Evaluation of Antimicrobial Activities of Nanoparticles and Nanostructured Surfaces In Vitro
Summary
We introduce four methods to evaluate the antimicrobial activities of nanoparticles and nanostructured surfaces using in vitro techniques. These methods can be adapted to study the interactions of different nanoparticles and nanostructured surfaces with a broad range of microbial species.
Abstract
The antimicrobial activities of nanoparticles and nanostructured surfaces, such as silver, zinc oxide, titanium dioxide, and magnesium oxide, have been explored previously in clinical and environmental settings and in consumable food products. However, a lack of consistency in the experimental methods and materials used has culminated in conflicting results, even amongst studies of the same nanostructure types and bacterial species. For researchers who wish to employ nanostructures as an additive or coating in a product design, these conflicting data limit their utilization in clinical settings.
To confront this dilemma, in this article, we present four different methods to determine the antimicrobial activities of nanoparticles and nanostructured surfaces, and discuss their applicability in different scenarios. Adapting consistent methods is expected to lead to reproducible data that can be compared across studies and implemented for different nanostructure types and microbial species. We introduce two methods to determine the antimicrobial activities of nanoparticles and two methods for the antimicrobial activities of nanostructured surfaces.
For nanoparticles, the direct co-culture method can be used to determine the minimum inhibitory and minimum bactericidal concentrations of nanoparticles, and the direct exposure culture method can be used to assess real-time bacteriostatic versus bactericidal activity resulting from nanoparticle exposure. For nanostructured surfaces, the direct culture method is used to determine the viability of bacteria indirectly and directly in contact with nanostructured surfaces, and the focused-contact exposure method is used to examine antimicrobial activity on a specific area of a nanostructured surface. We discuss key experimental variables to consider for in vitro study design when determining the antimicrobial properties of nanoparticles and nanostructured surfaces. All these methods are relatively low cost, employ techniques that are relatively easy to master and repeatable for consistency, and are applicable to a broad range of nanostructure types and microbial species.
Introduction
In the US alone, 1.7 million individuals develop a hospital-acquired infection (HAI) annually, with one in every 17 of these infections resulting in death1. In addition, it is estimated that the treatment costs for HAIs range from $28 billion to $45 billion annually1,2. These HAIs are predominated by methicillin-resistant Staphylococcus aureus (MRSA)3,4 and Pseudomonas aeruginosa4, which are commonly isolated from chronic wound infections and usually require extensive treatment and time to produce a favorable patient outcome.
Over the past several decades, multiple antibiotic classes have been developed to treat infections related to these and other pathogenic bacteria. For example, rifamycin analogs have been used to treat MRSA, other gram-positive and gram-negative infections, and Mycobacterium spp. infections5. In the 1990s, to effectively treat an increasing number of M. tuberculosis infections, additional drugs were combined with rifamycin analogs to increase their effectiveness. However, approximately 5% of M. tuberculosis cases remain resistant torifampicin5,6, and there is increasing concern regarding multi-drug resistant bacteria7. Currently, the use of antibiotics alone may not be sufficient in the treatment of HAIs, and this has provoked an ongoing search for alternative antimicrobial therapies1.
Heavy metals, such as silver (Ag)8,9,10 and gold (Au)11, and ceramics, such as titanium dioxide (TiO2)12 and zinc oxide (ZnO)13, in nanoparticle (NP) form (AgNP, AuNP, TiO2NP, and ZnONP, respectively) have been examined for their antimicrobial activities and have been identified as potential antibiotic alternatives. In addition, bioresorbable materials, such as magnesium alloys (Mg alloys)14,15,16, magnesium oxide nanoparticles17,18,19,20,21, and magnesium hydroxide nanoparticles [nMgO and nMg(OH)2, respectively]22,23,24, have also been examined. However, the previous antimicrobial studies of nanoparticles used inconsistent materials and research methods, resulting in data that are difficult or impossible to compare and are sometimes contradictory in nature18,19. For example, the minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) of silver nanoparticles varied significantly in different studies. Ipe et al.25 evaluated the antibacterial activities of AgNPs with an average particle size of ~26 nm to determine the MICs against gram-positive and gram-negative bacteria. The identified MICs for P. aeruginosa, E. coli, S. aureus, and MRSA were 2 µg/mL, 5 µg/mL, 10 µg/mL, and 10 µg/mL, respectively. In contrast, Parvekar et al.26 evaluated AgNPs with an average particle size of 5 nm. In this instance, the AgNP MIC and a MBC of 0.625 mg/mL were found to be effective against S. aureus. In addition, Loo et al.27 evaluated AgNPs with a size of 4.06 nm. When E. coli was exposed to these nanoparticles, the MIC and MBC were reported at 7.8 µg/mL. Finally, Ali et al.28 investigated the antibacterial properties of spherical AgNPs with an average size of 18 nm. When P. aeruginosa, E. coli, and MRSA were exposed to these nanoparticles, the MIC was identified at 27 µg/mL, 36 µg/mL, 27 µg/mL, and 36 µg/mL, respectively, and the MBC was identified at 36 µg/mL, 42 µg/mL, and 30 µg/mL, respectively.
Although the antibacterial activity of nanoparticles has been extensively studied and reported during recent decades, there is no standard for the materials and research methods used to allow for direct comparisons across studies. For this reason, we present two methods, the direct co-culture method (method A), and the direct exposure method (method B), to characterize and compare the antimicrobial activities of nanoparticles while keeping the materials and methods consistent.
In addition to nanoparticles, nanostructured surfaces have also been examined for antibacterial activities. These include carbon-based materials, such as graphene nanosheets, carbon nanotubes, and graphite29, as well as pure Mg and Mg alloys. Each of these materials has exhibited at least one antibacterial mechanism, including physical damage imposed on cell membranes by carbon-based materials and damage to metabolic processes or DNA through the release of reactive oxygen species (ROS) when Mg degrades. In addition, when zinc (Zn) and calcium (Ca) are combined in the formation of Mg alloys, the refinement of the Mg matrix grain size is enhanced, which leads to a reduction in bacterial adhesion to substrate surfaces in comparison to Mg-only samples14. To demonstrate antibacterial activity, we present the direct culture method (method C), which determines bacterial adhesion on and around nanostructured materials over time through the quantification of bacterial colony-forming units (CFUs) with direct and indirect surface contact.
The geometry of nanostructures on surfaces, including the size, shape, and orientation, could influence the bactericidal activities of materials. For example, Lin et al.16 fabricated different nanostructured MgO layers on the surfaces of Mg substrates through anodization and electrophoretic deposition (EPD). After a period of exposure to the nanostructured surface in vitro, the growth of S. aureus was substantially reduced in comparison to non-treated Mg. This indicated a greater potency of the nanostructured surface against bacterial adhesion versus the nontreated metallic Mg surface. To reveal the different mechanisms of the antibacterial properties of various nanostructured surfaces, a focused-contact exposure method (method D) that determines the cell-surface interactions within the area of interest is discussed in this article.
The objective of this article is to present four in vitro methods that are applicable to different nanoparticles, nanostructured surfaces, and microbial species. We discuss key considerations for each method to produce consistent, reproducible data for comparability. Specifically, the direct co-culture method17 and direct exposure method are used for examining the antimicrobial properties of nanoparticles. Through the direct co-culture method, the minimum inhibitory and minimum bactericidal concentrations (MIC and MBC90–99.99, respectively) can be determined for individual species, and the most potent concentration (MPC) can be determined for multiple species. Through the direct exposure method, the bacteriostatic or bactericidal effects of nanoparticles at minimum inhibitory concentrations can be characterized by real-time optical density readings over time. The direct culture14 method is suitable for examining bacteria directly and indirectly in contact with nanostructured surfaces. Finally, the focused-contact exposure16 method is presented to examine the antibacterial activity of a specific area on a nanostructured surface through the direct application of bacteria and the characterization of bacterial growth at the cell-nanostructure interface. This method is modified from the Japanese Industrial Standard JIS Z 2801:200016, and is intended to focus on microbe-surface interactions and exclude the effects of bulk sample degradation in microbial culture on antimicrobial activities.
Protocol
To present the direct co-culture and direct exposure methods, we use magnesium oxide nanoparticles (nMgO) as a model material to demonstrate bacterial interactions. To present the direct culture and focused-contact exposure methods, we use an Mg alloy with nanostructured surfaces as examples.
1. Sterilization of nanomaterials
NOTE: All the nanomaterials must be sterilized or disinfected prior to microbial culture. The methods that can be used include heat, pressure, radiation, and disinfectants, but the tolerance of the materials for each method must be identified prior to the in vitro experiments.
- Nanoparticles
NOTE: Samples can be stored in an organic solvent such as ethanol or packaged into a dish or box followed by sterilization or disinfection in an appropriate manner. The nanoparticles were sterilized using the following method for demonstration of the direct co-culture method17 and direct exposure method.- Sterilize MgO nanoparticles in a 200 °C convection oven30 for 60 min prior to each in vitro experiment.
NOTE: This method was chosen because water steam and UV light may affect the structures of the MgO nanoparticles (nMgO)17.
- Sterilize MgO nanoparticles in a 200 °C convection oven30 for 60 min prior to each in vitro experiment.
- Bulk materials
NOTE: The nanostructured materials were sterilized by the following method for demonstration of the direct culture method.- For the direct culture method14, disinfect as-prepared ZC21, Mg, and T64 samples using ultraviolet (UV) radiation for 4 h before initiating the in vitro studies.
- For the focused-contact exposure method16, disinfect all the samples using UV radiation for 2 h before initiation of the in vitro studies.
- Alternatively, use ethylene oxide (EtO) for the sterilization of heat-sensitive materials.
2. Direct co-culture method (method A)
NOTE: In method A, bacteria in a lag-phase seeding culture are directly mixed with nanoparticles of certain concentrations. For the examination of nanoparticle antimicrobial activities, we follow a protocol described by Nguyen et al.17.
- Characterization of nanoparticles and nanostructures
NOTE: Nanostructure composition and shape are confirmed using X-ray powder diffraction.- Measurement of nanoparticles
- Weigh the nanoparticles at 9x the desired mass per mL to accommodate 3 mL samples in triplicate. For example, weigh 0.2 mg/mL nMgO at 1.8 mg/mL.
NOTE: This ensures the consistent distribution of nanoparticles in the bacterial cultures and broths. - Measure all the nanoparticles in a 5.0 mL microcentrifuge tube that has been pre-weighed and tared using an analytical balance.
- Weigh the nanoparticles at 9x the desired mass per mL to accommodate 3 mL samples in triplicate. For example, weigh 0.2 mg/mL nMgO at 1.8 mg/mL.
- Measurement of nanoparticles
- Preparing and growing bacteria cultures
- For each in vitro experiment, retrieve bacteria cell stocks from storage at -80 °C. Add 10 µL of each bacteria cell stock into 5 mL of an appropriate growth medium.
- Place the inoculated medium into an incubator-shaker at 37 °C and 250 rpm for approximately 16 h overnight.
- If Staphylococcus species are grown, subculture by placing 400 µL of each overnight culture into 20 mL of fresh growth medium (100 µL/5 mL medium) and incubate with shaking at 37 °C and 250 rpm for an additional 4-6 h.
- Washing and counting the bacteria cells to determine the seeding density
NOTE: The desired seeding density of 7.8 × 106 CFU/mL was identified as the cell number greater than what is required to confirm a urinary tract infection.- Collect cultured bacteria by aliquoting 1 mL of the overnight bacteria culture into 1.5 mL microcentrifuge tubes. Create sufficient numbers of aliquots from overnight bacterial cultures and centrifuge them for 10 min at 1,956 × g to reach the desired seeding densities.
- After centrifugation, pipet to remove the supernatant from the pelleted cells and place the supernatant into a collection receptacle. Resuspend the cell pellets in 0.5 mL of fresh growth medium. Combine two 0.5 mL suspensions to create 1 mL of suspension to reduce the number of 1 mL aliquots of resuspended cells to six. Repeat the centrifugation for 10 min at 1,956 × g.
- Complete a second wash cycle using fresh growth medium, as in step 2.3.2, to reduce the number of 1 mL aliquots of resuspended washed cells to three.
- Complete the third cycle, as in step 2.3.2, except resuspend the cell pellets in 0.33 mL of Tris buffer (hydroxymethyl) aminomethane buffer (Tris buffer, pH 8.5). Combine the three suspensions, each of 0.33 mL volume, into one 1.5 mL microcentrifuge. Repeat the centrifugation for 10 min at 1,956 × g.
NOTE: Tris buffer does not contain Mg2+ or Ca2+ ions31. This is important for the use of inductively coupled plasma-optical emission spectrometry (ICP-OES) to measure Mg2+ and Ca2+ ions in post-culture broths. In contrast, the phosphates present in phosphate-buffered saline (PBS)32, for example, sequester both Mg2+ or Ca2+ ions33,34,35 and cause confusion in the interpretation of the ICP-OES data. - Remove the supernatant from the pelleted cells. Resuspend the cell pellet in 1 mL of fresh Tris buffer, known as the cell suspension-a lag-phase bacterial seeding culture.
- Determine the cell suspension concentration (cells/mL) using a hemocytometer.
- Creation of the seeding bacteria culture
NOTE: The sample has a total volume of 3 mL and is completed in triplicate (9 mL total).- Determine the total volume of bacteria seeding culture needed. To create the seeding culture, use C1V1 = C2V2 to calculate the volume of the cell suspension needed to create a seeding culture of 7.8 × 106 cells/mL. Add the calculated volume of the cell suspension (V1) to the required volume of growth media (V2).
- Determine the actual bacteria seeding density in CFU/mL. Create a tenfold serial dilution to 10-4. Spread-plate the 10-4 dilution onto the appropriate growth agar and incubate overnight. After incubation, count the number of colonies, and calculate the CFU/mL.
- Formation of bacteria and nanoparticle cultures
- In 12-well, non-tissue treated polystyrene plates, aliquot 2 mL of the bacterial seeding culture in each well for the number of wells needed.
- For each 5 mL microcentrifuge tube containing pre-weighed nMgO, add 3 mL of the bacterial seeding culture. Vortex briefly to mix the nMgO with the bacteria. Aliquot 1 mL of the mixture into three separate wells to create the triplicate samples of each pre-measured weight of nMgO. Pipet the bacteria/nMgO mixture 2-3x before each 1 mL aliquot is made to maintain nMgO in suspension.
NOTE: Control samples will consist of 3 mL of cells only, 3 mL of media only, and 3 mL containing the lowest and highest concentrations of nanoparticles used. These are prepared as in step 2.5.2. All the control samples are completed in triplicate. - Incubate all the 12-well plates at 37 °C and 120 rpm for 24 h.
- Determining the bacterial cell growth after exposure to nMgO
- After overnight incubation of the bacterial cultures, collect every 3 mL sample by pipetting into an individual 15 mL conical tube.
- Use a 96-well plate to create 1:10 serial dilutions. Determine the number of columns necessary to serially dilute all the samples containing bacteria. Add 180 µL of Tris buffer to each well in row B to row G for the appropriate number of columns.
- Briefly vortex each 15 mL conical tube containing bacterial samples, and add 50 µL to an individual well in row A.
- For each well in row A, transfer 20 µL into row B (e.g., A1 to B1), and briefly mix by pipetting to obtain a volume of 200 µL. Using a sterile pipet tip, transfer 20 µL from the well in row B to the well in row C. Continue this pattern until the well in row G contains 200 µL, completing a serial dilution from 10−1 in row B to 10−6 in row G.
- Pipet 100 µL of each well onto an appropriate growth agar plate and spread across the plate to disperse the cell culture. Place the plates into an incubator-shaker overnight at 37 °C. Incubate the plates at 37 °C for 24 h.
- Examine the plates and count those that have approximately 25-300 colonies. If possible, choose plates of the same dilution value. Use the number of colonies per plate to calculate the CFU/mL.
- Evaluation of the post-incubation pH
NOTE: Follow the manufacturer's instructions for each pH meter model.- Pre-calibrate a pH meter using standardizing solutions of pH 4, pH 7, and pH 10. Read each bacterial sample by placing the pH probe into the 15 mL conical tube.
- Preparation of samples for ICP-OES
NOTE: ICP-OES is used to determine the concentration of Mg2+ and Ca2+ ions. These cations are considered important, as each participates in cellular metabolism.- Centrifuge each 15 mL conical tube prepared in step 2.6.1 for 5 min at 5,724 × g to pellet the cellular and nanoparticle debris.
- Using a 15 mL conical tube, dilute 30 µL of the supernatant into 2.97 mL of 18.2 Ω filtered water to create a 1:100 dilution.
NOTE: The dilution factor can be adjusted based on the projected ion concentrations and the detection limit of the spectrometer instrument. - Read the samples using ICP-OES.
3. Direct exposure method (method B)
NOTE: If the growth rate of the chosen bacteria is unknown, then a standardization of growth curve must be completed prior to implementing this method.
- Determine the real-time bacteriostatic and bactericidal activities upon exposure to the nanoparticles of interest.
- Prepare the desired concentrations of nanoparticles using the methods described in step 2.1-2.1.2.2. Prepare two sets of each weight of nanoparticles to be used: one to be added to 3 mL aliquots of bacteria culture in the logarithmic growth phase, and the other to be added to 3 mL aliquots of growth medium without bacteria as control groups.
- Determine the appropriate bacteriostatic and bactericidal antibiotics for the bacteria species to be tested.
NOTE: Knowledge of the minimum inhibitory concentration for each antibiotic is required.
- Calculate the total volume of bacterial culture needed for triplicate 3 mL samples for all the nanoparticle, antibiotic, and control samples.
NOTE: An equal volume of sterile growth medium is required. If the use of spectrophotometry is planned, this requires adjustments to be made to the total volume needed for accommodation of the 0.5 mL to 1.0 mL volume needed for cuvettes. - Day 1: For each in vitro experiment, create overnight stocks following step 2.2.1-2.2.2.
- Day 2: Confirm the plate reader settings can accommodate a 96-well plate with an optical density of 600 nm. Measure the samples in 200 µL aliquots using a 96-well plate.
- Determine an initial bacterial culture OD600 at 0.01-the density used to begin the initial incubation period prior to the addition of nanoparticles and antibiotics.
- Collect the overnight bacteria culture from the incubator-shaker.
- For each material (e.g., pre-measured nanoparticles or antibiotics) to be tested, create a separate bacterial culture at an OD600 of 0.01. Use a container large enough to accommodate the volume of cell culture needed (e.g., a sterilized Erlenmeyer flask or 50 mL conical tubes).
- Dilute the overnight bacterial samples with growth medium by placing three 200 µL aliquots of the growth medium into three wells and three 200 µL aliquots of the bacterial culture into additional wells (e.g., A1 through A6).
- Place the 96-well plate into the plate reader and scan it.
NOTE: The readings are averaged for each scanned well to produce a mean value. - To calculate the density of bacteria in suspension, average the mean value for each well containing a triplicate sample.
- To determine the optical density of the bacteria, subtract the broth sample average from the bacterial average.
- If it is necessary to continue to adjust the bacterial suspension, continue to add broth or overnight bacterial culture as needed, and repeat scanning in the plate reader as necessary until an OD600 of approximately 0.01 is reached.
- Incubate at 37 °C with shaking at 150 rpm until logarithmic growth is reached.
- Remove the bacterial cultures from the incubator shaker.
- Aliquot 3 mL of the OD600 0.01 bacterial culture into labeled 15 mL conical tubes for test and control samples in triplicate.
- Aliquot 200 µL of the bacterial culture into three wells of a 96-well plate and measure the samples using the plate reader.
- Calculate the readings as described previously in step 3.4.2.2 and step 3.4.2.6-the "-0.5 h reading".
- Creation and measurement of bacteria/nanoparticle mixtures
- Aliquot 3 mL of sterile growth medium in equal numbers to the bacterial cultures into labeled 15 mL conical tubes for control samples (in triplicate).
- To prepare bacteria/nanoparticle suspensions and sterile media/nanoparticle suspensions, remove 1 mL of bacterial culture or medium from each triplicate set of 15 mL conical tubes.
- Place the 1 mL aliquots into a 5 mL centrifuge tube containing pre-measured nanoparticles, and vortex briefly to mix.
- From the 5 mL centrifuge tube, return 1 mL aliquots of the bacteria/nanoparticle or medium/nanoparticle suspensions to the 15 mL conical tube to distribute the nanoparticles homogeneously.
NOTE: Pipet the bacteria/nanoparticle mixture 2x-3x before each 1 mL aliquot is pipetted to maintain the nanoparticles in suspension.
- Creation and measurement of bacteria/antibiotic mixtures
- To create the bacterial/antibiotic cultures, aliquot 3 mL of the incubated bacterial culture into the correspondingly labeled 15 mL conical tubes.
- After all the samples have been prepared, aliquot 200 µL of each sample into the individual wells of a 96-well plate.
- Place the plate(s) into the plate reader, and initiate scanning-the "0 h" reading.
- Place the 15 mL conical tubes containing the samples into an incubator-shaker at 37 °C and 150 rpm. Record all the data as described previously in step 3.4.2.2 and step 3.4.2.6.
- Approximately 15 min after completing the 0 h reading, repeat the steps described in step 3.7.2-3.7.3-the "0.5 h" reading.
- After the established 0 h, repeat the steps described in step 3.7.2-step 3.7.3 every 90 min for six cycles; complete in 9 h.
- After the sixth cycle has been completed, place the samples in the incubator-shaker for an additional 15 h at 37 °C and 150 rpm. Repeat the steps described in step 3.7.2-3.7.3 to obtain the 24 h reading. Record all the data as described previously in step 3.4.2.2 and step 3.4.2.6.
4. Direct culture method (method C)
NOTE: In method C, bacteria in a lag-phase seeding culture are placed directly on the nanostructured surfaces of interest. For examination of the nanostructure antimicrobial activities, we follow a protocol described by Zhang et al.14. To demonstrate this direct culture method, ZC21 (Mg-Zn-Ca Alloy) and Mg pins were used as samples.
- Preparing and growing the bacteria cultures
- For each in vitro experiment, create overnight stocks following step 2.2.1-2.2.2.
- Washing and counting the bacteria to determine the seeding density
NOTE: The desired seeding density of 7.5 × 105 CFU/mL was identified as the reported cell concentration that causes orthopedic infections14.- Retrieve the overnight bacterial culture from the incubator-shaker.
- Wash and collect the bacteria, following step 2.3.2-2.3.2.3, but replace the fresh medium with revised simulated body fluid (rSBF) for each washing step.
NOTE: rSBF is a buffer solution that mimics the ionic concentration of human blood plasma. However, rSBF contains only inorganic compounds, and is without any bioorganic compounds such as proteins. In order to resolve this, fetal bovine serum (FBS) is combined with rSBF to simulate the composition of human blood to better mimic the natural formation of apatite in calcified tissues under experimental conditions36. - Remove the supernatant from the pelleted cells. Resuspend the cell pellet in 1 mL of rSBF supplemented with 10% FBS-the cell suspension.
- Determine the cell suspension concentration (cells/mL) using a hemocytometer. Dilute the cell suspension to a concentration of 7.5 × 105 cells/mL in rSBF supplemented with 10% FBS. Create a seeding culture following step 2.4.1.
- Determine the actual seeding density following step 2.4.2.
- Distribute bacterial cell suspension onto the samples in the culture wells.
- Place both the samples and control samples into the individual wells of a 48-well, non-tissue treated polystyrene plate.
- Aliquot 0.75 mL of the cell suspension into each well containing the samples and controls. Place the 48-well plate(s) into an incubator-shaker for 24 h at 37 °C with shaking at 120 rpm.
- Characterization of the bacterial concentration
- After 24 h of incubation, collect the samples and place them into separate, labeled collection tubes.
- Collect two samples from each group and place them individually into labeled 5.0 mL microcentrifuge tubes. Add 2 mL of rSBF to each tube.
- Place the samples collected in step 4.4.2 into a sonication bath, and sonicate for 10 min. After each 5 min period, vortex each sample for 5 s.
- Collect the rSBF containing the newly detached bacterial cells and place the suspension into a labeled, fresh microcentrifuge tube.
- Serial dilutions and plating the bacteria
- Serially dilute and plate the bacterial suspension following step 2.6.2-2.6.6.
- Evaluate the post-incubation pH of culture media by following step 2.7.
- Prepare the post-culture medium for ICP-OES by following the methods described in step 2.8.
5. Focused-contact exposure method (method D)
NOTE: In method D, bacteria on a nitrocellulose filter paper are put in direct contact with an area of interest on the nanostructured surfaces. This method minimizes the interference of bulk sample degradation in bacterial cultures with the bacterial activities. To examine nanosurface antimicrobial activities, we follow a protocol described by Lin et al.16.
- Follow the procedures in step 2.2.1-2.2.2.1 to create a bacterial seeding culture. Confirm the actual seeding density following step 2.4.2.
- Prepare the samples to a size of 1 × 1 cm2 in a square shape. Prepare all the sample types in triplicate. If necessary, place the samples on a three-dimensional (3D) holder that is 15 mm in diameter by 10 mm in height. Place the sample and holder into a well of a non-tissue culture-treated polystyrene well plate.
- Prepare sterilized nitrocellulose papers by trimming each to a diameter of 1 cm. Place the prepared nitrocellulose papers onto an agar plate containing the appropriate medium.
- Pipet 50 µL of the diluted bacterial culture onto the filter paper.
- Pipet 50 µL of an appropriate medium onto the center of each sample surface.
- Using sterilized tweezers, pick up the nitrocellulose paper from the surface of the agar. Carefully flip the nitrocellulose paper and place it onto the sample surface so that the bacteria are in contact with the 50 µL of medium and the nanostructured surface of interest.
- Add 1 mL of Tris buffer to each well containing a sample to maintain the humidity. Incubate the polystyrene well plate containing all the samples at 37 °C for 24 h.
- Collect the nitrocellulose paper from each sample surface. Place each into 5 mL of Tris buffer. Vortex the collected filter papers and nanosurfaced material samples for 5 s.
- Sonicate each sample for 10 min. Vortex for 5 s after 5 min and again after 10 min.
- Collect the Tris buffer suspensions from all the samples and place each volume into individual fresh collection tubes.
- Serially dilute and plate the samples following step 2.6.2-2.6.6.
6. Post-culture characterization of bacteria and nanomaterials
- Examination of bacterial adhesion and morphology using scanning electron microscopy (SEM)
- After overnight incubation, add 10% glutaraldehyde into each well of the 48-well, non-tissue treated polystyrene plate until the samples are fully covered. Incubate the samples for 1 h to fix the bacterial cells.
- Aspirate the glutaraldehyde into a waste bottle and rinse all the samples 3x with Tris buffer solution to remove non-adherent bacteria.
- Dehydrate the samples using 30%, 75%, and 100% ethanol for 30 min each16.
NOTE: It is recommended to use a critical point dryer to dry the samples. It is not ideal to air-dry the samples at room temperature for 24 h. - Use conductive adhesive tapes, such as copper or carbon tapes, to fix the samples onto an SEM stub. Coat the sample surfaces via sputter coating to provide conductivity prior to the imaging (e.g., coat Mg plates with adhered bacteria under platinum/palladium (Pt/Pd) for 45 s at 20 mA16).
NOTE: The coating materials, processing time, and operation current may vary depending on the sample types. - Take images of the bacteria on the samples using SEM with an appropriate working distance and accelerating voltage and at the desired magnifications. For example, use an SEM with a secondary electron detector to take images at a working distance of 5 mm and an accelerating voltage of 10 kV16.
- Examine surface morphology of post-culture nanomaterial samples using SEM.
- For nanomaterials, wash the samples using Tris buffer 3x to remove the free bacteria that are not attached to the sample. For nanoparticles, disperse the particles into a solvent that does not affect the sample properties through ultrasonication to achieve better dispersion when needed.
- Dry the samples after the washing procedure.
- Use carbon or copper double-sided adhesive tapes to adhere the samples onto the SEM stub.
- Create a conductive surface layer of Pt/Pd using a sputter coater, as described in step 6.1.4, prior to the imaging. Take the sample images as described in step 6.1.5.
- Analyze the surface elemental composition using EDS with an appropriate accelerating voltage and at the desired magnifications (e.g., at 10 kV after performing the SEM analysis16).
- Fluorescence imaging of bacterial adhesion
- Prepare the samples to be visualized using fluorescence microscopy by first washing them with Tris buffer 3x. Air-dry the samples at room temperature.
- Stain each sample with 10 µM thioflavin T fluorescence dye following the established protocol37.
- Perform fluorescence imaging of bacterial adhesion using an inverted microscope coupled with an electron-multiplying charge-coupled device digital camera.
Representative Results
The identification of the antibacterial activity of magnesium oxide nanoparticles and nanostructured surfaces has been presented using four in vitro methods that are applicable across different material types and microbial species.
Method A and method B examine bacterial activities when exposed to nanoparticles at a lag phase (method A) and log phase (method B) for a duration of 24 h or longer. Method A provides results regarding the MIC and MBC, while method B determines the inhibitory versus bactericidal effects of nanoparticles. Method C examines the bacterial activities with direct and indirect contact with nanostructured surfaces, and method D examines the bacterial activities on a select area of a cell-nanostructure interface for a duration of 24 h or longer.
The methods used are described in Figure 1 and Figure 2, and their results are presented in Figure 3, Figure 4, Figure 5, Figure 6, Figure 7, and Figure 8. Representative experimental results that quantify antimicrobial activity of nMgO against gram-negative and gram-positive bacteria and yeast can be viewed in Figure 3. The antibacterial activities of nMgO and nMg(OH)2 against MRSA can be viewed in Figure 4. The antibacterial activities of nanostructured surfaces against MRSA can be viewed in Figure 5 and Figure 6. Finally, the antibacterial activity of magnesium alloys against E. coli can be viewed in Figure 8B.
Using method A, it is possible to determine the MICs and MBCs for bacteria exposed to nanoparticles using consistent methods and materials. This consistency allows comparisons to be made between species using the identified MBCs to determine the most potent concentration of nanoparticles tested. In addition, this method can be applied to other classifications of microorganisms for comparisons of MICs and minimal lethal concentrations (MLCs17) as well. Here, sterilized nanoparticles were pre-measured at a quantity allowing triplicates of each concentration required. These nanoparticles were suspended in a lag-phase mono-bacteria seeding culture with a density of 6 × 106-8 × 106 cells/mL. The method described to suspend the pre-measured nanoparticles with seeding culture or broth successfully created triplicate samples that were relatively homologous in nanoparticle distribution, which may reduce the deviation in CFU/mL within triplicate samples. The experimental workflow for this method is illustrated in Figure 1A. A demonstration of the antimicrobial effects of nMgO on gram-negative and gram-positive bacteria and Candida spp. using method A can be seen in Figure 3. Here, an MIC of 1.0 mg/mL nMgO for gram-negative Escherichia coli and Pseudomonas aeruginosa, as well as an MBC99.99 of 1.0 mg/mL and 1.6 mg/mL nMgO, respectively, were identified for these species (Figure 3A). Gram-positive S. epidermidis, S. aureus, and MRSA demonstrated MICs of 0.5 mg/mL, 0.7 mg/mL, and 1.0 mg/mL nMgO, respectively. MBC99.99 values of 1.6 mg/mL and 1.2 mg/mL nMgO were identified for S. epidermidis and S. aureus, respectively, while MRSA was not reduced beyond MBC90 (Figure 3B). In drug-sensitive and drug-resistant Candida spp., C. albicans and C. albicans FR, MICs of 1.2 mg/mL and 1.0 mg/mL nMgO were identified, respectively. In contrast, nMgO demonstrated MIC values of 1.0 and 0.7 mg/mL for C. glabrata and C. glabrata ER, respectively. In addition, each Candida species reached an MBC90 of 0.7-1.2 mg/mL nMgO, but only C.glabrata ER was reduced to MBC99.99 at 1.2 mg/mL nMgO (Figure 3C). In addition, in most species tested, the identification of the most potent concentration (MPC) of nMgO was determined. The MPC indicates the nanoparticle concentration that is most effective across poly-microbial communities17.
Method B exposes bacteria in the logarithmic growth phase to nanoparticles of certain concentrations to determine if the nanoparticles are bacteriostatic (inhibitory) or bactericidal using the MBCs identified in method A. This method uses spectrophotometry (OD600) measurements over discrete time periods to identify alterations in bacterial growth in response to nanoparticle exposure. In addition, bacteria exposed to bacteriostatic or bactericidal antibiotics are concurrently grown with the nanoparticle-exposed bacteria in separate wells to provide a reference in the identification of these nanoparticle activities. The concentrations of nMgO and nMg(OH)2 used were derived from a previous study using bacteria of the logarithmic growth phase exposed to nMgO and nMg(OH)2. Concentrations of nMgO and nMg(OH)2 were identified using mM equivalents ranging from 5 mM to 50 mM. The results showed that nMgO was bactericidal to MRSA at 30 mM (1.2 mg/mL nMgO), while nMg(OH)2 was bacteriostatic at 50 mM (2.9 mg/mL nMg(OH)2). The concentrations of antibiotics used were identified from the literature and confirmed in our own experiments. The experimental workflow for this method is illustrated in Figure 1B. Representative results for method B can be seen in Figure 4. Here, MRSA unexposed to nanoparticles grew exponentially to an OD600 of 0.85. When exposed to 1.2 mg/mL nMgO and 6.25 µg/mL trimethoprim, bacterial growth was reduced to an OD600 of 0.18 (80.2% and 81.6%, respectively), and 2.9 mg/mL nMg(OH)2 reduced bacterial growth to an OD600 of 0.25 (70.3%). Exposure to 1.0 µl/mL vancomycin resulted in a 99.99% reduction in the growth of MRSA, suggesting that the concentrations of nMgO and nMg(OH)2 used here were bacteriostatic in activity.
Method C examines the antibacterial activity of nanostructured surfaces. A bacterial culture in lag-phase growth was seeded directly onto a nanostructured surface to measure CFUs with or without direct contact with the surface. Using this method, it was possible to determine the surface effects on bacterial adhesion and viability under direct contact conditions. Bacteria in direct contact with the nanostructured surfaces and in indirect contact (in culture suspension) were collected and quantified in CFU/mL to determine the bacterial growth under each condition. These data obtained may be useful in downstream applications of nanostructured materials on surfaces for clinical use, where a reduction in bacterial colonization on surface structures is desired. The experimental workflow for this method is illustrated in Figure 2A. Representative results for method C can be seen in Figure 5. Here, there was no inhibition of bacterial growth with indirect contact. However, bacterial adhesion was reduced for all substrates, most significantly with the ZC21 alloy, followed by T64 (titanium), magnesium, glass only, and the ZSr41 alloy, respectively. These results indicate that ZC21 had the strongest antibacterial activity against the adhesion and growth of MRSA for all the samples tested14.
Method D examines antibacterial activity at a select area of interest at the cell-nanostructure interface through the direct placement of bacteria in filter paper onto a nanostructured surface. Using this method, a correlation between antibacterial activity and surface properties, such as the surface chemistry, roughness, and area of the nanostructured surface, can be identified16. The experimental workflow for this method is illustrated in Figure 2B. Representative results for method D can be seen in Figure 6. Here, no viable S. aureus was identified on the 1.9A, 1.9 AA, and EPD samples or their paired filter papers. However, exposure to the EPD samples after annealing (A-EPD) reduced bacterial growth to a few cells on the nanostructured surface and the paired filter paper. The sample containing magnesium (Mg) also had no bacterial growth, but viable bacteria were isolated from the paired filter paper. A reduction in the growth of S. aureus was not seen with the titanium and glass samples or their paired filter papers16.
In addition to the use of the methods described above to determine antimicrobial activities, SEM and fluorescence microscopy can be used to characterize the morphological changes that occur in microorganisms after exposure to nanoparticles and nanostructured materials. The representative SEM images showing the post-exposure gram-negative Escherichia coli are presented in Figure 7A, and those showing the post-exposure Gram-positive S. aureus are presented in Figure 7B. Using the SEM imaging technique with a 5,000x magnification, phenotypic changes in E. coli exposed to concentrations of 0.5 mg/mL nMgO or higher could be identified17. Fluorescence microscopy can also be used to characterize morphological changes in microorganisms, but with potentially lower costs and greater accessibility than SEM. Figure 8 shows representative fluorescence images of E. coli exposed to the nanostructured material MgY_O for 1 day, 2 days, and 3 days, as well as reference E. coli. Only one bacterial species, E. coli, was used initially. In the future, we will consider exposing additional bacterial species to the same materials of interest to produce a more comprehensive understanding of bacterial responses to MgY_O exposure. In these results, individual E. coli cells and colonies could be viewed and imaged using thioflavin T staining and a fluorescence microscope for the qualitative analysis of the antibacterial activity of MgY_O15.
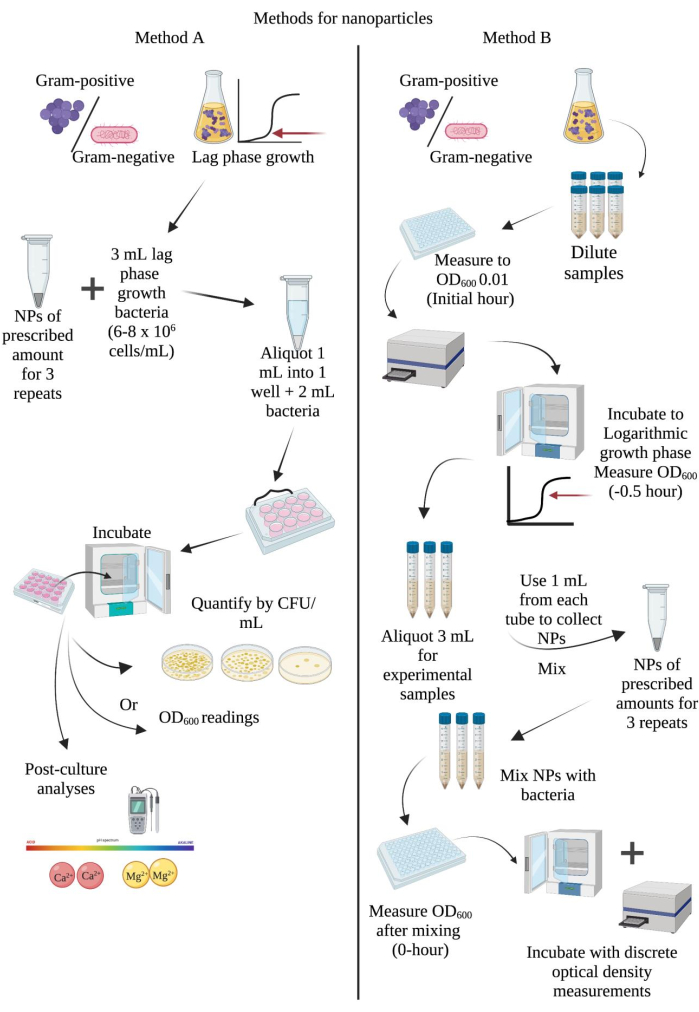
Figure 1: Schematic diagrams to determine nanoparticle MIC or MBC90-99.99 and bacteriostatic or bactericidal activities in post-exposure cell cultures. (A) Direct co-culture method17. (B) Direct exposure method. Abbreviations: CFU = colony-forming units; NPs= nanoparticles; OD600 = optical density at 600 nm. Please click here to view a larger version of this figure.
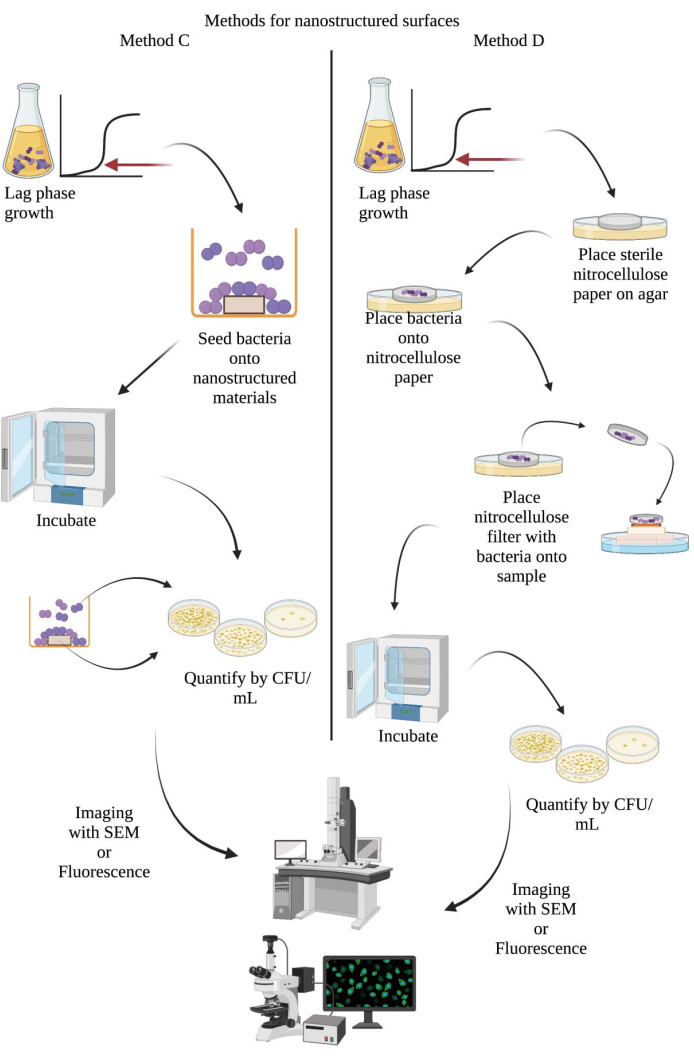
Figure 2: Schematic diagrams to determine bacterial growth with direct and indirect contact and at the cell-nanostructure interface of nanostructured surfaces. (A) Direct culture method14. (B) Focused-contact exposure method16. Abbreviations: CFU = colony-forming units; SEM = scanning electron microscopy. Please click here to view a larger version of this figure.
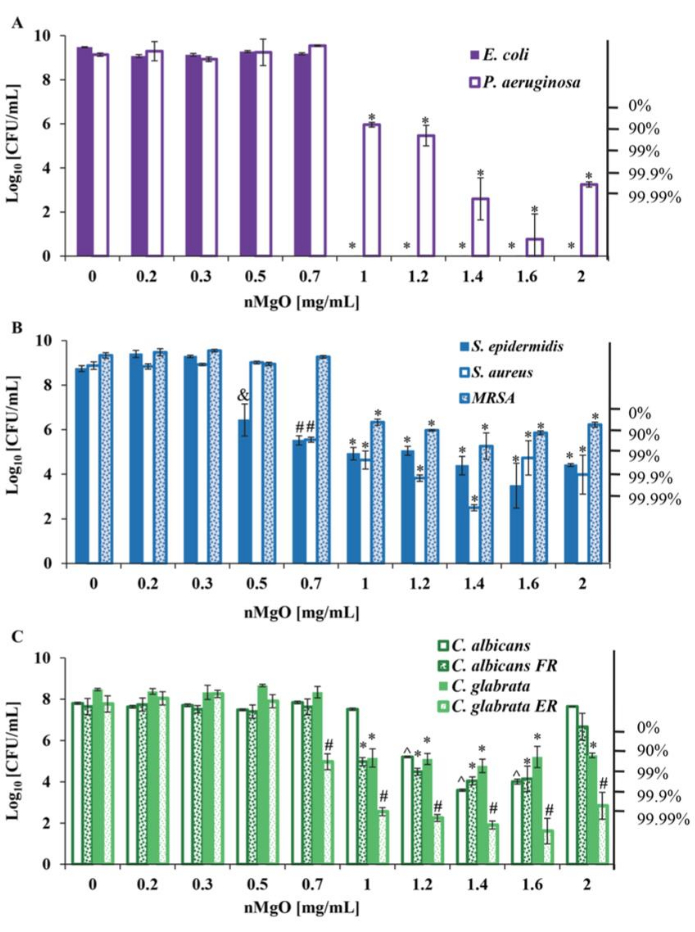
Figure 3: Representation of viable bacteria and yeasts quantified by CFU/mL 24 h post-exposure to 0-2.0 mg/mL nMgO. (A) CFU/mL of gram-negative bacteria, including E. coli and P. aeruginosa. (B) CFU/mL of gram-positive bacteria, including S. epidermidis, S. aureus, and methicillin-resistant S. aureus. (C) CFU/mL of drug-susceptible C. albicans, drug-resistant C. albicans FR, drug-susceptible C. glabrata, and drug-resistant C. glabrata ER. Data are represented as mean ± standard deviation (N = 9). *p ≤ 0.05, significantly lower than the groups at 0-0.7 mg/mL nMgO for the respective bacterium or yeast. ^p ≤ 0.05, significantly lower than the groups at 0-1 mg/mL nMgO for the respective microorganism. #p ≤ 0.05, significantly lower than the groups at 0-0.5 mg/mL nMgO for the respective microorganism. &p ≤ 0.05, significantly lower than the groups at 0-0.3 mg/mL nMgO for the respective microorganism17. This figure is modified from Nguyen et al.17. Abbreviations: FR = fluconazole-resistant; ER = echinocandin-resistant; nMgO = magnesium oxide nanoparticles; CFU = colony-forming units. Please click here to view a larger version of this figure.
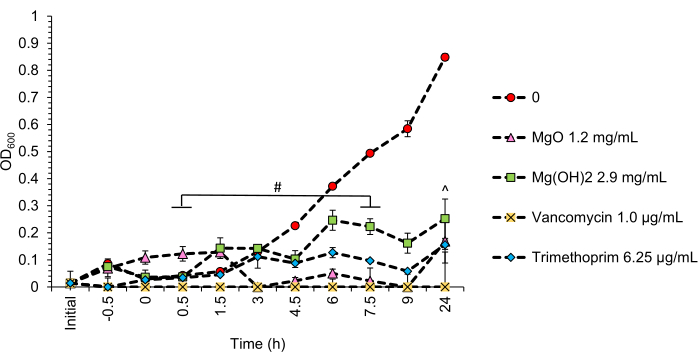
Figure 4: Representation of optical density readings (OD600) taken in real-time for methicillin-resistant S. aureus. MRSA exposure to 1.2 mg/mL nMgO, 2.9 mg/mL nMg(OH)2, 6.25 µg/mL trimethoprim, and 1.0 µg/mL vancomycin for 24 h. p ≤ 0.025: the growth of MRSA exposed to 1.0 µg/mL vancomycin at 0.5 h is significantly lower than the growth of MRSA exposed to 2.9 mg/mL Mg(OH)2 at 7.5 h. ^p ≤ 0.025: the growth of MRSA exposed to 1.0 µg/mL vancomycin at 24 h post-inoculation is significantly lower than the growth of MRSA exposed to 2.9 mg/mL Mg(OH)2 at 24 h. Data are presented as the mean ± the standard error of the mean (SEM). Please click here to view a larger version of this figure.
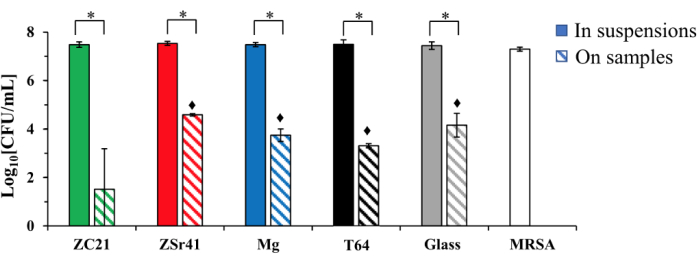
Figure 5: Representation of viable methicillin-resistant S. aureus quantified by CFU/mL at 24 h post-exposure to samples and controls. Bacteria were seeded at an actual density of 7.5 × 105 cells/mL and confirmed by CFU/mL. Data are represented as the mean ± standard deviation, n = 3; *p < 0.05.  p < 0.05 when compared with the CFU on ZC21 samples. This figure was modified from Zhang et al.14. Please click here to view a larger version of this figure.
p < 0.05 when compared with the CFU on ZC21 samples. This figure was modified from Zhang et al.14. Please click here to view a larger version of this figure.
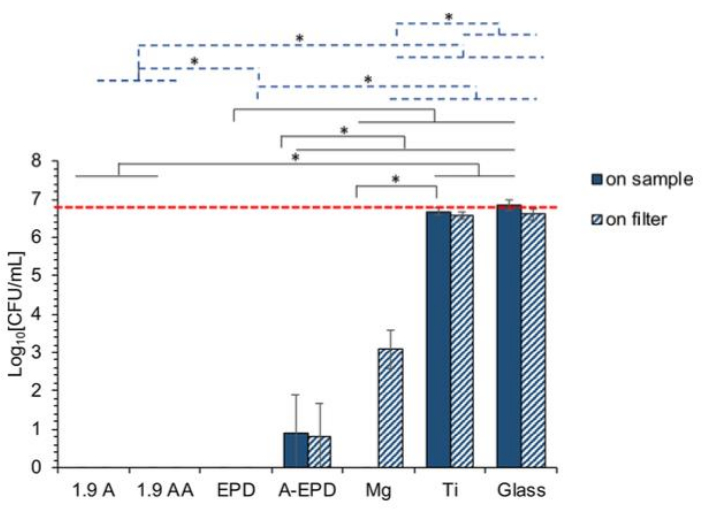
Figure 6: Representative quantification by CFU/mL of the bacterial seeding density 24 h post-exposure to surface-treated Mg samples of 1.9 A, 1.9 AA, EPD, and A-EPD, and Mg and Ti control samples. Bacteria were seeded at an actual density of 6 × 106 CFU/mL, as represented by the red dashed line. Data are represented as the mean ± standard deviation; n = 3. *p < 0.05. The solid black line indicates the statistical analysis results for the bacterial density on the sample surfaces. The blue dashed line indicates the statistical analysis results for the bacterial density on the filters covering the sample surfaces. Abbreviations: CFU = colony-forming units; A-EPD = electrophoretic deposition after annealing; EPD = electrophoretic deposition before annealing; A = anodization before annealing; AA = anodization after annealing. This figure was modified from Lin et al.16. Please click here to view a larger version of this figure.
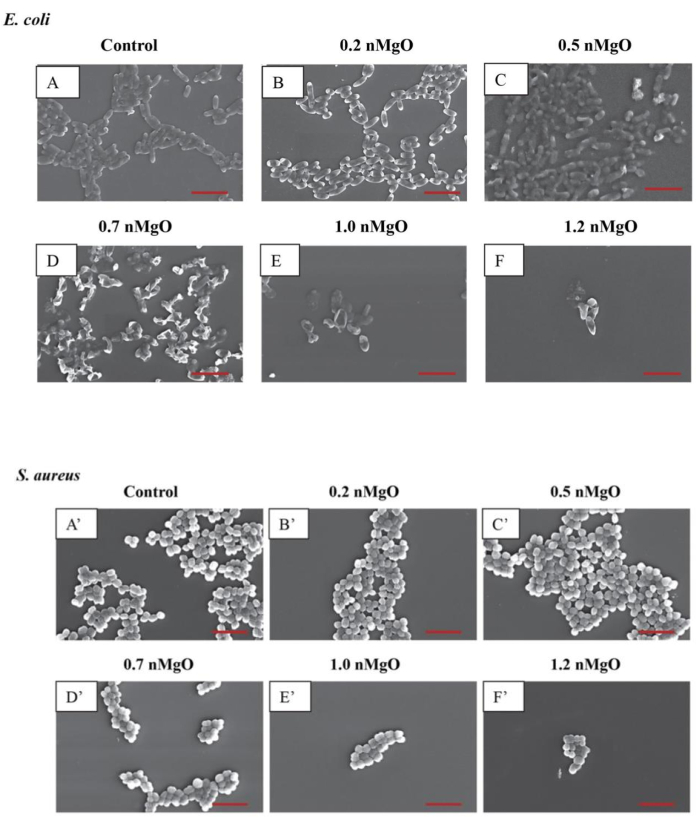
Figure 7: Representative SEM images of gram-negative E. coli and gram-positive S. aureus exposed to 0–1.2 mg/mL nMgO. (A) E. coli showed similar morphology at 1.2 mg/mL to 2.0 mg/mL nMgO. For this reason, one image at 1.2 mg/mL is shown as a representative of E. coli exposed to higher nMgO concentrations. (B) S. aureus also showed similar morphology at 1.2 mg/mL to 2.0 mg/mL nMgO. For this reason, one image at 1.2 mg/mL is shown as a representative of S. aureus exposed to higher nMgO concentrations17. Scale bars = 5 µm. nMgO = magnesium oxide nanoparticles. This figure was modified from Nguyen et al.17. Please click here to view a larger version of this figure.
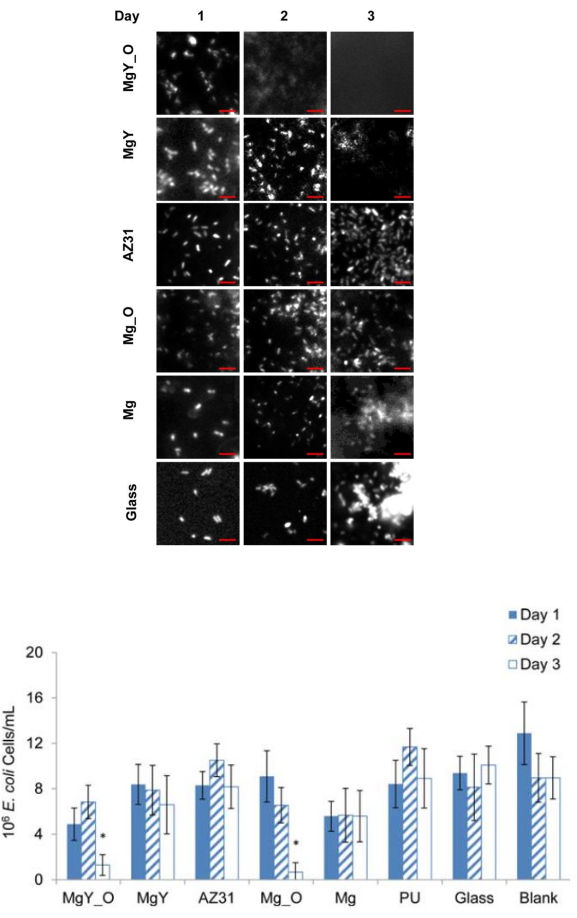
Figure 8: Representative fluorescence images of E. coli adhered onto the substrates with quantification at 1 day, 2 days, and 3 days of incubation. (A) Fluorescence images of E. coli stained with 10 µM thioflavin T. (B) Quantification of E. coli exposed to materials of interest using cells/mL. There was no bacterial adherence to MgY_O after 2 days of incubation, and decreased adherence to MgY after 3 days of incubation. Scale bar = 10 µm. This figure was modified from Lock et al.15. Please click here to view a larger version of this figure.
Discussion
We have presented four in vitro methods (A-D) to characterize the antibacterial activities of nanoparticles and nanostructured surfaces. While each of these methods quantifies bacterial growth and viability over time in response to nanomaterials, some variation exists in the methods used to measure the initial bacterial seeding density, growth, and viability over time. Three of these methods, the direct co-culture method (A)17, the direct culture method (C)14, and the focused-contact exposure method (D)16, quantify bacterial growth and viability after a period of exposure to nanomaterials by counting the CFUs per unit volume. In comparison, the direct exposure method (B) quantifies bacterial growth via discrete real-time optical density (OD600) measurements. This method can also be applied to the direct co-culture, direct culture, and focused-contact exposure methods if there are time constraints for quantifying CFUs, and/or a real-time bacterial density reading is needed for kinetic studies. If effective antibiotics are not known for the bacterial species to be tested in the direct exposure method, a literature search must be carried out to determine the correct antibiotics and their MIC values to serve as references for the nanomaterial groups. A confirmation of the MIC values can be made by completing a standardization curve, which plots the relationship between the spectrophotometry data and the quantification of colony-forming units at specific time points. This process correlates the number of cells present in culture at specific points with the respective optical density. In addition, bacteria at lag-stage growth are introduced to nanoparticles in the direct co-culture method and nanostructured surfaces in the direct culture and focused-contact exposure methods. In contrast, bacteria at the logarithmic growth stage are introduced to nanoparticles in the direct exposure method. Furthermore, bacterial seeding densities are quantified as cells/mL in the direct co-culture method, the direct culture method, and the focused-contact exposure method, whereas the optical density (OD600) of bacteria is measured to track real-time growth in the direct exposure method.
Although each of these methods is similar in framework, they also present unique characteristics to distinguish the effects of nanoparticles or nanostructured surfaces on bacteria. For example, the direct co-culture method characterizes bacterial responses to increasing pre-measured concentrations of nanoparticles by CFU/mL quantification (Figure 3). This method generates minimum inhibitory and minimum bactericidal concentrations (MIC and MBC) of nanoparticles for microbial species of interest and, if multiple species are examined, the most potent concentration (MPC) of nanoparticles17. The identification of the MIC, MBC90-99.99, and MPC is essential for the application of nanoparticles in downstream applications. For example, these data can be applied to in vivo examinations of nanoparticle antibacterial activity while maintaining cytocompatibility when less-than-lethal concentrations have also been identified. In contrast, the direct exposure method characterizes nanoparticle activity as bacteriostatic or bactericidal in a manner that is similar to the classification of antibiotics, where synergistic or undesirable effects must be identified prior to use in research or clinical settings.
The characterization of antibacterial compounds as bacteriostatic or bactericidal is relevant to clinical use and in vitro and in vivo research38. For this reason, when designing methods to examine these characteristics, any negative effects that may impact the results must be identified. Here, the direct exposure method categorizes nanoparticles as bacteriostatic or bactericidal for specific bacterial species using real-time measurements of optical density at 600 nm (Figure 4). Bacteria in the logarithmic growth phase are mixed with nanoparticles at the MIC level to create bacteria and nanoparticle suspensions. These suspensions are discreetly measured in real time to determine the ongoing effects of nanoparticle exposure on the bacteria. To verify the results obtained, concurrently grown bacteria in suspension with bacteriostatic and bactericidal antibiotics act as a reference to confirm the growth rates of the nanoparticle-exposed bacteria.
The incorporation of nanoparticles into bioengineering, clinical, and environmental sciences may have far-reaching possibilities, but using the antibacterial data of nanoparticles for in vitro and in vivo studies on nanostructured surfaces is challenging to accomplish. For this reason, we presented the direct culture and focused-contact exposure methods to characterize the in vitro antibacterial activities of nanostructured surfaces. The direct culture method examines the antibacterial effects of nanostructured surfaces when bacteria are in direct contact with the materials of interest14. Here, bacteria suspended in rSBF plus 10% FBS are exposed to nanostructured surfaces, followed by the quantification of bacterial growth over time for bacteria in direct contact with the surface and in suspension surrounding the material of interest (indirect contact) using CFU/mL. Previously, experimental outcomes involving magnesium alloy nanostructures (ZC21 and ZSr41 alloys) have indicated that antibacterial activities increase when the bacteria are in direct contact with nanostructured surfaces in comparison with indirect contact, where the bacteria remain in suspension with little to no contact14 (Figure 5). Finally, the focused-contact method characterizes antibacterial activity at the interface of bacteria and a nanostructured surface16. Here, we examined magnesium oxide anodized or electrophoretically deposited onto the surface of a bioresorbable magnesium alloy to determine the potential disruption of bacterial adhesion and biofilm formation over time. The disruption of biofilm formation can be critical for patient healing processes as biofilms tend to evade host immune responses and antibiotic therapies39 and result in infections that are extremely difficult to treat.
Here, we present a method adapted from the Japanese Industrial Standard JIS Z 2801:200016. This method ensures that bacteria are attached to sterile nitrocellulose filter paper followed by exposure to a fixed area of the nanostructured surface. This limits the characterization of the surface effects on antibacterial activity to the testing area only. In the representative results, the anodization before annealing (1.9A), anodization after annealing (1.9AA), and electrophoretic deposition before annealing (EPD) samples showed significant effects on reducing bacterial growth as well as their corresponding filter papers when compared to the magnesium, titanium, and glass control samples. The electrophoretic deposition after annealing (A-EPD) samples were also effective in reducing bacterial growth, but this result was less significant in comparison to the results obtained for the 1.9A, 1.9AA, and EPD samples16 (Figure 6).
In summary, each in vitro method presented here is reasonably easy to master and incorporates inexpensive and readily available materials. These methods can be used to characterize interactions between a wide range of nanomaterials and microbial species that are of interest to researchers. Understanding the similarities and differences among these methods is necessary to design optimal experiments to meet the research objectives. Here, the scientific knowledge gained from these in vitro methods can be applied to downstream medical applications where antibiotics must be limited or represent an unfavorable approach.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors appreciate the financial support from the U.S. National Science Foundation (NSF CBET award 1512764 and NSF PIRE 1545852), the National Institutes of Health (NIH NIDCR 1R03DE028631), the University of California (UC) Regents Faculty Development Fellowship, the Committee on Research Seed Grant (Huinan Liu), and the UC-Riverside Graduate Research Mentorship Program Grant awarded to Patricia Holt-Torres. The authors appreciate the assistance provided by the Central Facility for Advanced Microscopy and Microanalysis (CFAMM) at UC-Riverside for the use of SEM/EDS and Dr. Perry Cheung for the use of XRD. The authors would also like to thank Morgan Elizabeth Nator and Samhitha Tumkur for their assistance with the experiments and data analyses. Any opinions, findings, conclusions, or recommendations expressed in this article are those of the authors and do not necessarily reflect the views of the National Science Foundation or the National Institutes of Health.
Materials
| 1.5 mL microcentrifuge tube | Milipore Sigma | Z336777 | |
| 80 L NTRL Certified Convection Drying Oven | MTI Corporation | BPG-7082 | https://www.mtixtl.com/BPG-7082.aspx |
| (hydroxymethyl) aminomethane buffer pH 8.5; Tris buffer | Sigma-Aldrich | 42457 | |
| AnaSpec THIOFLAVIN T ULTRAPURE GRADE | Fisher Scientific | 50-850-291 | |
| Electron-multiplying charge-coupled device digital camera | Hamamatsu | C9100-13 | |
| Falcon 15 mL conical tubes | Fisher Scientific | 14-959-49B | |
| Gluteraldehyde | Sigma-Aldrich | G5882 | |
| Hemocytometer | Brightline, Hausser Scientific | 1492 | |
| Inductively coupled plasma – optical emission spectrometry (ICP-OES) | PerkinElmer | 8000 | |
| Inverse microscope | Nikon | Eclipse Ti-S | |
| Luria Bertani Broth | Sigma Life Science | L3022 | |
| Luria Bertani Broth + agar | Sigma Life Science | L2897 | |
| MacroTube 5.0 | Benchmark Scientific | C1005-T5-ST | |
| Magnesium oxide nanoparticles | US Research Nanomaterials, Inc | Stock #: US3310 M | MgO, 99+%, 20 nm |
| MS Semi-Micro Balance | Mettler Toledo | MS105D | |
| Nitrocellulose paper | Fisherbrand | 09-801A | |
| Non-tissue treated 12-well polystyrene plate | Falcon Corning Brand | 351143 | |
| Non-tissue treated 48-well polystyrene plate | Falcon Corning Brand | 351178 | |
| Non-tissue treated 96-well polystyrene plate | Falcon Corning Brand | 351172 | |
| Petri dish 100 mm | VWR | 470210-568 | |
| Petri dish, 15 mm | Fisherbrand | FB0875713A | |
| pH meter | VWR | SP70P | |
| Scanning electron microscopy (SEM) | TESCAN | Vega3 SBH | |
| Sonicator | VWR | 97043-936 | |
| Table top centrifuge | Fisher Scientific | accuSpin Micro 17 | |
| Table top centrifuge | Eppendorf | Centrifuge 5430 | |
| Tryptic Soy Agar | MP | 1010617 | |
| Tryptic Soy Broth | Sigma-Aldrich | 22092-500G | |
| UV-Vis spectrophotometer | Tecan | Infinite 200 PRO | https://lifesciences.tecan.com/plate_readers/infinite_200_pro |
| VWR Benchmark Incu-shaker 10L | VWR | N/A | |
| X-ray power defraction | Panalytical | N/A | PANalytical Empyrean Series 2 |
References
- Haque, M., Sartelli, M., McKimm, J., Abu Bakar, M. Health care-associated infections – An overview. Infection and Drug Resistance. 11, 2321-2333 (2018).
- O’Connell, K. M. G. Combating multidrug-resistant bacteria: Current strategies for the discovery of novel antibacterials. Angewandte Chemie. 52 (41), 10706-10733 (2013).
- Li, B., Webster, T. J. Bacteria antibiotic resistance: New challenges and opportunities for implant-associated orthopedic infections. Journal of Orthopaedic Research. 36 (1), 22-32 (2018).
- Yung, D. B. Y., Sircombe, K. J., Pletzer, D. Friends or enemies? The complicated relationship between Pseudomonas aeruginosa and Staphylococcus aureus. Molecular Microbiology. 116 (1), 1-15 (2021).
- Adams, R. A. Rifamycin antibiotics and the mechanisms of their failure. The Journal of Antibiotics. 74 (11), 786-798 (2021).
- Harding, E. WHO global progress report on tuberculosis elimination. The Lancet. Respiratory Medicine. 8 (1), 19 (2020).
- Baptista, P. V. Nano-strategies to fight multidrug resistant bacteria-"A battle of the titans". Frontiers in Microbiology. 9, 1441 (2018).
- Seong, M., Lee, D. G. Silver nanoparticles against Salmonella enterica serotype typhimurium: Role of inner membrane dysfunction. Current Microbiology. 74 (6), 661-670 (2017).
- Dasgupta, N., Ramalingam, C. Silver nanoparticle antimicrobial activity explained by membrane rupture and reactive oxygen generation. Environmental Chemistry Letters. 14, 477-485 (2016).
- Su, H. L. The disruption of bacterial membrane integrity through ROS generation induced by nanohybrids of silver and clay. Biomaterials. 30 (30), 5979-5987 (2009).
- Cui, Y. The molecular mechanism of action of bactericidal gold nanoparticles on Escherichia coli. Biomaterials. 33 (7), 2327-2333 (2012).
- Ranjan, S., Ramalingam, C. Titanium dioxide nanoparticles induce bacterial membrane rupture by reactive oxygen species generation. Environmental Chemistry Letters. 14, 487-494 (2016).
- Kadiyala, U., Turali-Emre, E. S., Bahng, J. H., Kotov, N. A., VanEpps, J. S. Unexpected insights into antibacterial activity of zinc oxide nanoparticles against methicillin resistant Staphylococcus aureus (MRSA). Nanoscale. 10 (10), 4927-4939 (2018).
- Zhang, C. Antimicrobial bioresorbable Mg-Zn-Ca alloy for bone repair in a comparison study with Mg-Zn-Sr alloy and pure Mg. ACS Biomaterials Science and Engineering. 6 (1), 517-538 (2020).
- Lock, J. Y. Degradation and antibacterial properties of magnesium alloys in artificial urine for potential resorbable ureteral stent applications. Journal of Biomedical Materials Research. Part A. 102 (3), 781-792 (2014).
- Lin, J., Nguyen, N. -. Y. T., Zhang, C., Ha, A., Liu, H. H. Antimicrobial properties of MgO nanostructures on magnesium substrates. ACS Omega. 5 (38), 24613-24627 (2020).
- Nguyen, N. -. Y. T., Grelling, N., Wetteland, C. L., Rosario, R., Liu, H. Antimicrobial activities and mechanisms of magnesium oxide nanoparticles (nMgO) against pathogenic bacteria, yeasts, and biofilms. Scientific Reports. 8 (1), 16260 (2018).
- Bindhu, M. R., Umadevi, M., Micheal, M. K., Arasu, M. V., Al-Dhabi, N. A. Structural morphological and optical properties of MgO nanoparticles for antibacterial applications. Materials Letters. 166, 19-22 (2016).
- He, Y. Study on the mechanism of antibacterial action of magnesium oxide nanoparticles against foodborne pathogens. Journal of Nanobiotechnology. 14 (1), 54 (2016).
- Zhang, K., An, Y., Zhang, L., Dong, Q. Preparation of controlled nano-MgO and investigation of its bactericidal properties. Chemosphere. 89 (11), 1414-1418 (2012).
- Hayat, S. In vitro antibiofilm and anti-adhesion effects of magnesium oxide nanoparticles against antibiotic resistant bacteria. Microbiology and Immunology. 62 (4), 211-220 (2018).
- Dong, C. Investigation of Mg(OH)2 nanoparticles as an antibacterial agent. Journal of Nanoparticle Research. 12, 2101-2109 (2010).
- Halbus, A. F., Horozov, T. S., Paunov, V. N. Controlling the antimicrobial action of surface modified magnesium hydroxide nanoparticles. Biomimetics. 4 (2), 41 (2019).
- Pan, X. Investigation of antibacterial activity and related mechanism of a series of Nano-Mg(OH)2. ACS Applied Materials and Interfaces. 5 (3), 1137-1142 (2013).
- Ipe, D. S., Kumar, P. T. S., Love, R. M., Hamlet, S. M. Silver nanoparticles at biocompatible dosage synergistically increases bacterial susceptibility to antibiotics. Frontiers in Microbiology. 11, 1074 (2020).
- Parvekar, P., Palaskar, J., Metgud, S., Maria, R., Dutta, S. The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) of silver nanoparticles against Staphylococcus aureus. Biomaterial Investigations in Dentistry. 7 (1), 105-109 (2020).
- Loo, Y. Y. In vitro antimicrobial activity of green synthesized silver nanoparticles against selected gram-negative foodborne pathogens. Frontiers in Microbiology. 9, 1555 (2018).
- Ali, K. Microwave accelerated green synthesis of stable silver nanoparticles with Eucalyptus globulus leaf extract and their antibacterial and antibiofilm activity on clinical isolates. PLoS One. 10 (7), e0131178 (2015).
- Al-Jumaili, A., Alancherry, S., Bazaka, K., Jacob, M. V. Review on the antimicrobial properties of carbon nanostructures. Materials. 10 (9), 1066 (2017).
- MTI Corporation. . 80L NTRL Certified Convection Drying Oven (18″x16″x18″, 250°C) with Digital Temperature Controller (SSP) – BPG-7082. , (2022).
- CHEBI. . Tris (CHEBI:9754). , (2023).
- . Cold Spring Harbor Protocols. Phosphate buffer. Cold Spring Harbor Laboratory Press. , (2016).
- Barat, R., Montoya, T., Seco, A., Ferrer, J. Modelling biological and chemically induced precipitation of calcium phosphate in enhanced biological phosphorus removal systems. Water Research. 45, 3744-3752 (2011).
- Carlsson, H., Aspegren, H., Lee, N., Hilmer, A. Calcium phosphate precipitation in biological phosphorus removal systems. Water Research. 31 (5), 1047-1055 (1997).
- Gonzalez, J., Hou, R. Q., Nidadavolu, E. P. S., Willumeit-Römer, R., Feyerabend, F. Magnesium degradation under physiological conditions – Best practice. Bioactive Materials. 3 (2), 174-185 (2018).
- Oyane, A., et al. Preparation and assessment of revised simulated body fluids. Journal of Biomedical Materials Research. Part A. 65 (2), 188-195 (2003).
- Xia, B., et al. Amyloid histology stain for rapid bacterial endospore imaging. Journal of Clinical Microbiology. 49 (8), 2966-2975 (2011).
- Pankey, G. A., Sabath, L. D. Clinical relevance of bacteriostatic versus bactericidal mechanisms of action in the treatment of Gram-positive bacterial infections. Clinical Infectious Diseases. 38 (6), 864-870 (2004).
- Ribeiro, M., Monteiro, F. J., Ferraz, M. P. Infection of orthopedic implants with emphasis on bacterial adhesion process and techniques used in studying bacterial-material interactions. Biomatter. 2 (4), 176-194 (2012).

