Fabrication of 3D Cardiac Microtissue Arrays using Human iPSC-Derived Cardiomyocytes, Cardiac Fibroblasts, and Endothelial Cells
Summary
Here, we describe an easy-to-use methodology to generate 3D self-assembled cardiac microtissue arrays composed of pre-differentiated human-induced pluripotent stem cell-derived cardiomyocytes, cardiac fibroblasts, and endothelial cells. This user-friendly and low cell requiring technique to generate cardiac microtissues can be implemented for disease modeling and early stages of drug development.
Abstract
Generation of human cardiomyocytes (CMs), cardiac fibroblasts (CFs), and endothelial cells (ECs) from induced pluripotent stem cells (iPSCs) has provided a unique opportunity to study the complex interplay among different cardiovascular cell types that drives tissue development and disease. In the area of cardiac tissue models, several sophisticated three-dimensional (3D) approaches use induced pluripotent stem cell-derived cardiomyocytes (iPSC-CMs) to mimic physiological relevance and native tissue environment with a combination of extracellular matrices and crosslinkers. However, these systems are complex to fabricate without microfabrication expertise and require several weeks to self-assemble. Most importantly, many of these systems lack vascular cells and cardiac fibroblasts that make up over 60% of the nonmyocytes in the human heart. Here we describe the derivation of all three cardiac cell types from iPSCs to fabricate cardiac microtissues. This facile replica molding technique allows cardiac microtissue culture in standard multi-well cell culture plates for several weeks. The platform allows user-defined control over microtissue sizes based on initial seeding density and requires less than 3 days for self-assembly to achieve observable cardiac microtissue contractions. Furthermore, the cardiac microtissues can be easily digested while maintaining high cell viability for single-cell interrogation with the use of flow cytometry and single-cell RNA sequencing (scRNA-seq). We envision that this in vitro model of cardiac microtissues will help accelerate validation studies in drug discovery and disease modeling.
Introduction
Drug discovery and disease modeling in the field of cardiovascular research face several challenges due to a lack of clinically relevant samples and inadequate translational tools1. Highly complex pre-clinical models or oversimplified in vitro single-cell models do not exhibit pathophysiological conditions in a reproducible manner. Therefore, several miniaturized tissue-engineered platforms have evolved to help bridge the gap, with the goal of achieving a balance between ease of application in a high-throughput manner and faithful recapitulation of tissue function2,3. With the advent of induced pluripotent stem cell (iPSC) technology, tissue engineering tools can be applied to patient-specific cells with or without underlying cardiovascular disease state to answer research questions4,5,6. Such tissue engineered models with cellular composition similar to the heart tissue could be utilized in drug development efforts to test for cardiotoxicity and dysfunction induced by pathological changes in behavior of one or multiple cell types.
Self-assembled microtissues or organoids derived from human iPSCs are three-dimensional (3D) structures that are miniature tissue-like assemblies exhibiting functional similarities to their in vivo counterparts. There are several different approaches that allow formation of organoids in situ via directed differentiation of iPSCs or through the formation of embryoid bodies4. The resulting organoids are an indispensable tool to study morphogenetic processes that drive organogenesis. However, the presence of a variety of cell populations and differences in self-organization can lead to variability in outcomes between different organoids5. Alternatively, pre-differentiated cells that are self-assembled into microtissues with tissue-specific cell types to study local cell-cell interactions are excellent models, where it is feasible to isolate the self-assembled components. Particularly in human cardiac research, development of 3D cardiac microtissues with multicellular components has proven to be challenging when cells are derived from different patient lines or commercial sources.
To improve our mechanistic understanding of cell behaviors in a physiologically relevant, personalized, in vitro model, ideally all component cell types should be derived from the same patient line. In the context of a human heart, a truly representative cardiac in vitro model would capture the crosstalk among predominant cell types, namely, cardiomyocytes (CMs), endothelial cells (ECs), and cardiac fibroblasts (CFs)6,7. The faithful recapitulation of a myocardium not only requires biophysical stretch and electrophysiological stimulation, but also cell-cell signaling that arise from supporting cell types such as ECs and CFs8. CFs are involved in the synthesis of extracellular matrix and maintaining tissue structure; and in a pathological state, CFs can induce fibrosis and alter electrical conduction in the CMs9. Similarly, ECs can regulate contractile properties of CMs through paracrine signaling and supplying vital metabolic demands10. Hence, there is a need for human cardiac microtissues composed of all three major cell types to allow physiologically relevant high-throughput experiments to be conducted.
Here, we describe a bottom-up approach in fabrication of cardiac microtissues by derivation of human iPSC-derived cardiomyocytes (iPSC-CMs), iPSC-derived endothelial cells (iPSC-ECs), and iPSC-derived cardiac fibroblasts (iPSC-CFs) and their 3D culture in uniform cardiac microtissue arrays. This facile method of generating spontaneously beating cardiac microtissues can be utilized for disease modeling and rapid testing of drugs for functional and mechanistic understanding of heart physiology. Furthermore, such multicellular cardiac microtissue platforms could be exploited with genome editing techniques to emulate cardiac disease progression over time under chronic or acute culture conditions.
Protocol
1. Medium, reagent, culture plate preparation
- Cell wash solution for cell culture: Use 1x phosphate buffered saline (PBS) or Hanks balanced salt solution (HBSS) without calcium or magnesium.
- Cardiomyocyte differentiation media
- Prepare differentiation Medium #1 by adding 10 mL supplement (50x B27 plus insulin) to 500 mL cardiomyocyte basal medium (RPMI 1640).
- Prepare differentiation Medium #2 by adding 10 mL supplement (50x B27 minus insulin) to 500 mL cardiomyocyte basal medium (RPMI 1640).
- Prepare purification Medium #3 by adding supplement (50x B27 plus insulin) to 500 mL cardiomyocyte basal medium (RPMI 1640) minus glucose.
- Endothelial growth media and Magnetic-activated cell sorting (MACS) reagents
- Prepare endothelial growth medium (EGM) with commercially available endothelial cell growth medium supplement kit.
- Prepare cell separation sorting buffer solution by diluting bovine serum albumin (BSA) stock solution to 1:20 with rinsing solution.
- Cardiac fibroblast differentiation medium: Prepare cardiac fibroblast differentiation medium with fibroblast basal medium (DMEM-high glucose) and Serum-Free Fibroblast Life Factors kit.
- Cardiac microtissue fabrication and maintenance medium: Prepare filtered medium with cardiomyocyte basal medium (RPMI 1640) with supplement (50x B27 plus insulin) and EGM in 70/30 v/v% ratio.
- Small molecule and growth factor stock solutions
- For differentiation of all three cell types, prepare 200 µL aliquots of GSK3-beta inhibitor CHIR 99021 (6-[[2-[[4-(2,4-Dichlorophenyl)-5-(5-methyl-1H-imidazol-2-yl)-2-pyrimidinyl]amino]ethyl]amino]-3-pyridinecarbonitrile); Wnt inhibitor IWR-1-endo (4-(1,3,3a,4,7,7a-Hexahydro-1,3-dioxo-4,7-methano-2H-isoindol-2-yl)-N-8-quinolinyl-Benzamide; and transforming growth factor beta (TGF-β) inhibitor SB431542 (4-[4-(1,3-Benzodioxol-5-yl)-5-(2-pyridinyl)-1H-imidazol-2-yl]benzamid) at 10 mM concentration in dimethyl sulfoxide (DMSO).
- For human iPSC-EC differentiation, prepare 100 µL aliquots of basic fibroblast growth factor (bFGF) (20 µg/mL), vascular endothelial growth factor 165 (VEGF165; 50 µg/mL), and bone morphogenetic protein 4 (BMP4; 20 µg/mL) in 0.1% (w/v) BSA in ultrapure distilled water. Store the aliquots at -20 °C; for long-term storage, aliquots can be stored at -80 °C for up to 1 year.
- Pre-coating plates for maintenance of iPSC-CMs, iPSC-ECs, and iPSC-CFs.
- Prepare basement membrane matrix medium coated 6-well plates for iPSC-CM re-plating by diluting Growth Factor Reduced (GFR) basement membrane matrix medium in DMEM/F12 in a 1:200 ratio. Add 2 mL of diluted basement membrane matrix medium to each well of the 6-well plate and leave it to set for at least 2 to 4 h.
- Pre-coat 6-well plate or 10 cm dish with gelatin. Liquefy 2% gelatin solution in a water bath at 37 °C and prepare filtered 0.2% (v/v) gelatin in PBS in an appropriate volume as needed. Coat the culture plate with 10 mL of 0.2% gelatin for at least 30 min at 37 °C prior to use.
NOTE: Gelatin plates can be used for both iPSC-CFs and iPSC-ECs after MACS.
- Fabrication of microtissue molds: Prepare 2% low melting agarose solution in PBS in a dry 100 mL glass bottle. Sterilize the agarose at 121 °C, 15 psi for 20 min prior to use. Autoclave the casting silicone micro-molds at 121 °C, 15 psi for 15 min.
- Solutions for digestion, immunostaining, and flow cytometry
- For microtissue digestion, prepare an enzyme digestion buffer of Dispase I (1 U/mL) and Liberase (3 U/mL) in PBS. Keep this solution on ice.
- For both cell and microtissue fixation, use ice cold commercially available fixation reagent containing 4.2% paraformaldehyde (PFA).
- Prepare 0.25% Triton X-100 in PBS for permeabilization and 0.1% Tween-20 for washes. The solution can be stored at room temperature.
- For fluorescence-activated cell sorting (FACS) analyses, prepare FACS buffer [PBS or HBSS with 2% fetal bovine serum (FBS)] and store at 4 °C.
- For immunostaining, prepare 2% normal goat/donkey/rabbit serum in PBS. Choose the serum species based on the host in which the antibodies were raised.
2. Cardiac differentiation and purification
NOTE: All iPSCs should be maintained at ~75% to 80% confluency prior to cardiomyocyte differentiation. iPSCs used for this protocol were derived from peripheral blood mononuclear cells (PBMCs) using Sendai virus reprogramming performed at the Stanford Cardiovascular Institute (SCVI) Biobank.
- Prior to differentiation, culture iPSC colonies up to passage (P20).
- Remove the iPSC culture medium from the 6-well plate and wash the cells once with 2 mL PBS.
- On day 0, begin mesendoderm induction by adding 2 mL of differentiation Medium #1 with CHIR (6 µM final) in each well. The final concentration may vary between 6 to 9 µM for different iPSC lines. Hence, it is recommended to test different CHIR concentrations on a 6-well plate to identify optimal cardiac mesoderm induction.
- On day 2, recover the cells by replacing with fresh 2 mL Medium #1 in each well.
- On day 3, replace medium in each well with 2 mL differentiation Medium #1 with Wnt inhibitor IWR-1-endo (5 µM) to induce cardiac-lineage specification.
- On day 5, recover the cells by replacing medium with fresh 2 mL Medium #1 in each well.
- On day 7, replace medium with 2 mL Medium #2 in each well. Spontaneous beating of cardiomyocytes may be observed as early as day 8-9. For some lines, beating cells may appear as late as day 11.
- For purification of the differentiation culture, replace with 2 mL minus glucose Medium #3 in each well on day 10.
- On day 13, recover the cells by changing the medium with 2 mL plus glucose Medium #2 in each well.
- On day 16, perform second round of purification with 2 mL of Medium #3 in each well.
- Recover the cells on day 19 with 2 mL plus glucose Medium #2.
- On day 20, wash the wells once with 1 mL PBS and dissociate the cells using 1 mL 10x Trypsin for 6 min at 37 °C. After incubation, the cells are lifted off the well surface into single cells for less than 15 s and added to 15 mL conical tube with equal volume of Medium #2 to neutralize the enzyme. Centrifuge the cell suspension for 3 min at 270 x g.
- Resuspend the cell pellet in 3 mL Medium #3. Count the cells and add appropriate volume of Medium #3 to plate a total of ~3 million cells in each well of a new basement membrane matrix medium coated 6-well plate. On the second day after re-plating, replace the medium with fresh 2 mL Medium #2, and replenish with 2 mL Medium #2 every other day until trypsinization of cardiomyocytes for freezing or future experiments.
3. Endothelial cell differentiation and MACS
- At 75% to 80% iPSC confluency, wash the plate with PBS, 2 mL per well.
- Begin differentiation at day 0 by replacing with 2 mL of differentiation Medium #1 with 6 µM CHIR.
- On day 2, replace the medium with 2 mL of differentiation Medium #1 with 2 µM CHIR.
- On day 4, replace the medium with 2 mL of EGM supplemented with 20 ng bFGF-2, 50 ng VEGF165, and 20 ng BMP4.
- Change the medium with 2 mL of EGM supplemented with growth factors every 2 days from day 6 to day 10. Addition of TGFβ inhibitor (SB431542) at 8 µM concentration is optional to promote endothelial expansion and suppress differentiation of other mesenchymal origin cell types.
- On day 12, begin steps for MACS by rinsing each well with 1 mL of PBS followed by cell dissociation using 1 mL of 10x Trypsin in each well of a 6-well plate for 8 min at 37 °C.
- Prepare an equal volume of EGM medium in a 50 mL conical tube to neutralize the dissociation enzyme. Place a 40 µm cell strainer on the 50 mL conical tube and pass the dissociated cell suspension through the strainer. Change the filter for every 2 to 3 dissociated wells.
- Enumerate the total viable cells using 0.4% Trypan blue using a hemocytometer or an automated cell counter.
- Pellet the cell suspension at 300 x g for 5 min in a pre-cooled centrifuge set at 4 °C.
- Discard the supernatant and resuspend the cell pellet in appropriate volume of sorting buffer based on the total count in step 3.8.
NOTE: Add 80 µL of sorting buffer per 107 total cells. - For magnetic bead labeling, add 20 µL of FcR Blocking reagent per 107 cells and incubate for 5 min.
- Add 20 µL of CD144 or CD31 Microbeads per 107 cells and mix well. Incubate the cell suspension in dark at 4 °C for 15 min.
- Add 20 mL of sorting buffer to wash the labeled cells and pellet the cells 300 x g for 5 min in a pre-cooled centrifuge set at 4 °C.
- Resuspend the cell pellet in 3 mL of sorting buffer and leave it on ice.
- Prepare a magnetic separation column by placing the column in the separator notch.
- Equilibrate the column by rinsing with 3 mL of sorting buffer and collect the flow through in a waste conical tube.
- Once the rinsing buffer flows through, resuspend the cell suspension thoroughly to break any clumps and apply the cell suspension onto the column.
- After the cell suspension has flowed through, wash the column with 3 mL of sorting buffer three times to remove any unlabeled cells.
- Remove the column from the separator and place it in a 15 mL conical tube for CD144+/CD31+ cell elution.
- Add 5 mL of sorting buffer onto the column and immediately flush the magnetically labeled cells with the plunger.
- Determine the viable cell number using 0.4% Trypan blue using a hemocytometer or an automated cell counter.
- Centrifuge the cell suspension in a pre-cooled centrifuge at 300 x g for 3 min.
- Resuspend the cell pellet in appropriate volume of EGM with 5-8 µM of TGFβ inhibitor (SB431542) based on the total count in step 3.21.
- Plate this passage 0 (P0) cells at an appropriate cell density (4 x 104 cells/cm2) on a pre-coated 0.2% gelatin plate.
- For P0 and P1, continue to replenish the medium with fresh EGM every 2 days with TGFβ inhibitor. From P2, cells can be cultured in endothelial growth medium without TGFβ inhibitor.
4. Cardiac fibroblast differentiation
- Allow iPSCs to become 90% to 95% confluent; wash each well with 1 mL PBS.
- Begin differentiation at day 0 by adding 2 mL of differentiation Medium #1 with 11 µM CHIR. For sensitive iPSC lines, the concentration may vary between 9-10 µM.
- On day 1, observe the plate. It is normal to observe significant cell death with ~30% to 40% cells adhered to the plate.
- On day 3, add 2 mL of differentiation Medium #1 with 5 µM IWR-1-endo to promote expansion of cardiac progenitors.
- On day 4, replace the medium with 2 mL of cardiac fibroblast differentiation medium. Replace with fresh medium every 2 days till day 16.
- On day 18, detach the cells using 1 mL of 10x Trypsin in each well of a 6-well plate for 10 min at 37 °C.
- Disrupt the cell layer thoroughly and pass the cell suspension through a 70 µm cell strainer in a 50 mL conical tube containing equal volume of DMEM/F12 medium supplemented with 5% FBS.
- Determine the viable cell number using 0.4% Trypan blue using a hemocytometer or an automated cell counter.
- Centrifuge the cell suspension at 300 x g for 5 min to obtain a pellet.
- For the first re-plating, resuspend the cell pellet in an appropriate volume of differentiation medium and plate at a cell density (6 x 104 cells/cm2) on a 0.2% gelatin coated plate until 90% confluency.
- Split the plate and maintain the cardiac fibroblasts in regular DMEM/F12 with 10% serum on gelatin coated plates.
5. Casting of cardiac microtissue molds and cell seeding
- Melt the agarose in a microwave in a 100 mL glass bottle until boiling. Spray the agarose bottle and place it in the biosafety cabinet. Allow the agarose to cool down for ~3 min.
- Pipette 700 µL of molten agarose in a silicone micro-mold of 9 x 9 array. Avoid generating bubbles while pipetting.
- Carefully place the mold on a pre-cooled ice block to accelerate agarose gelation.
- Ensure that once the agarose is gelled, it becomes translucent. Carefully bend around the edges of the micro-mold to loosen the agarose replica. Then, gently peel the replica from all sides to detach the agarose replica from the silicone micro-mold.
- Transfer the agarose microtissue tray containing 81 circular recesses (800 µm diameter; 800 µm depth) into a sterile 12-well plate.
- Add 2 mL of PBS to the agarose microtissue tray and inspect under the microscope for any trapped bubbles or irregular shaped wells.
- Submerge the agarose tray in 2 mL 70% ethanol overnight, followed by UV treatment in the biosafety cabinet for 1 h.
- Prior to use, remove the 70% ethanol and wash twice with distilled water and a final wash with 2 mL PBS.
- Trypsinize, neutralize, and count iPSC-CMs, iPSC-ECs, and iPSC-CFs and place the cell suspensions on ice.
- Remove the PBS from the well and the cell seeding chamber carefully without touching the recesses.
- In a new tube, mix iPSC-CMs, iPSC-ECs, and iPSC-CFs in 7:2:1 ratio, respectively, to achieve a final cell density of 106 cells/mL. Higher cell densities will result in larger microtissues.
NOTE: Do not exceed 2 x 106 cells/mL. - Carefully add the 200 µL of the cell suspension dropwise in the seeding chamber.
- Allow the cells to settle at 37 °C in a CO2 incubator for 2 h.
- Add microtissue fabrication medium surrounding the agarose mold to just cover the surface of the inner chamber.
- After 24 h, the cells self-assemble and compact significantly in the circular recesses. Replace with fresh medium every 2 days for maintenance.
6. Fixation and permeabilization of cells and cardiac microtissues for immunostaining
- For each individual cell types, culture the cells separately on basement membrane matrix medium or gelatin coated chamber slides (approximately 2.5-3 x 105 cells/mL). Cardiac microtissues can be collected in 15 mL conical tube by gently flushing them out of the circular recesses.
- Aspirate the medium and rinse the cells or microtissues with 1 mL PBS; thereafter, fix with the fixation buffer containing 4.2% PFA for 20 min for the chamber slides and 1 h for microtissues at room temperature.
- Aspirate the PFA and add 1 mL of permeabilizing solution (0.25% Triton X-100 in PBS) for 5 to 7 min for chamber slides and 20 min for microtissues in 15 mL conical tube.
NOTE: From this step onwards, the samples can be gently rocked on a benchtop rocking platform. - Aspirate the permeabilizing solution and rinse once with 2 to 3 mL of PBS.
- Incubate the cells with 500-1,000 µL of blocking solution (2% to 5% normal goat serum or donkey serum) for at least 1 h for the chamber slides and 3 to 4 h for the microtissues.
- Incubate the cells or cardiac microtissues with conjugated antibodies prepared in the blocking solution sufficient to cover the sample. Incubate with anti-cardiac Troponin-T (cTnT2) (1:50), anti-CD31 (1:75), and anti-DDR2/Vimentin (1:50) for 1 h for the chamber slides and overnight at 4 °C for cardiac microtissues.
- Wash the chamber slides three times with 500 µL 0.1% Tween-20 for 5 min between each wash and a final wash with PBS.
- For the cardiac microtissues, wash with 2 mL 0.1% Tween-20 five times with 20 min duration between each wash. Perform a final wash step for an additional 20 min.
- Incubate the cells or microtissues with 4',6-diamidino-2-phenylindole (DAPI) (1 µg/mL) prior to confocal microscopy.
- For cardiac microtissues, transfer carefully to a 35 mm glass bottom dish and add PBS to submerge microtissues.
- For 3D imaging, using a 20x or 40x oil immersion objective gain center focus of the microtissue and adjust exposure parameters for each fluorophore.
- For obtaining a Z-stack, set first and last coordinates in Live mode with a total imaging depth of 100-200 µm with 5-10 µm slice interval.
7. Digestion of cardiac microtissues and preparation of cells for flow cytometry
- To digest microtissues, gently flush the microtissues with Medium #1 out of the circular recesses using a wide-bore 1 mL pipette tip into a 15 mL conical tube.
- Allow the microtissues to settle and aspirate the medium carefully and rinse the cells or microtissues with 1 mL PBS and add 200-300 µL of enzyme digestion buffer for 20 min at 37 °C. At 10 min, mix the microtissues gently for 1 min and incubate again at 37 °C for the remainder of time.
- After incubation, use a regular 1 mL pipette tip to mix the microtissues vigorously to obtain a turbid cell suspension.
- Once the microtissues are sufficiently digested into single cells, immediately neutralize the cell suspension with 5 mL of medium containing 5% FBS and strain the cell suspension through a 40 µm cell strainer. Count the total number of cells and centrifuge the single cell suspension at 300 x g for 5 min at 4 °C.
- Aspirate the supernatant and resuspend 1 x 105-1 x 106 cells in 100 µL annexin-binding buffer with FITC Annexin V and 100 µg/mL propidium iodide (PI) or To-Pro3 dead cell exclusion dye for 10 min on ice.
- After incubation, add 300 µL of the annexin binding buffer to the cell suspension and transfer to a round bottom FACS tube for flow cytometry analysis. Use the correct lasers and emission filters for the select fluorophores.
- For quantification of cell surface markers using fixed cells, perform the fixation and permeabilization of the cell pellet as described in steps 6.2 and 6.3.
- After permeabilization, rinse the cell pellet and incubate the cells with respective conjugated antibodies for 1 h. Wash the cell pellet in 4 mL FACS buffer (2% FBS in PBS) and centrifuge at 300 x g for 3 min. Repeat the wash step twice.
- Resuspend the cells in 200-300 µL FACS buffer for flow cytometry analysis.
- Adjust the forward and side scatter properties with an unstained sample and consider using an isotype control for each fluorophore to adjust the laser voltages. Collect a minimum of 20,000 events for data analysis.
8. Performing contraction analyses of spontaneously beating cardiac microtissues
- Record videos of cardiac microtissues to capture at least three beats. Set the recording resolution to at least 1280 x 720 pixels at frame rate >30 frames per second and save the video in .AVI format.
- Run the MotionGUI script11 in a MATLAB environment to launch the user interface.
- Find the .AVI file location in your folder to load the video. Then, enter the frame rate at which the video was captured in the Input panel.
- In the advanced input panel, a pixel size may be specified based on the resolution and the capture magnification.
- A suitable macroblock pixel size can be specified (default 16) and a detectable pixel motion depending on the strength of microtissue contraction.
- After adjusting the parameters, click on Get Motion Vectors to begin the analysis.
- Select a region of interest with the Choose AOI radio button to exclude areas surrounding the single microtissue in the circular recess.
- Use the function Map Time Ave to generate a mean contraction heatmap based on motion detected on X and Y axes.
- For peak tracing data, use the function Get Contraction Data to automatically measure the contraction and relaxation peaks.
- In the event of a low signal-to-noise ratio, apply a peak height and distance threshold to correctly annotate the maximum contraction velocity (blue dot) and maximum relaxation velocity (red triangle).
- After setting the correct thresholds, select Analyze Peaks to obtain the contraction and relaxation values with beat rate and beat interval.
- Obtain measurements from a minimum of 25 individual microtissues for statistical analyses.
Representative Results
Immunostaining and flow cytometry characterization of iPSC-derived CMs, ECs, and CFs
To generate cardiac microtissues composed of iPSC-CMs, iPSC-ECs, and iPSC-CFs, all three cell types are differentiated and characterized individually. In vitro differentiation of iPSCs to iPSC-CMs has improved over the past several years. However, the yield and purity of iPSC-CMs differ from line to line. The current protocol yields over 75% pure iPSC-CMs that spontaneously start beating around day 9 (Figure 1A). Further purification steps from day 9 to day 14 can improve iPSC-CM purity to over 80% as previously described12. Similarly, high-purity iPSC-ECs can be generated using previously published protocols13,14 that include the addition of several vascular growth factors that polarize endothelial progenitors arising from the mesoderm around days 4-5 (Figure 1B) to form phenotypically well-defined iPSC-ECs. iPSC-CFs are a highly heterogenous population based on their location and are characterized based on their morphology and expression of extracellular matrix proteins. Here, using published protocols15,16,17 with modifications, human iPSC-CFs are obtained from cardiac mesoderm progenitor cells (Figure 1C).
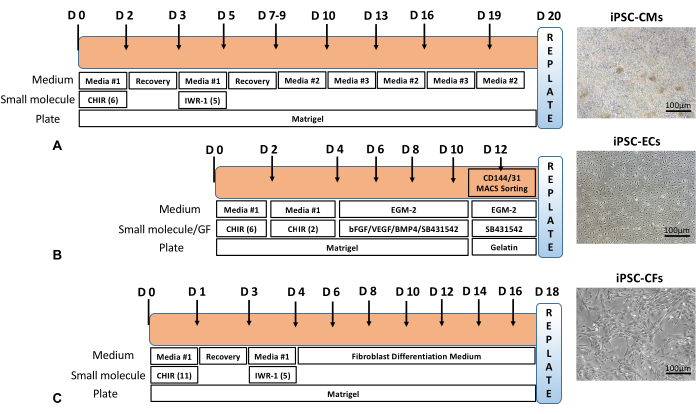
Figure 1: Differentiation timeline. Overall schematic of (A) iPSC-CM, (B) iPSC-EC, and (C) iPSC-CF differentiation timeline with representative phase contrast images of cells after differentiation and purification steps. Scale bar = 100 µm. Please click here to view a larger version of this figure.
The purity of iPSC-CMs, iPSC-ECs, and iPSC-CFs were determined by immunostaining and flow cytometry using cTnT2, platelet endothelial cell adhesion molecule (PECAM1/CD31) and vimentin (VIM), respectively (Figure 2A). Quantitative analysis showed a purified population containing over 90% cTnT2 cells at day 20. iPSC-ECs obtained at day 12 after MACS using CD31 beads were identified with immunostaining against PECAM1/CD31 endothelial cell surface marker. MACS yielded highly pure endothelial cells as evidenced by over 95% CD31+ cells at P0. However, it must be noted that the purity of endothelial cells decreased with higher passage numbers due to de-differentiation. Similarly, at day 20, flow cytometry analyses revealed that over 95% iPSC-CFs expressed the fibroblast marker VIM (Figure 2B).
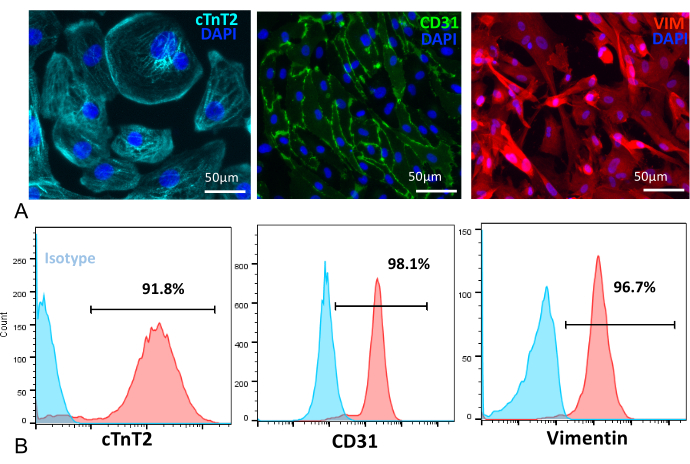
Figure 2: Immunofluorescence and flow cytometry characterization. (A) [Left to right panel] iPSC-CMs at day 25 stained with cTnT2 (cyan), iPSC-ECs stained with PECAM1/CD31 (green), iPSC-CFs stained with VIM (red), and nuclei stained with DAPI (blue). Scale bar = 50 µm. (B) [Left to right panel] Flow cytometry quantification showed a high percent purity of iPSC-CMs (91.8%), iPSC-ECs (98.1%), and iPSC-CFs (96.7%) following differentiation. Please click here to view a larger version of this figure.
Fabrication of cardiac microtissue cultures and size analyses
Single cell suspensions of iPSC-CMs, iPSC-ECs, and iPSC-CFs were mixed in 7:2:1 ratio and carefully dispensed into the cell seeding chamber of the sterilized agarose replica mold (Figure 3A,B). The cells uniformly settled inside the circular recesses in 2 h. Around day 3, the self-assembled cells organize into uniform sized spontaneously beating cardiac microtissues (Figure 3C). Arrays of different sized microtissues can be fabricated by tuning the final cell density (Figure 3D,E). Cardiac microtissues fabricated with a final cell density of 1 x 106 cells/mL is ~300-350 µm in diameter, 2 x 106 cells/mL is ~600 µm in diameter, and 4 x 106 cells/mL is over 800 µm in diameter. Microtissue assembly was obtained with a cell density of 1 x 106 cells/mL, which is typically used for experiments. These microtissue cultures can be maintained in culture for up to 6 weeks.
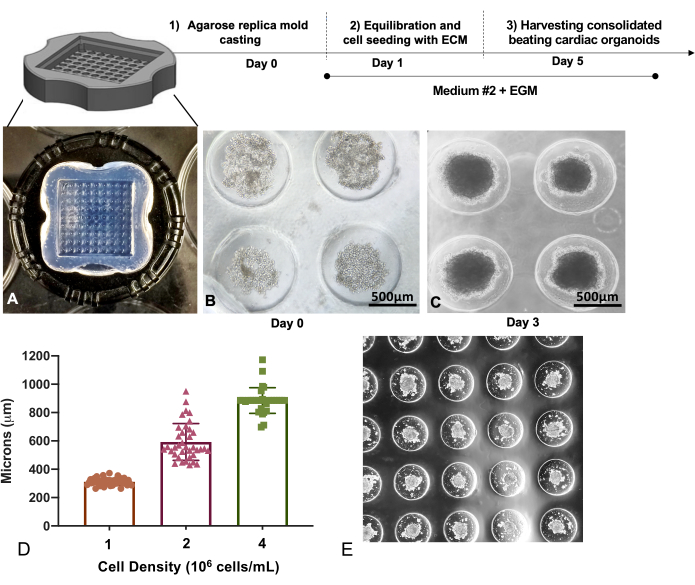

Figure 3: Replica molding technique to generate multicellular cardiac microtissues. (A) Replica molded agarose microwell trays of (B) iPSC-CM, iPSC-EC, and iPSC-CF mixtures captured inside the microwells for self-assembly. Scale bar 500 = µm. (C) Micrograph showing compaction of cardiac microtissues on day 3. Scale bar = 500 µm. (D) Cardiac microtissue sizes formed show a linear relationship with initial seeding densities, with higher cell densities resulting in larger microtissues. (E) A representative figure showing the self-assembled cardiac microtissues in the microwell array. Please click here to view a larger version of this figure.
Immunostaining and viability after enzymatic digestion
Immunofluorescence staining of day 12 post-fabrication using antibodies against cTnT2 for iPSC-CMs, CD31 for iPSC-ECs, and DDR2 for iPSC-CFs revealed a unique cell distribution in cardiac microtissues. iPSC-CMs, the heaviest of all three cell types, occupied the center, whereas iPSC-ECs were interspersed throughout the microtissues, and iPSC-CFs were observed to predominantly occupy the periphery (Figure 4A). Short and rapid digestion of the microtissues achieved using Dispase I and Liberase TL resulted in overall highly viable cell proportion (Figure 4B, top panel) with less than 5% apoptotic cells after 2 weeks in culture. This was followed by a brief 1 h exposure of cardiac microtissues to a high concentration (5 µM) of Doxorubicin, a chemotherapeutic drug that is known to induce dose-dependent cardiotoxicity (Figure 4B, bottom panel).
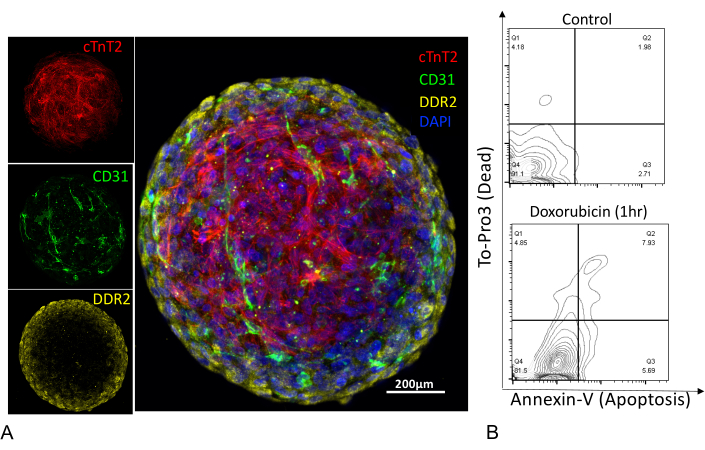
Figure 4: Immunostaining of cardiac microtissues and assessment of cell viability. (A) Confocal z-stack images of human cardiac microtissue stained for iPSC-CMs (cTnT2), iPSC-ECs (CD31), iPSC-CFs (DDR2), and nuclei stained with (DAPI). Scale bar = 200 µm. (B) Flow cytometry plots of cardiac microtissues enzymatically digested into single cell suspension and stained with Annexin V (apoptotic marker) and dead cell exclusion dye (To-Pro3). Digested single cell suspension showed a high cell viability (91%) with <5% apoptotic cell population (top panel), compared to single cell suspension treated with an apoptosis-inducing chemotherapeutic drug, Doxorubicin, for 1 h at 5 µM concentration (bottom panel). Please click here to view a larger version of this figure.
Computational contractility analysis
Contractility analyses of individual cardiac microtissues can be performed with the help of a MATLAB-based image analyses tool. Video recordings of spontaneously beating cardiac microtissues were obtained at 30 fps for analysis. As described previously11, the block-matching method employs motion tracking algorithm to capture movement of a block of pixels for the total frames acquired as a time series of motion vectors. The contractility of the microtissues and movement of the vectors generate a pseudo heatmap that illustrates mean or average contraction profile across the microtissue (Figure 5A). Contractile motion of the cardiac microtissues generates positive peaks that are measured as contraction velocity (blue circle), relaxation velocity (red triangle), and beat rate, the last of which is calculated as the time between two contraction cycles (Figure 5B). Furthermore, the contractility of the cardiac microtissues do not change significantly over 4 weeks in culture (Figure 5C).
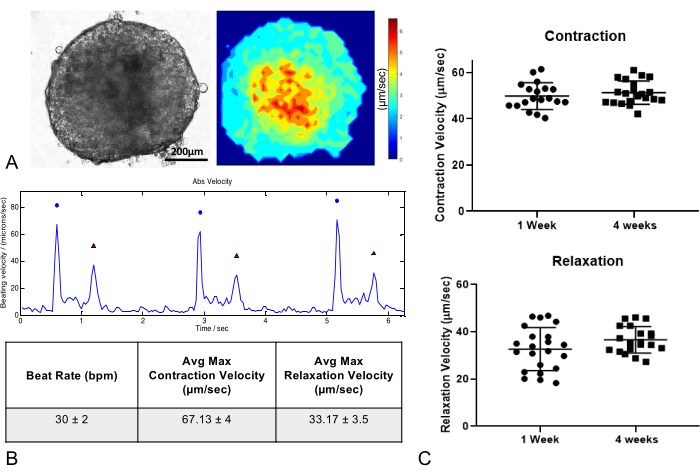
Figure 5: Contraction analysis of cardiac microtissues. (A) Phase contrast image and contraction map of cardiac microtissues 1 week after fabrication. Scale bar = 200 µm. (B) Cardiac microtissues show regular contraction and relaxation profiles and beat rates. Table shows representative values of beat rate, maximum contraction, and relaxation velocities. (C) Long-term culture of cardiac microtissues for up to 4 weeks does not significantly influence contractility parameters (n = 20/group). Please click here to view a larger version of this figure.
Discussion
To generate cardiac microtissues from pre-differentiated iPSC-CMs, iPSC-ECs, and iPSC-CFs, it is essential to obtain a highly pure culture for better control of cell numbers after contact-inhibited cell compaction within the cardiac microtissues. Recently, Giacomelli et. al.18 have demonstrated the fabrication of cardiac microtissues using iPSC-CMs, iPSC-ECs, and iPSC-CFs. Cardiac microtissues generated using the described method consist of ~5,000 cells (70% iPSC-CMs, 15% iPSC-ECs, and 15% iPSC-CFs). In this method, both cardiomyocytes and endothelial cells were co-differentiated followed by separation of endothelial progenitors using CD34+ marker. The current protocol described here consists of 12,000 cells (70% iPSC-CMs, 20% iPSC-ECs, and 10% iPSC-CFs) per microtissue, which yields higher cell numbers per construct for downstream single cell analyses. Furthermore, since all three cell types are differentiated separately there is limited heterogeneity in cell populations that is unique to understanding cell-specific responses to treatments.
For cardiomyocyte differentiation, we and others have previously demonstrated the derivation of pure cardiomyocytes using chemically defined culture conditions12,19,20. Briefly, differentiation begins with mesendoderm induction, followed by modulation of Wnt/β-catenin signaling that promotes cardiac lineage specification. Further purification of the cardiomyocytes in a glucose-deprived medium allows the elimination of non-cardiomyocytes. Here, it is important to note that prolonged culture in purification medium can hamper the quality of cardiomyocytes. A recently published protocol shows the proliferative capacity of early-stage cardiomyocytes can be harnessed to obtain large number of cells. The expansion of human iPSC-CMs is induced with reintroduction of CHIR, a potent mitogen during the early proliferative phase21. Although the precise molecular mechanism is still unknown, this technique can significantly improve iPSC-CM yield by ten-fold or hundred-fold. Furthermore, to improve the fidelity of iPSC-CMs for disease modeling, they can be cultured in a maturation medium to enhance electrophysiological, mechanical, and structural maturation22. A thorough overview of iPSC-CM maturation techniques is reviewed elsewhere23.
Vascular endothelial cells have been generated from pluripotent stem cells with different purification variable efficiencies13,24,25. The current protocol provides high differentiation efficiency and selection of phenotypically stable iPSC-ECs with MACS26. The differentiation methodology to generate functional cardiac fibroblasts follows modulation of the Wnt pathway and fibroblast growth factor (FGF) signaling to generate iPSC-CFs17. In vivo CFs originate from epicardium, endocardium, and neural crest progenitors16,27,28,29. Here, the CF lineage is generated from committed mesodermal cardiac progenitors without generating epicardial cells as an intermediate. Overall, the protocols described here to generate iPSC-CMs, iPSC-ECs, and iPSC-CFs take into account the ease of reproducibility and high purity based on phenotypic characteristics. To obtain high quality microtissues, it is important to obtain >90% purity for each individual cell type. Higher purity in the cellular phenotype will also ensure detection of changes in cellular trajectory due to different treatments.
After successful derivation of all three cell types, the cells are carefully dispensed into the agarose seeding chamber to allow settling of the cells in the circular recesses. Critical parameters include achieving homogenous single cell suspension and preventing cell aggregates or clumps that may cause variability in size distribution of the cardiac microtissues. From an overall structure perspective, 3D constructs are often limited by diffusion of nutrients, and as the cell density increases, the size and thickness of the constructs also increase linearly. Tissue engineered constructs in the form of spherical structures have a uniform nutrient consumption rate due to isotropic diffusivity and fixed distance from the core to the surface30,31. The final size of the microtissue dictates the concentration gradient of nutrients and oxygen diffusion to the center. In a static culture system, constructs over 350-400 µm may lead to a necrotic cell core when cultured over longer periods of time. Hence, careful consideration should be given to seeding density. A limitation of cardiac microtissue model is that it does not allow cardiomyocyte alignment offered by methodologies that involve geometric confinement of single cells. Hence, quantification of parameters such as sarcomere alignment or length remains limited due to random multidimensional cellular assembly. Despite the limitation, the technique allows rapid dose-dependent assessment of drug or small molecule toxicities on cardiovascular cells32. In a recent study, we employed 3D microtissues composed of iPSC-CMs to model secondary iron overload-induced cardiomyopathy, and we observed a significant reduction in microtissue size due to a high concentration of iron treatment33.
With regard to immunostaining, unlike 2D cultures, longer permeabilization and primary antibody incubation duration is necessary to allow diffusion of antibodies throughout the microtissues. The incubation time for adequate antibody penetration may require further optimization for microtissues larger than ~400 µm in diameter. Another important aspect is the longer blocking step with serum proteins to reduce background fluorescence resulting from non-specific binding. Typically, a blocking serum containing high IgG levels is preferred, such as goat or donkey serum. In order to maintain structural integrity of the cardiac microtissues, we have not probed for proteins involved in specific cell-cell interactions and their maintenance over the culture period. For cellular phenotyping, the microtissues must be digested over a short period of time to ensure high cell viability and preservation of target extracellular antigens. A combination of enzymatic digestion and mechanical disruption with a wide-bore pipette tip allows for an efficient and mild treatment of the microtissues during the digestion process. High quality viable single cells obtained through this rapid digestion protocol can be used to elucidate complex biological cell-cell interactions with the help of scRNA-seq34,35,36.
In addition to morphological and structural characterization, it is important to assess functional properties of the cardiac microtissues to assess the efficacy, toxicity, and disease state of the tissue assembly in vitro. Quantitative analysis of tissue contraction is a relevant parameter to assess cardiac function. Several non-invasive video microscopy techniques combined with motion tracking algorithms have enabled real-time monitoring with a robust measure of cardiac tissue contraction linked to phenotypic characteristics. To quantify the changes in phenotype, the cardiac microtissues can be maintained in vitro for up to 4 weeks without significant changes in contractile parameters. Most software algorithms work on the principles of optical-flow and vector mapping, which are superimposed over captured series of video frames11,37,38. The optical flow analysis may lead to the detection of artifacts; hence, the user should avoid or minimize inadvertent surrounding disturbances that could result in sample movement or vibrations transmitted to the stage of the microscope. It must be noted that the contractility analyses may not detect passive tension or loading effects due to suspended form of the microtissues, in contrast to microtissues formed between flexible posts of pre-defined stiffness. However, in both cases, it is important to note that the contraction profiles of engineered cardiac muscle tissues do not achieve contraction forces generated at an organ level. The main advantages of image-based assays are that they are non-destructive and allow measurement of acute and chronic drug exposure without extensive calibration. These tools can be applied over a variety of 2D and 3D platforms to measure changes in cardiac contractility due to treatments or underlying genetic diseases, and to evaluate tissue maturation strategies. Future investigation of this microtissue model may involve combining electrophysiological measurements that will allow simultaneous recording of calcium- or voltage-sensitive dyes to obtain multiple independent recordings of each microtissue in an array in a high-throughput manner.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We thank Dr. Amanda Chase for her helpful feedback on the manuscript. Funding support was provided by the Tobacco-Related Disease Research Program (TRDRP) of the University of California, T29FT0380 (D.T.) and 27IR-0012 (J.C.W.); American Heart Association 20POST35210896 (H.K.) and 17MERIT33610009 (J.C.W.); and National Institutes of Health (NIH) R01 HL126527, R01 HL123968, R01 HL150693, R01 HL141851, and NIH UH3 TR002588 (J.C.W).
Materials
| 12-well plates | Fisher Scientific | 08-772-29 | |
| 3D micro-molds | Microtissues | 12-81 format | |
| 6-well plates | Fisher Scientific | 08-772-1B | |
| AutoMACS Rinsing Solution | Thermo Fisher Scientific | NC9104697 | |
| B27 Supplement minus Insulin | Life Technologies | A1895601 | |
| B27 Supplement plus Insulin | Life Technologies | 17504-044 | |
| BD Cytofix | BD Biosciences | 554655 | |
| BD Matrigel, hESC-qualified matrix | BD Biosciences | 354277 | |
| Cardiac Troponin T Antibody | Miltenyi | 130-120-403 | |
| CD144 (VE-Cadherin) MicroBeads | Miltenyi | 130-097-857 | |
| CD31 Antibody | Miltenyi | 130-110-670 | |
| CD31 Microbeads | Miltenyi | 130-091-935 | |
| CHIR-99021 | Selleckchem | S2924 | |
| DDR2 | Santa Cruz Biotechnology | sc-81707 | |
| Dead Cell Apoptosis Kit with Annexin V FITC and PI | Thermo Fisher Scientific | V13242 | |
| Dispase I | Millipore Sigma | 4942086001 | |
| DMEM, high glucose (4.5g/L) no glutamine medium | 11960044 | ||
| DMEM/F-12 basal medium | Gibco | 11320033 | |
| Dulbecco's phosphate buffered saline (DPBS), no calcium, no magnesium | Life Technologies | 14190-136 | |
| EGM2 BulletKit | Lonza | CC-3124 | |
| Fetal bovine serum | Life Technologies | 10437 | |
| FibroLife Serum-Free Fibroblast LifeFactors Kit | LifeLIne Cell Technology | LS-1010 | |
| Glucose free RPMI medium | Life Technologies | 11879-020 | |
| Goat serum | Life Technologies | 16210-064 | |
| Human FGF-basic | Thermo Fisher Scientific | 13256029 | |
| Human VEGF-165 | PeproTech | 100-20 | |
| IWR-1-endo | Selleckchem | S7086 | |
| Liberase TL | Millipore Sigma | 5401020001 | |
| LS Sorting Columns | Miltenyi | 130-042-401 | |
| MACS BSA Stock solution | Miltenyi | 130-091-376 | |
| MACS Rinsing Buffer | Miltenyi | 130-091-222 | |
| MidiMACS Separator | Miltenyi | 130-042-302 | |
| RPMI medium | Life Technologies | 11835055 | |
| SB431542 | Selleckchem | S1067 | |
| TO-PRO 3 | Thermo Fisher Scientific | R37170 | |
| Triton X-100 | Millipore Sigma | X100-100ML | |
| TrypLE Select 10X | Thermo Fisher Scientific | red | |
| Vimentin Alexa Fluor® 488-conjugated Antibody | R&D Systems | IC2105G |
References
- Neofytou, E., O’Brien, C. G., Couture, L. A., Wu, J. C. Hurdles to clinical translation of human induced pluripotent stem cells. Journal of Clinical Investigation. 125 (7), 2551-2557 (2015).
- Lancaster, M. A., Knoblich, J. A. Organogenesis in a dish: Modeling development and disease using organoid technologies. Science. 345 (6194), 1247125 (2014).
- Liu, C., Oikonomopoulos, A., Sayed, N., Wu, J. C. Modeling human diseases with induced pluripotent stem cells: from 2D to 3D and beyond. Development. 145 (5), 156166 (2018).
- Yin, X., Mead, B. E., Safaee, H., Langer, R., Karp, J. M., Levy, O. Engineering stem cell organoids. Cell Stem Cell. 18 (1), 25-38 (2016).
- Giacomelli, E., et al. Three-dimensional cardiac microtissues composed of cardiomyocytes and endothelial cells co-differentiated from human pluripotent stem cells. Development. 144 (6), 1008-1017 (2017).
- Kurokawa, Y. K., George, S. C. Tissue engineering the cardiac microenvironment: Multicellular microphysiological systems for drug screening. Advances in Drug Delivery Reviews. 96, 225-233 (2016).
- Cartledge, J. E., et al. Functional crosstalk between cardiac fibroblasts and adult cardiomyocytes by soluble mediators. Cardiovascular Research. 105 (3), 260-270 (2015).
- Ravenscroft, S. M., Pointon, A., Williams, A. W., Cross, M. J., Sidaway, J. E. Cardiac non-myocyte cells show enhanced pharmacological function suggestive of contractile maturity in stem cell derived cardiomyocyte microtissues. Toxicology Science. 152 (1), 99-112 (2016).
- Ieda, M., et al. Cardiac fibroblasts regulate myocardial proliferation through beta1 integrin signaling. Developmental Cell. 16 (2), 233-244 (2009).
- Brutsaert, D. L. Cardiac endothelial-myocardial signaling: its role in cardiac growth, contractile performance, and rhythmicity. Physiological Reviews. 83 (1), 59-115 (2003).
- Huebsch, N., et al. Automated video-based analysis of contractility and calcium flux in human-induced pluripotent stem cell-derived cardiomyocytes cultured over different spatial scales. Tissue Engineering Part C Methods. 21 (5), 467-479 (2015).
- Burridge, P. W., et al. Chemically defined generation of human cardiomyocytes. Nature Methods. 11 (8), 855-860 (2014).
- Gu, M., et al. Pravastatin reverses obesity-induced dysfunction of induced pluripotent stem cell-derived endothelial cells via a nitric oxide-dependent mechanism. European Heart Journal. 36 (13), 806-816 (2015).
- Williams, I. M., Wu, J. C. Generation of endothelial cells from human pluripotent stem cells. Arteriosclerosis, Thrombosis and Vascular Biology. 39 (7), 1317-1329 (2019).
- Zhang, H., Shen, M., Wu, J. C. Generation of quiescent cardiac fibroblasts derived from human induced pluripotent stem cells. Methods in Molecular Biology. , 1-7 (2020).
- Zhang, H., et al. Generation of quiescent cardiac fibroblasts from human induced pluripotent stem cells for in vitro modeling of cardiac fibrosis. Circulation Research. 125 (5), 552-566 (2019).
- Zhang, J., et al. Functional cardiac fibroblasts derived from human pluripotent stem cells via second heart field progenitors. Nature Communications. 10 (1), 2238-2315 (2019).
- Giacomelli, E., et al. Human-iPSC-derived cardiac stromal cells enhance maturation in 3d cardiac microtissues and reveal non-cardiomyocyte contributions to heart disease. Cell Stem Cell. 26 (6), 862-879 (2020).
- Burridge, P. W., Holmström, A., Wu, J. C. Chemically defined culture and cardiomyocyte differentiation of human pluripotent stem cells. Current Protocols in Human Genetics. 87 (1), 1-15 (2015).
- Lian, X., et al. Directed cardiomyocyte differentiation from human pluripotent stem cells by modulating Wnt/β-catenin signaling under fully defined conditions. Nature Protocols. 8 (1), 162-175 (2013).
- Buikema, J. W., et al. Wnt activation and reduced cell-cell contact synergistically induce massive expansion of functional human iPSC-derived cardiomyocytes. Cell Stem Cell. 27 (1), 50-63 (2020).
- Feyen, D. A. M., et al. Metabolic maturation media improve physiological function of human iPSC-derived cardiomyocytes. Cell Reports. 32 (3), 107925 (2020).
- Karbassi, E., et al. Cardiomyocyte maturation: Advances in knowledge and implications for regenerative medicine. Nature Reviews Cardiology. 17 (6), 341-359 (2020).
- Li, Z., Hu, S., Ghosh, Z., Han, Z., Wu, J. C. Functional characterization and expression profiling of human induced pluripotent stem cell- and embryonic stem cell-derived endothelial cells. Stem Cells Development. 20 (10), 1701-1710 (2011).
- Lian, X., et al. Efficient differentiation of human pluripotent stem cells to endothelial progenitors via small-molecule activation of WNT signaling. Stem Cell Report. 3 (5), 804-816 (2014).
- Gu, M. Efficient differentiation of human pluripotent stem cells to endothelial cells. Current Protocols in Human Genetics. 98 (1), 64 (2018).
- Acharya, A., et al. The bHLH transcription factor Tcf21 is required for lineage-specific EMT of cardiac fibroblast progenitors. Development. 139 (12), 2139-2149 (2012).
- Wessels, A., et al. Epicardially derived fibroblasts preferentially contribute to the parietal leaflets of the atrioventricular valves in the murine heart. Development Biology. 366 (2), 111-124 (2012).
- Ali, S. R., et al. Developmental heterogeneity of cardiac fibroblasts does not predict pathological proliferation and activation. Circulation Research. 115 (7), 625-635 (2014).
- McMurtrey, R. J. Analytic and numerical models of oxygen and nutrient diffusion, metabolism dynamics, and architecture optimization in three-dimensional tissue constructs with applications and insights in cerebral organoids. Tissue Engineering Part C Methods. (3), 221-249 (2015).
- Thomas, D., O’Brien, T., Pandit, A. Toward customized extracellular niche engineering: progress in cell-entrapment technologies. Advanced Materials. 30 (1), 1703948 (2018).
- Thomas, D., Shenoy, S., Sayed, N. Building Multi-dimensional induced pluripotent stem cells-based model platforms to assess cardiotoxicity in cancer therapies. Front Pharmacol. 12, 39 (2021).
- Rhee, J. -. W., et al. Modeling secondary iron overload cardiomyopathy with human induced pluripotent stem cell-derived cardiomyocytes. Cell Reports. 32 (2), 107886 (2020).
- Paik, D. T., Cho, S., Tian, L., Chang, H. Y., Wu, J. C. Single-cell RNA sequencing in cardiovascular development, disease, and medicine. Nature Reviews Cardiology. 17 (8), 457-473 (2020).
- Nguyen, Q. H., Pervolarakis, N., Nee, K., Kessenbrock, K. Experimental considerations for single-cell RNA sequencing approaches. Frontiers in Cell and Developmental Biology. 6, 108 (2018).
- Lau, E., Paik, D. T., Wu, J. C. Systems-wide approaches in induced pluripotent stem cell models. Annual Reviews in Pathology. 14 (1), 395-419 (2019).
- Maddah, M., et al. A non-invasive platform for functional characterization of stem-cell-derived cardiomyocytes with applications in cardiotoxicity testing. Stem Cell Report. 4 (4), 621-631 (2015).
- Sala, L., et al. Musclemotion: A versatile open software tool to quantify cardiomyocyte and cardiac muscle contraction in vitro and in vivo. Circulation Research. 122 (3), 5-16 (2018).

