "true" base call, giving the false impression of an absence of SNPs. - Perform a multiple sequence alignment using the alignment software, SeqMan Pro, listed in the Table of Materials or other open-source alignment software, such as ClustalW16 or T-Coffee17.
- Open the alignment software | click on Add Sequences | select the desired sequences and click on Add | once all sequences are chosen, click on Done.
- Click on Assemble to perform the alignment. To open the alignment, click on Contig 1 and analyze the alignment and identify the SNPs (Figure 1).
NOTE: As an alternative to steps 1.1.3-1.1.6, PCR can be performed from isolated genomic DNA obtained from bulk samples (derived from >10 individuals). Sequenced amplicons can be analyzed directly for the presence of SNPs appearing as double peaks in the electropherogram, though rare SNPs will be more difficult to detect.
- Design 3-5 single guide RNAs (sgRNAs), avoiding regions containing any SNP identified above, following the protocol described in18.
- Select the target exon to disrupt proper polypeptide translation.
- HRMA primer design
- Test the exon sequence for the possible formation of secondary structures during PCR using mFold19.
- Go to the UNAFold Web Server | click on mFold | on the dropdown menu, click on Applications | DNA Folding Form.
- Enter the sequence name in the box and paste the exon.
- Change the folding temperature to 60 °C; on the Ionic conditions, change [Mg++] to 1.5 and on Units switch to mM; click on Fold DNA.
- On Output, below Structure 1, click on pdf to open the Circular Structure Plots.
- Go to NCBI Blast – Primer Blast15 for primer design.
- Copy and paste the selected exon sequence determined by sequencing in step 1.1 (the sequence that contains the lowest number of SNPs or no SNPs) in the box at the top of the page.
- Use the symbol < > to mark and exclude sequences that contain SNPs, the target site, and regions with possible formation of secondary structure.
- Select the PCR product size to be between 80 and 150 bp and choose the organism.
NOTE: Larger fragment sizes can be successfully used (~300 bp). However, longer amplicons may decrease sensitivity between sequences differing in one or just a few base pairs. - Click on Get Primers. Select 2–3 pairs of primers to be tested (ideally, primer sites are ≥20–50 bp away from any CRISPR target sites).
- Test the exon sequence for the possible formation of secondary structures during PCR using mFold19.
- Primer validation
- Perform a gradient PCR using gDNA from a single individual.
- Prepare a master mix and remove one sample for the non-template control (NTC) in a separate tube. Add the template to the remaining master mix and aliquot into a 96-well plate.
- Follow the cycling parameters: 98 °C 30 s, 34 cycles of 98 °C for 10 s, 55–65 °C 30 s, 72 °C 15 s; final extension of 72 °C for 10 min.
- Generate thermal melt profiles following the parameters: denaturation step 95 °C for 1 min, annealing 60 °C for 1 min, melt curve detection between 75 °C and 95 °C in 0.2 °C increments, with a hold time of 10 s at each temperature.
NOTE: Only annealing temperatures with a single thermal melt profile should be used.
- Perform a gradient PCR using gDNA from a single individual.
- Proceed to generate the mutant lines with embryo injections as described in12.
2. Preparation of genomic DNA from mosquito legs
- Separate the G1 mosquitoes by sex at the pupal stage so that they do not mate before genotyping, and make sure that age-matched, wild-type control mosquitoes will be available to be used for references and backcrosses. Sex-separate them likewise.
- Gather the materials needed (Figure 2A) and prepare a 96 well PCR plate with 0.5 µL of the DNA-release reagent and 20 µL of dilution buffer (from the gDNA release kit) per individual to be genotyped (Figure 2B) and leave it on ice. Reserve two reactions for NTC.
- Label a tray for mosquito vials (Drosophila vials) so each well on the 96-plate corresponds to the respective mosquito vial in the tray.
- Anesthetize the G1 mosquitos with CO2 and place them in a glass Petri dish to keep them sedated (Figure 2C,D). Anesthetize and place 8 wild-type mosquitoes in a second Petri dish.
- Wipe a pair of tweezers with 70% ethanol and remove one of the mosquito hind legs (Figure 2E,F).
- Submerge the leg in the DNA-release reagent solution, place the mosquito in the corresponding vial, and close it with a sponge (Figure 2G,H).
- Wipe the tweezers again with 70% ethanol and proceed with removing the leg from the next mosquito. Repeat steps 2.5-2.7 until the 96-well plate is completed.
NOTE: It is important to wipe the tweezers with 70% ethanol. This minimizes DNA cross-contamination. - Seal the plate with an optical PCR plate seal (Figure 2I) and incubate the 96-well plate containing the legs at room temperature (RT) for 2-5 min and then at 98 °C for 2 min. Allow the plate to cool down to RT while preparing the PCR mix.
NOTE: The entire process typically takes 3-4 h. If the mosquitoes must be kept in the vials for more time than the expected duration (especially ≥1 day), place a small piece of raisin (or source of sugar and water) with each mosquito to ensure the survival of the mosquitoes during extended incubations.
3. HRMA
- Perform PCR.
- Prepare a master mix containing the following components per each reaction: 10 µL of 2x buffer (from the gDNA release kit), 0.5 µM of each primer, 1 µL of EvaGreen dye, 0.4 µL of polymerase (from gDNA release kit), and complete to 19 µL with molecular-grade water.
- Using a multichannel pipet, transfer 19 µL of the master mix into each well. Transfer 1 µL of the DNA release solution containing mosquito DNA prepared in section 2 to the plate (Figure 3A). Seal the plate with an optical PCR plate seal.
- Perform the PCR following the cycling parameters: 98 °C for 5 min, 39 cycles of 98 °C for 10 s, the chosen annealing temperature (72 °C) for 30 s; final extension 72 °C for 2 min.
- Generate thermal melt profiles following the parameters: denaturation step 95 °C for 1 min, annealing 60 °C for 1 min, melt curve detection between 75 °C and 95 °C in 0.2 °C increments, with a hold time of 10 s at each temperature. See the Supplemental Material and Supplemental Figure S1-S5 for a detailed description of the software setup for HRMA run using the CFX96 Real-Time System.
- Examine the melt profiles (Figure 3B). Assign wild-type control to the reference cluster.
NOTE: The software (see the Table of Materials) automatically normalizes the data and designates clusters with colors for different melt curves. - Mark the different clusters with corresponding colors on the 96-well template (Figure 3C).
- Select the individuals with curves of interest, remove them from the tubes, and backcross (Figure 3D). Blood-feed the mated females and collect G2 eggs.
4. Sequence verification by Sanger sequencing
- Purify the PCR product from the wells with selected mosquitoes (from the plate prepared in section 3.1), using an enzyme to degrade the residual PCR primers, and dephosphorylate excess dNTPs. Alternatively, use any column clean-up kit and proceed with sequencing the samples.
NOTE: Direct sequencing may be more challenging when the PCR product size is small (≤200 bp). In such cases, design primers to amplify larger fragments encompassing the target site (≥250 bp) and amplify by PCR using the DNA in the 96-well plate (step 2.2). - Analyze and identify the indels using trace viewer software (Figure 4). Alternatively, Poly Peak Parser software20 can help detect the mutation in heterozygous individuals.
- Go to the Poly Peak Parser website, select the sequence from the mutant individuals on the Browse menu, and copy and paste the reference sequence on the box.
NOTE: The alignment between the alternate allele and the reference will appear automatically on the right side of the screen.
- Go to the Poly Peak Parser website, select the sequence from the mutant individuals on the Browse menu, and copy and paste the reference sequence on the box.
Representative Results
Mosquitoes containing mutations in the genes AaeZIP11 (putative iron transporter21) and myo-fem (a female-biased myosin gene related to flight muscles13) were obtained using CRISPR/Cas9 technology, genotyped using HRMA, and sequence-verified (Figure 5). Figure 5A and Figure 5C show the normalized fluorescence intensity from the HRM curves from AaeZIP11 and myo-fem mutant samples, respectively, along with wild-type controls. Figure 4B and Figure 4D show the magnification of the difference curves (from the AaeZIP11 and myo-fem mutant samples, respectively) between the melt profiles of the different clusters assigned by the software after subtracting each curve from the wild-type reference. Heterozygous and homozygous mutant AaeZIP11 individuals were placed in different clusters and are easily distinguished from the wild-type controls (Figure 5B). Heterozygous mutant myo-fem individuals are also distinct from the controls (Figure 5D). Note that two clusters within the wild-type controls were present in both cases, most likely due to the presence of SNPs in the target region (Figure 5), highlighting the need to use multiple control samples to avoid categorizing controls as mutants when SNPs cannot be avoided.
Figure 4A shows the sequence analysis of the AaeZIP11 mutant. The electropherogram from AaeZIP11 heterozygous mutants indicates the nucleotide position where the indel occurred. This is represented by a shift from single to double peaks as polymorphic positions will show both nucleotides concomitantly (Figure 4A). The number of base pairs deleted or inserted was calculated by counting the single peaks at the end of the run (Figure 4B) as one of the DNA strands will be shorter or longer than the other by the number of base pairs deleted or inserted, respectively. It is recommended to use sequence-verified gDNA from the mutants as a reference for the identification of the heterozygotes to help with HRMA analyses for subsequent generations. Manual assignment for the curves may be needed when similar curves are automatically assigned to different clusters. This can be done by comparing the temperature-shifted curves and difference curves. Manual adjustment of the software might be needed to properly assign samples to clusters. HRMA results from AaeZIP11 and a mutant called Aeflightin show that individual sample analyses were needed to successfully categorize heterozygotes, homozygotes, and trans-heterozygotes. Initially, the automatic cluster assignment from the software could not make a proper distinction between the groups (Figure 6A, Figure 6C, Figure 6E, and Figure 6G). Each sample was then analyzed individually and assigned to the correct groups based on similarity to reference samples of heterozygotes, homozygotes, and trans-heterozygotes (previously verified by sequencing) (Figure 6B, Figure 6D, Figure 6F, and Figure 6H).

Figure 1: SNP identification. Schematic representation of multiple sequence alignment of AaeZIP11 fragment from wild-type. In red are the SNPs, and in green are the fragments free of SNPs; this SNP-free region is suggested for sgRNA and primer design. Abbreviations: SNP = single nucleotide polymorphism; sgRNA = single guide RNA; LVP = Liverpool strain. Please click here to view a larger version of this figure.
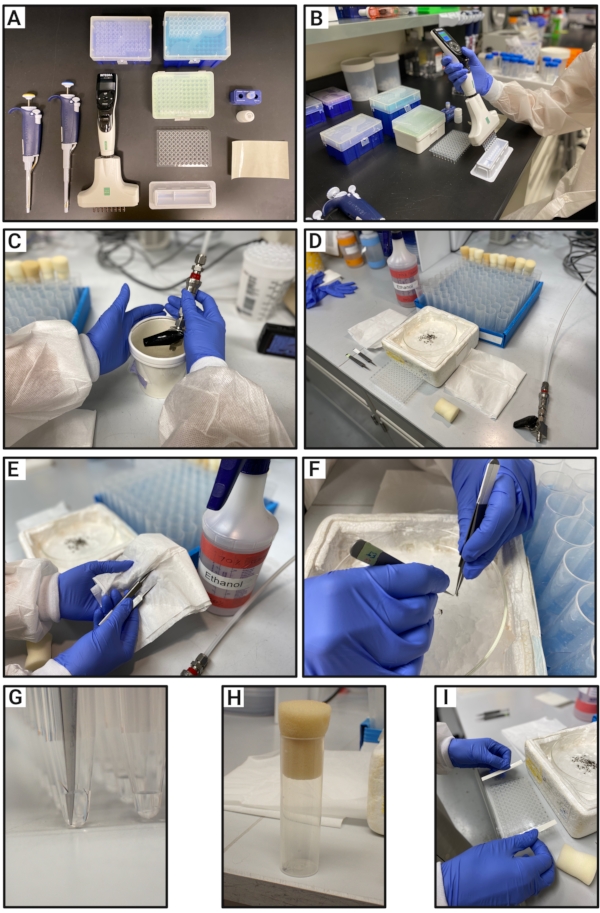

Figure 2: Experimental procedure for obtaining genomic DNA from mosquito legs for HRMA. (A) Materials for obtaining genomic DNA including pipettes, tips, PCR plate, optical seal, reservoir, the dilution buffer, and DNA release solution. (B) PCR plate preparation containing DNA release reagent and dilution buffer. (C) Mosquito anesthesia with CO2. (D) Wiping the tweezers with 70% ethanol to prevent contamination between samples. (E) Experimental setup including tweezers, rack for mosquito vials, Petri dish on top of an ice container with anesthetized mosquitoes, and the previously prepared PCR plate. (F) Removal of the mosquito leg. (G) Zoom view of the mosquito leg being submerged in the DNA release reagent solution. (H) Single mosquito placed in the vial. (I) Sealing the PCR plate containing the mosquito legs. Abbreviation: HRMA = high-resolution melt analysis. Please click here to view a larger version of this figure.
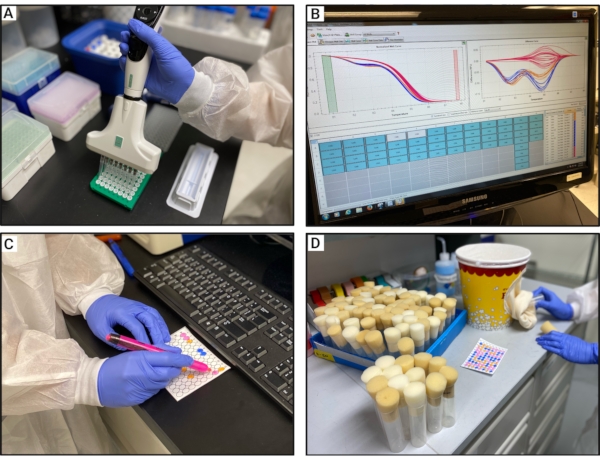
Figure 3: Experimental procedure for HRM analyses. (A) Transferring the released gDNA to a 96-well plate containing the PCR mix. (B) Visual inspection of the difference curves. (C) Marking each sample on the 96-well template with the same color as its respective difference curve color. (D) Backcrossing the individuals from the same cluster. Abbreviations: HRM = high-resolution melt; gDNA = genomic DNA. Please click here to view a larger version of this figure.
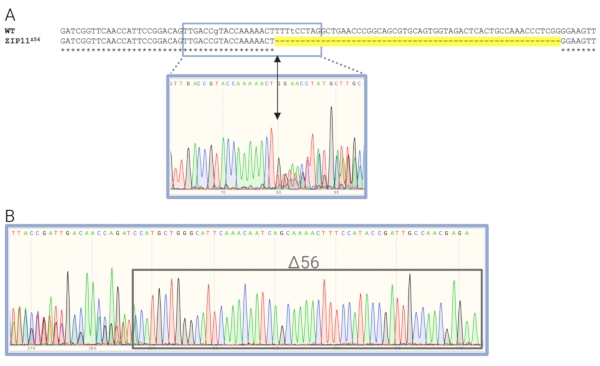
Figure 4: Sequence analysis of an AaeZIP11 mutant. (A) Nucleotide alignment of the AaeZIP11Δ56 mutant. Dashes highlighted in yellow are the deleted bases. In the box, the electropherogram transitions from single peaks to double peaks, depicting the position where the deletion occurred (arrow). (B) Electropherogram of the end of the sequencing run and the transition from double peaks to single peaks. Note that the number of single peaks represents the number of bases deleted (gray rectangle). Primer sequences are provided in the Supplemental Table S1. Please click here to view a larger version of this figure.
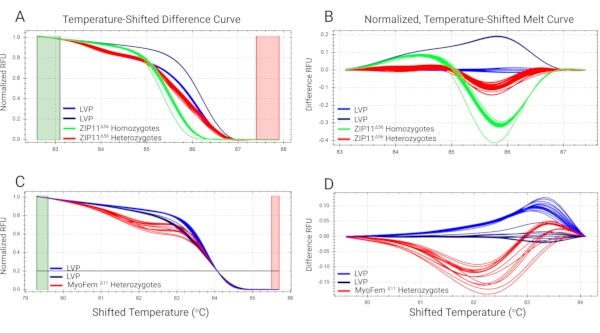
Figure 5: HRMA of DNA extracted from mosquito legs. DNA was extracted from a single leg from AaeZIP11 (A and B) and myo-fem (C and D) knockout Ae. aegypti mosquitoes and analyzed by HRMA. A and C denote the normalized fluorescence signals of the samples to relative values of 1.0 to 0. B and D denote the magnification of the curve differences by subtracting each curve from the wild-type reference (Liverpool strain). Abbreviations: HRMA = high-resolution melt analysis; LVP = Liverpool strain; RFU = relative fluorescence units. Primer sequences are provided in the Supplemental Table S1. Please click here to view a larger version of this figure.
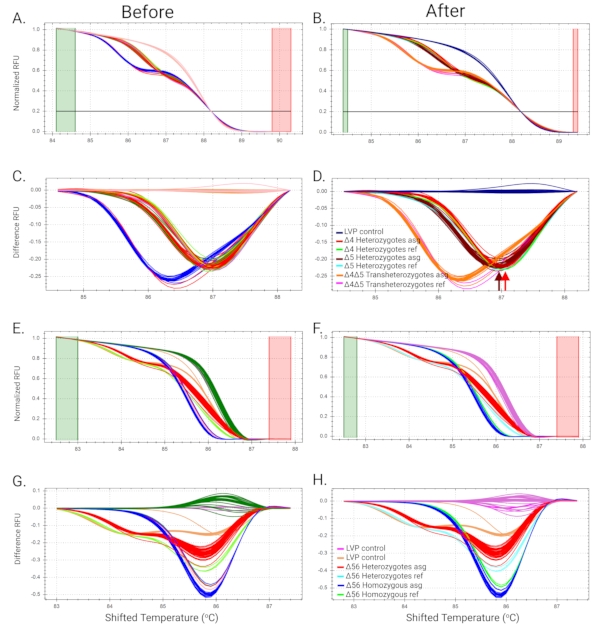
Figure 6: Examples of manual group assignments. (A–D) HRMA for Aeflightin mutants. (A and C) Melt curves (normalized linear scale curves and difference curves, respectively) are automatically grouped by the Precision Melt Analysis Software. (B) Normalization of differential curves altered manually. (D) After altering the normalization of the differential curves, each sample was assigned individually by the peak temperatures in the difference curves for corresponding positive controls (previously sequenced samples identified by Δ4 and Δ5 heterozygotes and Δ4Δ5 trans-heterozygotes), enabling the clear identification of the 3 groups. Red and brown arrows are added to highlight the peaks at different temperatures. (E–H) HRMA for AaeZIP11 mutants. (E and G) Melt curves are automatically assigned by the software. (F and H) Mutants in the second heterozygous self-cross were appropriately assigned to the correct groups by the similarity of the normalized and difference curves to previously determined references (color-coded in curves). Heterozygotes asg, Homozygotes asg, and Trans-heterozygotes asg: assigned melt curves by Precision Melt Analysis Software. Abbreviations: HRMA = high-resolution melt analysis; LVP = Liverpool strain; RFU = relative fluorescence units. Primer sequences are provided in Supplemental Table S1. Please click here to view a larger version of this figure.
Supplemental Material: Detailed protocol for setting up HRMA in the CFX96 Real-Time System (e.g., Bio-rad). Please click here to download this File.
Supplemental Figure S1: Step-by-step instructions for setting up the cycling protocol on Bio-rad CFX Manager. See Supplemental Material 2.1-2.3. Please click here to download this File.
Supplemental Figure S2: Step-by-step instructions for a plate setup on Bio-rad CFX Manager. See Supplemental Material 3.1-3.4. Please click here to download this File.
Supplemental Figure S3: Step-by-step instructions for a plate setup on Bio-rad CFX Manager. See Supplemental Material 3.5-3.6. Please click here to download this File.
Supplemental Figure S4: Step-by-step instructions for the run setup on Bio-rad CFX Manager. See Supplemental Material 4.1. Please click here to download this File.
Supplemental Figure S5: Step-by-step instructions for HRMA analysis on Bio-rad CFX Manager. See Supplemental Material 5.1-5.3. Abbreviation: HRMA = high-resolution melt analysis. Please click here to download this File.
Supplemental Table S1: List of primers. Please click here to download this Table.
Discussion
High-resolution melt analysis offers a simple and fast solution for the identification of indels generated by CRISPR/Cas9 technology in the vector mosquito Ae. aegypti. It provides flexibility, enabling the genotyping of mosquitoes mutated for a wide range of genes from flight muscle to iron metabolism and more13,14. HRMA can be performed in just a few hours from sample collection to the final analyses. Additional time is required for primer design and will also depend on the time to receive any needed sequencing results (for identification of the SNPs), sequence analyses, and primer ordering.
The protocol presented here details the step-by-step methods for successful analyses. One of the most critical steps in this process is the proper design of the primers. HRMA is sufficiently sensitive that the presence of even a single SNP within the fragment amplified can result in a strong difference curve that could be misinterpreted as a mutation. Sequencing laboratory colony samples prior to CRISPR mutagenesis allows for the detection and avoidance of SNP-rich regions when designing the target; the HRMA primers will help to prevent this issue. Designing primers for small amplicons (70-100 bp) can also improve the sensitivity and reproducibility of the experiment as larger fragments can present several melt domains, decreasing the chance to distinguish variants22. A limitation of the technique is that completely avoiding SNPs might not be feasible depending on the gene; thus, sequencing and further use of multiple wild-type control individuals may aid the analyses.
Specifically for this protocol, it is important to keep genotyped mosquitoes alive so they can be backcrossed for the establishment of mutant lines. Removing one of the mosquito legs to extract gDNA may be the least invasive way of achieving that; however, the amount of gDNA recovered from this tissue is minimal. The kit suggested in the Table of Materials was sufficient for obtaining DNA from a mosquito leg and performing the PCR to complete HRMA.
HRMA can offer more flexibility in terms of detection of a broad range of indel sizes, varying from single-base mismatches to multiple base pairs (AaeZIP11 had a 56 bp deletion) and surpassing methods such as the SNA or T7E1 (both enzyme mismatch cleavage assays) that are limited to single-base mismatch or small deletions (~20 bp)5,23. In addition, SNA does not detect homozygous mutants as the PCR products do not contain mismatches. Moreover, SNA results may also be masked by naturally occurring allelic polymorphisms when the nuclease-treated PCR products have a similar length to the mutant alleles. Neither of these limitations is applicable to HRMA. Ultimately, sequencing is necessary to identify the mutant DNA sequence. However, once a mutation in question is known, HRMA used for the identification of mutant individuals is much faster than direct sequencing. This method preserves valuable samples (more mosquitoes will survive, the less they have to wait for their genotype results) and reduces the need for sequencing of large numbers of samples, lowering the total cost of the experiment.
Disclosures
The authors have nothing to disclose.
Acknowledgements
All figures were created with Biorender.com under a license to Texas A&M University. This work was supported by funds from the National Institute of Allergy and Infectious Disease (AI137112 and AI115138 to Z.N.A.), Texas A&M AgriLife Research under the Insect Vectored Disease Grant Program, and the USDA National Institute of Food and Agriculture, Hatch project 1018401.
Materials
| 70% Ethanol | 70% ethanol solution in water | ||
| 96-well PCR and Real-time PCR plates | VWR | 82006-636 | For obtaining genomic DNA (from the mosquito leg) |
| 96-well plate templates | House-made printed, for genotype recording | ||
| Bio Rad CFX96 | Bio Rad | PCR machine with gradient and HRMA capabilities | |
| Diversified Biotech reagent reservoirs | VWR | 490006-896 | |
| Exo-CIP Rapid PCR Cleanup Kit | New England Biolabs | E1050S | |
| Glass Petri Dish | VWR | 89001-246 | 150 mm x 20 mm |
| Hard-shell thin-wall 96-well skirted PCR plates | Bio-rad | HSP9665 | For HRMA |
| Multi-channel pipettor (P10) | Integra Biosciences | 4721 | |
| Multi-channel pipettor (P300) | Integra Biosciences | 4723 | |
| Nunc Polyolefin Acrylate Sealing tape, Thermo Scientific | VWR | 37000-548 | To use with the 96-well PCR plates for obtaining genomic DNA |
| Optical sealing tape | Bio-rad | 2239444 | To use with the 96-well skirted PCR plates for HRMA |
| Phire Animal tissue direct PCR Kit (without sampling tools) | Thermo Fisher | F140WH | For obtaining genomic DNA and performing PCR |
| Plastic Fly Vial Dividers | Genesee | 59-128W | |
| Precision Melt Analysis Software | Bio Rad | 1845025 | Used for genotyping the mosquito DNA samples and analyzing the thermal denaturation properties of double-stranded DNA (see protocol step 3.3) |
| SeqMan Pro | DNAstar Lasergene software | For multiple sequence alignment | |
| Single-channel pipettor | Gilson | ||
| Tweezers Dumont #5 11 cm | WPI | 14098 | |
| White foam plugs | VWR | 60882-189 | |
| Wide Drosophila Vials, Polystyrene | Genesee | 32-117 | |
| Wide Fly Vial Tray, Blue | Genesee | 59-164B |
References
- WHO. Dengue Guidelines for diagnosis, treatment, prevention and control. WHO. , 160 (2009).
- WHO. Zika epidemiology update. WHO. , 1-13 (2019).
- WHO. Guidelines for prevention and control of Chikungunya fever. WHO. , (2009).
- World malaria report 2020: 20 years of global progress and challenges. World Health Organization Available from: https://www.who.int/publications/i/item/9789240015791 (2020)
- Qiu, P., et al. Mutation detection using Surveyor nuclease. Biotechniques. 36 (4), 702-707 (2004).
- Babon, J. J., McKenzie, M., Cotton, R. G. The use of resolvases T4 endonuclease VII and T7 endonuclease I in mutation detection. Molecular Biotechnology. 23 (1), 73-81 (2003).
- Erali, M., Voelkerding, K. V., Wittwer, C. T. High resolution melting applications for clinical laboratory medicine. Experimental and Molecular Pathology. 85 (1), 50-58 (2008).
- Reed, G. H., Wittwer, C. T. Sensitivity and specificity of single-nucleotide polymorphism scanning by high-resolution melting analysis. Clinical Chemistry. 50 (10), 1748-1754 (2004).
- Parant, J. M., George, S. A., Pryor, R., Wittwer, C. T., Yost, H. J. A rapid and efficient method of genotyping zebrafish mutants. Developmental Dynamics. 238 (12), 3168-3174 (2009).
- Tong, S. Y., Giffard, P. M. Microbiological applications of high-resolution melting analysis. Journal of Clinical Microbiology. 50 (11), 3418-3421 (2012).
- Simko, I. High-resolution DNA melting analysis in plant research. Trends in Plant Science. 21 (6), 528-537 (2016).
- Basu, S., et al. Silencing of end-joining repair for efficient site-specific gene insertion after TALEN/CRISPR mutagenesis in Aedes aegypti. Proceedings of the National Academy of Sciences of the United States of America. 112 (13), 4038-4043 (2015).
- O’Leary, S., Adelman, Z. N. CRISPR/Cas9 knockout of female-biased genes AeAct-4 or myo-fem in Ae. aegypti results in a flightless phenotype in female, but not male mosquitoes. PLoS Neglected Tropical Diseases. 14 (12), 0008971 (2020).
- Kojin, B. B., et al. Aedes aegypti SGS1 is critical for Plasmodium gallinaceum infection of both the mosquito midgut and salivary glands. Malaria Journal. 20 (1), 11 (2021).
- Ye, J., et al. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics. 13, 134 (2012).
- Larkin, M. A., et al. Clustal W and Clustal X version 2.0. Bioinformatics. 23 (21), 2947-2948 (2007).
- Notredame, C., Higgins, D. G., Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. Journal of Molecular Biology. 302 (1), 205-217 (2000).
- Bassett, A. R., Tibbit, C., Ponting, C. P., Liu, J. L. Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Reports. 6 (6), 1178-1179 (2014).
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Research. 31 (13), 3406-3415 (2003).
- Hill, J. T., et al. Poly peak parser: Method and software for identification of unknown indels using sanger sequencing of polymerase chain reaction products. Developmental Dynamics. 243 (12), 1632-1636 (2014).
- Tsujimoto, H., Anderson, M. A. E., Myles, K. M., Adelman, Z. N. Identification of candidate iron transporters from the ZIP/ZnT gene families in the mosquito Aedes aegypti. Frontiers in Physiology. 9, 380 (2018).
- Slomka, M., Sobalska-Kwapis, M., Wachulec, M., Bartosz, G., Strapagiel, D. High resolution melting (HRM) for high-throughput genotyping-limitations and caveats in practical case studies. International Journal of Molecular Sciences. 18 (11), 2316 (2017).
- Vouillot, L., Thelie, A., Pollet, N. Comparison of T7E1 and surveyor mismatch cleavage assays to detect mutations triggered by engineered nucleases. G3. 5 (3), 407-415 (2015).

