Establishment and Analysis of Three-Dimensional (3D) Organoids Derived from Patient Prostate Cancer Bone Metastasis Specimens and their Xenografts
Summary
Three-dimensional cultures of patient BMPC specimens and xenografts of bone metastatic prostate cancer maintain the functional heterogeneity of their original tumors resulting in cysts, spheroids and complex, tumor-like organoids. This manuscript provides an optimization strategy and protocol for 3D culture of heterogeneous patient derived samples and their analysis using IFC.
Abstract
Three-dimensional (3D) culture of organoids from tumor specimens of human patients and patient-derived xenograft (PDX) models of prostate cancer, referred to as patient-derived organoids (PDO), are an invaluable resource for studying the mechanism of tumorigenesis and metastasis of prostate cancer. Their main advantage is that they maintain the distinctive genomic and functional heterogeneity of the original tissue compared to conventional cell lines that do not. Furthermore, 3D cultures of PDO can be used to predict the effects of drug treatment on individual patients and are a step towards personalized medicine. Despite these advantages, few groups routinely use this method in part because of the extensive optimization of PDO culture conditions that may be required for different patient samples. We previously demonstrated that our prostate cancer bone metastasis PDX model, PCSD1, recapitulated the resistance of the donor patient’s bone metastasis to anti-androgen therapy. We used PCSD1 3D organoids to characterize further the mechanisms of anti-androgen resistance. Following an overview of currently published studies of PDX and PDO models, we describe a step-by-step protocol for 3D culture of PDO using domed or floating basement membrane (e.g., Matrigel) spheres in optimized culture conditions. In vivo stitch imaging and cell processing for histology are also described. This protocol can be further optimized for other applications including western blot, co-culture, etc. and can be used to explore characteristics of 3D cultured PDO pertaining to drug resistance, tumorigenesis, metastasis and therapeutics.
Introduction
Three-dimensional cultured organoids have drawn attention for their potential to recapitulate the in vivo architecture, cellular functionality and genetic signature of their original tissues1,2,3,4,5. Most importantly, 3D organoids established from patient tumor tissues or patient derived xenograft (PDX) models provide invaluable opportunities to understand mechanisms of cellular signaling upon tumorigenesis and to determine the effects of drug treatment on each cell population6,7,8,9,10,11,12,13. Drost et al.5 developed a standard protocol for establishment of human and mouse prostate organoids, which has been widely adopted in the field of urology. In addition, significant effort has been dedicated for further characterization of 3D organoids and to understand the detailed mechanisms of tumorigenesis and metastasis4,12,14,15. In addition to the previously established and widely accepted protocol for 3D organoids cultures, we describe here a step-by-step protocol for the 3D culture of PDO using three different doming methods in optimized culture conditions.
In this manuscript, 3D organoids were established as an ex vivo model of bone metastatic prostate cancer (BMPC). The cells used for these cultures came from the Prostate Cancer San Diego (PCSD) series and were derived directly from patient prostate cancer bone metastatic tumor tissues (PCSD18 and PCSD22) or patient derived xenograft (PDX) tumor models (samples named PCSD1, PCSD13, and PCSD17). Because spontaneous bone metastasis of prostate cancer cells is rare in genetically engineered mouse models16, we used direct intra-femoral (IF) injection of human tumor cells into male Rag2-/-γc-/- mice to establish the PDX models of bone metastatic prostate cancer17.
Once 3D organoids are established from heterogeneous patient tumor cells or patient derived xenografts, it is essential to confirm their identity as prostate tumor cells and to determine their phenotypes in the 3D organoid cultures. Immunofluorescence chemistry (IFC) allows the visualization of protein expression in situ in each cell, often indicating the potential functions for specific cell populations2,4. In general, IFC protocols for a vast majority of samples including tissues and cells are straightforward and fully optimized. However, the cell density and number of organoids can be significantly lower than that of conventional culture. Therefore, the IFC protocol for organoids requires additional steps to ensure proper processing and embedding in paraffin for all organoids in the samples. We describe additional steps for an agarose pre-embedding process and tips to label the location of sectioned organoids on the slide that increases the success rate of IFC on organoids especially when the samples of organoids have lower cell density than desired.
Protocol
This study was carried out in strict accordance with the recommendations in the Guide for the University of California San Diego (UCSD) Institutional Review Board (IRB). IRB #090401 Approval was received from the UCSD Institutional Review Board (IRB) to collect surgical specimen from patients for research purposes. An informed consent was obtained from each patient and a surgical bone prostate cancer metastasis specimen was obtained from orthopedic repair of a pathologic fracture in the femur. Animal protocols were performed under the University of California San Diego (UCSD) animal welfare and Institutional Animal Care and Use Committee (IACUC) approved protocol #S10298. Cells from mechanically and enzymatically dissociated patient tumor tissue were intra-femorally injected into 6 to 8 week old male Rag2-/-;γc-/- mice as previously described17. Xenograft tumor volume was determined using an in vivo bioluminescence imaging system and caliper measurements. Upon tumor growth up to 2.0 cm (the maximal allowable size approved by IACUC), the tumor was harvested for 3D organoids establishment.
NOTE: Figure 1 shows the workflow for establishing 3D Organoids and a protocol number for each step of the experimental procedures.
1. Processing of patient derived xenograft (PDX) tumor tissues
NOTE: This is an initial step for organoid establishment for a tumor derived from a xenograft mouse model. This protocol is adapted from a previous publication by Drost et al.5 and we have modified the media conditions to include serum supplementation to organoid media.
- Process the tumor specimen as described below.
- Mince tumor samples to 1-3 mm3 sized pieces and digest the samples with 10 mL of cell dissociation solution for 45 min at room temperature.
- To terminate digestion, add 20 mL of DMEM complete media to the samples.
- Filter the suspension through a 70 μm cell strainer. Use a sterile plunger flange to push any leftover tissue on the top of 70 μm cell strainer.
- Centrifuge at 300 x g for 5 min at 4 °C.
- Wash the cell pellet three times with fresh adDMEM complete media. Match the volume of media for wash and resuspension with the volume of media suggested in Table 3. For example, for a 24 well plate culture condition, the volume for the wash should be 500 µL.
NOTE: As shown in Table 3, the protocol is applicable to different culture conditions. - Determine final cell counts using Trypan blue dye and a hemocytometer.
- After obtaining cell counts, re-suspend cell pellet in 80 µL of 2% FBS in PBS per 2 x 106 tumor cells.
- Add 20 µL of Mouse Cell Depletion Cocktail per 2 x 106 tumor cells. Mix well and incubate for 15 min at 2-8 °C.
- Adjust the volume to 500 µL with 2% FBS in PBS buffer per 2 x 106 tumor cells.
NOTE: Up to 1 x 107 tumor cells in 2.5 mL of cell suspension can be processed on one LS column. - Load the LS columns on the magnetic column separator and place a 15 mL conical tube on a rack underneath to collect the flow-through.
- Rinse each column with 3 mL of 2% FBS in PBS buffer. Discard the conical tube with the wash flow-through and replace with a new, sterile 15 mL conical tube.
- Add the cell suspension (up to 2.5 mL of 1 x 107 tumor cells) onto the column. Collect flow-through that will be the enriched with human tumor cells.
- Wash the column twice with 1 mL of 2% FBS in PBS buffer.
NOTE: It is important to perform wash steps as soon as the column is empty. Also, try to avoid forming air bubbles.
- Aliquot the appropriate volume of cell suspension to a 1.5 mL tube for the desired culture set up (Table 3).
- Centrifuge the 1.5 mL tube at 300 x g and 4 °C for 5 min.
- Carefully remove and discard the supernatant.
2. Processing of patient primary tumor tissues
NOTE: This is an initial step for organoid establishment.
- Follow all of step 1 except the mouse cell depletion process, which is not necessary for processing of patient primary tumor tissues.
3. Forming an attached round dome on the plate
NOTE: This manuscript describes three ways to make a dome from a mixture of the cell pellet and the basement membrane (e.g., Matrigel) as shown in Figure 1 and Figure 2. In Steps 2-4, the cells and the basement membrane should be kept on ice to prevent solidification of the basement membrane.
- Resuspend the cell pellet in the appropriate volume of basement membrane (e.g., 40 µL) for a 24 well plate set up (Table 3).
- Pipette up and down gently to ensure that the cells are re-suspended well in the basement membrane.
- Pipette the appropriate volume (Table 3) of the cell-basement membrane mixture (and optional 10 µL of adDMEM complete media) into the center of the pre-warmed tissue culture plate.
- Invert the plate and immediately place the plate upside down in the CO2 cell culture incubator set at 5% CO2, 37 °C for 15 min. This prevents cells from settling and adhering to the plate bottom while allowing the basement membrane to solidify.
- Pipette the appropriate volume (Table 3) of pre-warmed medium containing 10 μM Y-27632 dihydrochloride into each well.
- Place the plate right side up inside the CO2 cell culture incubator (5% CO2, 37 °C).
- Change the media every 3-4 days. After 5-7 days, use culture medium without 10 μM Y-27632 dihydrochloride to maintain the cultures.
4. Forming a floating dome from an attached round dome on the plate
- After step 3.7, detach the dome using a cell scraper.
5. Forming floating beads
NOTE: This protocol is named as floating beads since the mixture of basement membrane, media, and organoids look like beads.
- Cut a 2 inch x 4 inch piece of paraffin film.
- Place the paraffin film on the top of the divots of an empty tip-holding rack from a 1000 µL plastic pipette tip box.
- Gently press down on the paraffin film to trace the divots using a gloved index finger but without breaking through the paraffin film.
- Spray the paraffin film with 70% ethanol and turn on the UV lamp in the cell culture hood to sterilize the prepared paraffin film for at least 30 min.
- Prepare a mixture of cells and 20 µL of basement membrane. Seeding density can be 50,000 – 250,000 cells per dome.
- Pipette the mixture of cells processed from step 1 or 2, and 20 µL of basement membrane into the mold of the divot formed in the prepared paraffin film.
- Resuspend the cell pellet in basement membrane and pipette the cell suspension in the prepared paraffin filmed divots.
- Place the solidified beads and paraffin film into a 6-well plate. One well in a 6-well plate can fit up to 5 beads.
- Pipette 3-5 mL of pre-warmed medium containing 10 μM Y-27632 dihydrochloride into each well while gently brushing beads off of the paraffin film.
NOTE: As a minimum volume, 3 mL is recommended. For a maximum number of beads (N=5) per well, 5 mL of medium is recommended. - Place the plate inside a CO2 incubator (5% CO2, 37 °C).
- Change the organoid media every 3-4 days. After 5-7 days, use culture medium without 10 μM Y-27632 dihydrochloride to maintain the cultures.
6. In vivo organoids image stitching using microscope8
NOTE: Certain microscopes are unable to reach the outer perimeter of the cell plate (edge wall); therefore, we suggest using the wells close to the perimeter of the cell plate when image stitching.
- Place the cell culture plate in an upward position into the plate holder in the Keyence microscope.
- Place the lens on the center of the target dome.
- Set up the automatic stitching process by selecting number of the frames. For examples, 3 x 3 or 5 x 5 can be chosen to generate 9 images or 25 images total.
- Press the capture button to initiate imaging process.
- Open the image viewer software and load a group of images taken by step 4.
- Click Image Stitching to create a high-resolution stitched image.
NOTE: Capturing of serial 9 or 25 images can be performed either by manual or automatic set up to focus the cells.
7. Organoid processing for histology: the agarose spin down method
NOTE: This protocol is adapted from a previous publication by Vlachogiannis et al.7. We have added a step involving agarose embedding to successfully embed all populations of organoids.
- Remove existing media from the well. Be careful not to aspirate the basement membrane domes.
- Add an equal (equal to the volume of media removed from step 1) volume of cell recovery solution and incubate for 60 min at 4 °C.
- Dislodge the basement membrane dome using a pipette and crush the basement membrane dome using a pipet tip. Collect the dissociated dome and cell recovery solution in a 1.5 mL tube.
- Centrifuge at 300 x g and 4 °C for 5 min.
- Remove the supernatant (cell recovery solution). Save all supernatants in separate tubes until the end when the presence of organoids is confirmed in the final pelleting step.
- Add desired volume (Table 3) of cold PBS and gently pipette up and down to mechanically disturb pellet.
- Centrifuge at 300 x g and 4 °C for 5 min.
- Remove the supernatant (PBS).
- Fix the pellet in a matched volume (e.g., 500 µL for one pellet from the 24 well plate culture condition, Table 3) of 4% PFA for 60 min at room temperature.
- Following fixation, centrifuge at 300 x g and 4 °C for 5 min.
- Remove the supernatant (PFA).
- Wash with matched volume (e.g., 500 µL for one pellet from the 24 well plate culture condition, Table 3) of PBS and centrifuge at 300 x g and 4 °C for 5 min.
- Prepare warm agarose (2% agarose in PBS).
NOTE: Here, cell pellets for frozen sections can be directly re-suspended in 200 µL of OCT compound without further steps in Protocol 7. - Re-suspend the cell pellet in 200 µL of agarose (2% in PBS).
- Immediately after adding agarose, gently detach the cell pellet from the wall of the 1.5 mL tube using the 25 G needle attached to 1 mL syringe. As shown in Figure 3, if the cell pellet is not physically detached from the wall of the 1.5 mL tube, then there is a risk of losing all or part of the cell pellet during the agarose embedding process.
- Wait until the 2% agarose in PBS is completely solidified.
- Detach the solidified agarose block from the 1.5 mL tube using a 25 G needle attached to the 1 mL syringe.
- Transfer the detached agarose block containing the cell pellet to a new 1.5 mL tube.
- Fill the tube with 70% EtOH and proceed further using the conventional protocol for tissue dehydration and paraffin embedding.
8. Histology and Immunofluorescent cytochemistry (IFC) of organoids
- Select the slide(s) for histology or IFC.
- Before initiating the staining process, find out where the cells are located on the slide and draw a circle around the cells on the slide using a marker.
- Draw the perimeter around the edge or boarder of the slide and where circles are located on the slide in a laboratory notebook to record their locations.
- Perform desired staining.
NOTE: During this process, marked circles disappear since regular maker is not resistant to the chemicals. Even some histology permanent markers may be erased during staining process. - After the staining process, place the slide over the drawing in the laboratory notebook to find the locations of cells on the slide.
Representative Results
3D organoids were successfully established from a patient derived xenograft (PDX) model of bone metastatic prostate cancer (BMPC) as well as directly from patient bone metastatic prostate cancer tissue (Figure 4). Briefly, our PDX models of BMPC were established by intra-femoral (IF) injection of tumor cells into male Rag2-/- c-/- mice and then PDX tumors were harvested and processed as described in this manuscript. As shown in Figure 4, PDX tumor tissues from the PCSD series resulted in 3D organoids with differential phenotypes that manifested as cysts, spheroids and higher complexity organoids that formed using this protocol.
A stitched image from 25 high resolution 10x magnification images showed an entire dome of basement membrane and organoids (Figure 5). Using image analysis software, one has the option to sort out the cells or cell clusters that are larger than a certain size or to manually select spheroids or cysts.
The steps for agarose embedding of the organoid cultures and tips to label the location of sectioned organoids on the slide increase the success of performing IFC on the organoids especially when the samples of organoids have a lower cell density than desired. Figure 6 shows an example of IFC on a 5 μm thick paraffin section of organoids targeting cytokeratin 5 (CK5, basal epithelial cell marker), cytokeratin 8 (CK8, luminal epithelial cell marker) and DAPI.
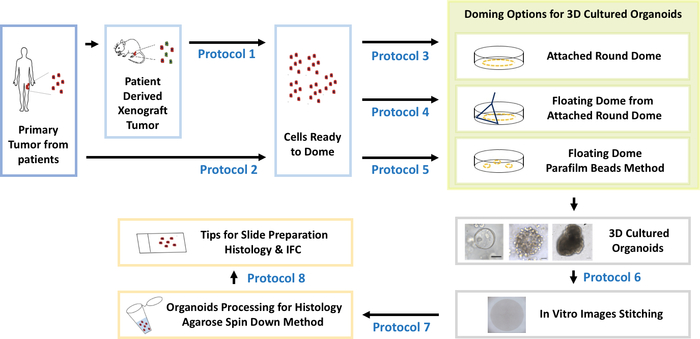
Figure 1: Establishment and characterization for 3D cultured organoids derived either from patient tumor tissues or patient derived xenograft (PDX) tumor tissues. Please click here to view a larger version of this figure.
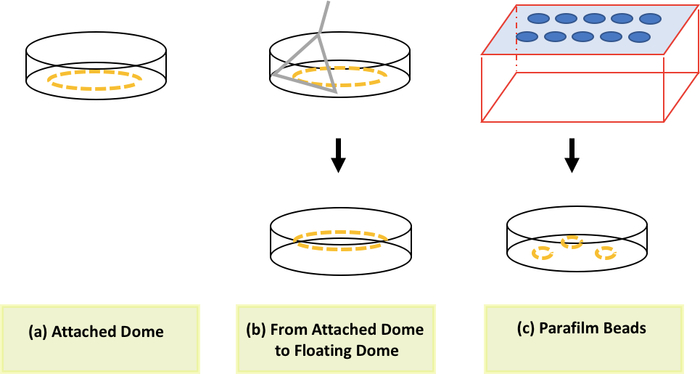
Figure 2: Methodologies to form a mixture of cell pellets and basement membrane. An attached dome (a), a floating dome (b) and floating beads (c). Please click here to view a larger version of this figure.
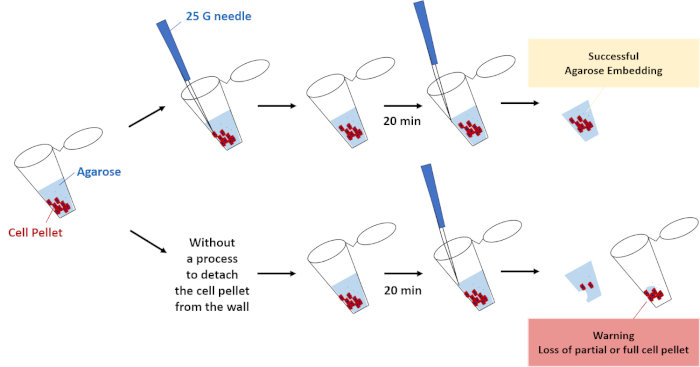
Figure 3: A key step for successful agarose embedding. A process to detach the cell pellet from the wall of the 1.5 mL tube. Please click here to view a larger version of this figure.
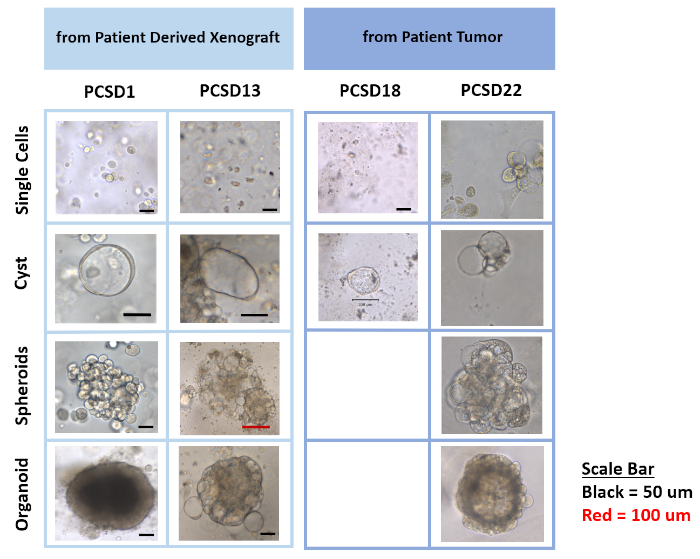
Figure 4: Differential phenotypes of organoids from a series of PCSD PDX tumors, which includes single cells, cysts, spheroids and higher complexity organoids. Please click here to view a larger version of this figure.
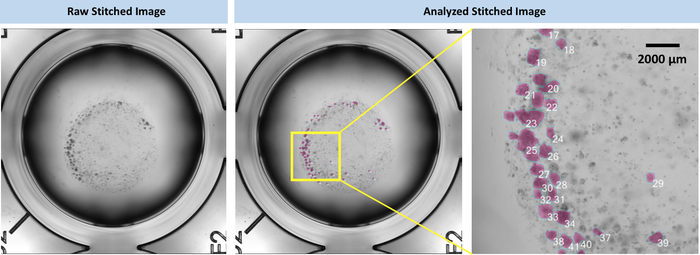
Figure 5: Example of a raw stitched image from total of 25 high resolution images and its application for a follow up analysis to count the number of cells or measure the area of cells/cell clusters. Please click here to view a larger version of this figure.
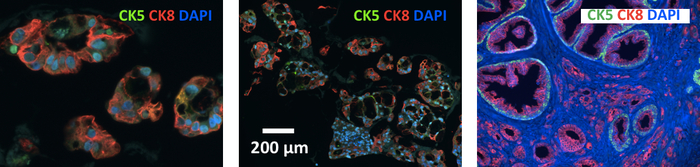
Figure 6: Image showing CK5 and CK8 IFC stained PCSD1 organoids (5 μm thickness of paraffin section). Please click here to view a larger version of this figure.
| Stock Component | Final Concentration | Vol (mL) needed for 500 ml solution |
| 1000 mM HEPES | 10 mM | 5 |
| 200 mM Glutamax | 2 mM | 5 |
| 100x Pen-Strep | 1x | 5 |
| adDMEM/F12 +/+/+ | 485 |
Table 1: adDMEM/F12 +/+/+ Preparation. This table has been previously published by Drost et al.5.
| Stock Component | Final Concentration | Vol (µL) needed for 50 mL solution | Vol (µL) needed for 25 mL solution |
| 50x B27 | 50x Diluted | 1000 | 500 |
| 500 mM N-Acetylcysteine | 1.25 mM | 125 | 62.5 |
| 0.5 mg/mL EGF | 5 ng/mL | 0.5 | 0.3 |
| 100 ug/mL Noggin | 100 ng/mL | 50 | 25 |
| R-Spondin 1 | 10% conditioned medium | 5000 | 2500 |
| 5 mM A83-01 | 500 nM | 5 | 2.5 |
| 0.1 mg/mL FGF10 | 10 ng/mL | 5 | 2.5 |
| 50 µg/mL FGF2 | 5 ng/mL | 5 | 2.5 |
| 10 mM Prostaglandin E2 | 1 µM | 5 | 2.5 |
| 1M Nicotinamide | 10 mM | 500 | 250 |
| 30 mM SB202190 | 10 µM | 16.7 | 8.4 |
| FBS | 10% | 5000 | 2500 |
| 1 µM DHT | 1 nM | 50 | 25 |
| adDMEM/F12 +/+/+ | Bring up to 50 ml | Bring up to 25 mL | |
| 100 mM Y-27632 Dihydrochloride | 10 µM | 5 | 2.5 |
Table 2: Components for C/S Human Media + 10 % FBS. This table has been previously published by Drost et al.5.
| Culture Plate | Seeding Density (cells) | Basement Membrane Volume (μL) | Medium (μL) |
| 48 well | 25,000-50,000 | 20 | 250 |
| 24 well | 50,000-250,000 | 40 | 500 |
| 6 well | 50,000-250,000 | 40 | 2,000 |
Table 3: Seeding density, basement membrane volume, and medium volume needed for one dome. This table is modified from a previous publication by Drost et al.5.
Discussion
3D organoids derived from patient bone metastasis prostate cancer cells are still relatively rare. Here, we describe strategies and further optimized protocol to successfully established serial 3D patient derived organoids (PDOs) of BMPC. In addition, protocols are described to secure the organoids in samples with lower cell density for IFC and IHC analysis. Differential phenotypes in the form of cyst, spheroids and more complex organoids indicate that this protocol provides culture conditions that allows for heterogonous tumor cell populations in patients to retain their cellular phenotype and structure. Thus, the culture conditions described here, which were adapted from Drost et al.5 modified with FBS serum supplementation, has been shown to be optimal for the cultures of patient BMPC-derived cells such that they retain their intra-subject and inter-subject heterogeneity (manuscript in preparation).
Our optimized protocol is practical for an experimental set up with multiple endpoints from limited starting material from primary or PDX tumor tissues to set up 3D organoids cultures. In general, establishment of 3D organoids from patient primary tumor tissues is less successful than establishment of 3D organoids from PDX tumor tissues. Therefore, it is recommended to have a higher seeding density (up to four times higher) of cells for the establishment of 3D organoids from patient primary tumor tissues than from PDX tumors). One method to increase cell yield during tumor tissue processing is to recover any tumor tissue leftover on the top of 70 μm cell strainer by passing the filtered cell suspension through the strainer again.
Three different methods to form a 3D organoid culture (Figure 2) can be used and easily adapted depending on the follow up endpoint analysis. For example, if one desired to have the stitched images after establishing 3D organoids, an attached dome is highly recommended since the image stitching process requires a serial movement of microscopic objective to acquire the desired number of images (either 9 or 25). This process results in a subtle vibration to the holder of cell culture plate. Both types of the floating dome will freely move due to the vibration while acquiring the image. In addition, both types of floating domes can be easily transferred from one well to the another, making them suitable for co-culture with adherent cells after setting up 3D organoid cultures and the adherent cells in the separate wells. The attached dome and floating dome released from being attached have been shown to retain organoids for up to 10 weeks. Interestingly, a floating dome from the floating beads method has been shown to retain organoids for up to 4 months. The floating beads method to produce basement membrane spheres introduced in our protocol involves more steps and skills but can be considered for a long-term culture. In parallel, despite their use for establishment of 3D organoids from different tumors and metastasis, the role of the doming method and the basement membrane shape on organoids formation have not been deeply determined. In that sense, the paraffin film method can be considered for use when the other two methods have been unsuccessful.
For all three different methods, this protocol recommends using a lower volume of media and a higher volume of basement membrane for the resuspension process because the domes will last longer in culture. A dome is formed with a 1:4 ratio of media and basement membrane. Of note, this method increases the formation of bubbles while pipetting the cell suspension up and down since the added volume of media does not circumvent the viscosity of the basement membrane. Therefore, it is recommended to set up 1.5 mL tubes such that there is one cell pellet per each dome instead of having one cell pellet for several domes in one 1.5 mL tube. In doing so, the formation of bubbles in one tube for one dome will not serially affect other domes. If the duration of the culture condition has been previously optimized and is known to be short (up to a month), the basement membrane can be diluted more to circumvent the viscosity issue of the basement membrane. A cell pellet resuspension for several wells can be prepared in one tube, utilizing a higher volume of media to the desired volume of basement membrane to minimize cell loss and variability.
To minimize bubble formation during the pipetting process, pipette a lower volume than the actual volume of the cells, media and basement membrane mixture. For example, for a dome within 24 well plates, 40 µL of basement membrane and up to 10 µL of adDMEM complete media are mixed with the cell pellet. In this case, the pipette can be set to handle a volume of 40 µL instead of 50 µL while pipetting up and down in order to minimize bubble formation. For this second method, the cell pellet resuspension for several wells can be prepared in one tube to minimize cell loss and variability. The liquid status of commercially available basement membrane is often variable. Therefore, new batches of basement membrane need to be tested for the doming process. If the basement membrane is more viscous than usual, then up to additional 10 µL of adDMEM complete media (details written in step 1) can be added to dilute the mixture of cell pellet and basement membrane.
Here, we present a protocol to establish, image and process cultured 3D organoid with detailed tips for successful completion of the desired procedures. Culture media for 3D organoids introduced in our manuscript is specifically for prostate-derived cells. However, other details described in our protocol are applicable to different types of tumor tissues and metastatic sites.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This study was supported by the Leo and Anne Albert Charitable Foundation and JM Foundation. We thank the University of California San Diego Moores Cancer Center members, Dr. Jing Yang and Dr. Kay T. Yeung for allowing us the use of their microtome and Randall French, Department of Surgery for technical expertise.
Materials
| 1 mL Pipettman | Gilson | F123602 | |
| 1 mL Syringe | BD Syringe | 329654 | |
| 1.5 mL tube | Spectrum Lab Products | 941-11326-ATP083 | |
| 25G Needle | BD PrecisionGlide Needle | 305122 | |
| 4% Paraformaldehyde (PFA) | Alfa Aesar | J61899 | |
| 70% Ethanol (EtOH) | VWR | BDH1164-4LP | |
| A83-01 | Tocris Bioscience | 2939 | |
| Accumax | Innovative Cell Technologies, Inc. | AM105 | |
| adDMEM | Life Technologies | 12634010 | |
| Agarose | Lonza | 50000 | |
| Antibody -for Cytokeratin 5 | Biolegend | 905901 | |
| Antibody for Cytokeratin 8 | Biolegend | 904801 | |
| B27 | Life Technologies | 17504044 | |
| Bioluminescence imaging system, IVIS 200 | Perkin Elmer Inc | IVIS 200 | |
| Cell Culture Plate – 24 well | Costar | 3524 | |
| Cell Culture Plate – 48 well | Costar | 3548 | |
| Cell Culture Plate – 6 well | Costar | 3516 | |
| Cell Dissociation Solution, Accumax | Innovative Cell Technologies, Inc. | AM105 | |
| Cell Recovery Solution | Corning | 354253 | |
| Cell Scraper | Sarstedt | 83.180 | |
| Cell Strainer | Falcon (Corning) | 352350 | |
| CO2 incubator | Fisher Scientific | 3546 | |
| DAPI | Vector Vectashield | H-1200 | |
| DHT | Sigma-Aldrich | D-073-1ML | |
| dPBS | Corning/Cellgro | 21-031-CV | |
| EGF | PeproTech | AF-100-15 | |
| FBS | Gemini Bio-Products | 100-106 | |
| FGF10 | PeproTech | 100-26 | |
| FGF2 | PeproTech | 100-18B | |
| Forceps | Denville Scientific | S728696 | |
| Glutamax | Gibco | 35050-061 | |
| HEPES | Gibco | 15630-080 | |
| LS Columns | Miltenyi | 130-0420401 | |
| Magnetic Column Seperator: QuadroMACS Separator | Miltenyi | 130-090-976 | |
| Marker | VWR | 52877-355 | |
| Matrigel (Growth Factor Reduced) | Mediatech Inc. (Corning) | 356231 | |
| Matrigel (High Concentration) | BD (Fisher Scientific) | CB354248 | |
| Microscope Imaging Software, Keyence | BZ-X800 (newest software) BZ-X700 (old software) | ||
| Microscope, Keyence | BZ-X700 (model 2016-2017)/BZ-X710 (model 2018-2019) | ||
| Mouse Cell Depletion Kit | Miltenyi | 130-104-694 | |
| N-Acetylcysteine | Sigma-Aldrich | A9165-5G | |
| Nicotinamide | Sigma-Aldrich | N0636-100G | |
| Noggin | PeproTech | 120-10C | |
| OCT Compound | Tissue-Tek | 4583 | |
| Parafilm | American National Can | N/A | |
| Pen-Strep | Mediatech Inc. (Corning) | 30-002-CI-1 | |
| Pipette tipes for 1 mL (Blue Tips) | Fisherbrand Redi-Tip | 21-197-85 | |
| Plunger (from 3 mL syringe) | BD Syringe | 309657 | |
| Prostaglandin E2 | Tocris Bioscience | 2296 | |
| R-Spondin 1 | Trevigen | 3710-001-01 | |
| SB2021190 | Sigma-Aldrich | S7076-25MG | |
| Small Table Top Centrifuge | ThermoFisher Scientific | 75002426 | |
| Water Bath | Fisher Sci | 2320 | |
| Y-27632 Dihydrochloride | Abmole Bioscience | M1817 |
References
- Fatehullah, A., Tan, S. H., Barker, N. Organoids as an in vitro model of human development and disease. Nature Cell Biology. 18 (3), 246-254 (2016).
- Tushir, J. S., et al. Unregulated ARF6 activation in epithelial cysts generates hyperactive signaling endosomes and disrupts morphogenesis. Molecular Biology of the Cell. 21 (13), 2355-2366 (2010).
- Karthaus, W. R., et al. Identification of multipotent luminal progenitor cells in human prostate organoid cultures. Cell. 159 (1), 163-175 (2014).
- McCray, T., Richards, Z., Marsili, J., Prins, G. S., Nonn, L. Handling and Assessment of Human Primary Prostate Organoid Culture. Journal of Visualized Experiments. (143), 59051 (2019).
- Drost, J., et al. Organoid culture systems for prostate epithelial and cancer tissue. Nature Protocols. 11 (2), 347-358 (2016).
- Gao, D., et al. Organoid cultures derived from patients with advanced prostate cancer. Cell. 159 (1), 176-187 (2014).
- Vlachogiannis, G., et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science. 359 (6378), 920-926 (2018).
- Cheung, K. J., Gabrielson, E., Werb, Z., Ewald, A. J. Collective invasion in breast cancer requires a conserved basal epithelial program. Cell. 155 (7), 1639-1651 (2013).
- Abou-Kheir, W. G., Hynes, P. G., Martin, P. L., Pierce, R., Kelly, K. Characterizing the contribution of stem/progenitor cells to tumorigenesis in the Pten-/-TP53-/- prostate cancer model. Stem Cells. 28 (12), 2129-2140 (2010).
- Beshiri, M. L., et al. A PDX/Organoid Biobank of Advanced Prostate Cancers Captures Genomic and Phenotypic Heterogeneity for Disease Modeling and Therapeutic Screening. Clinical Cancer Research. 24 (17), 4332-4345 (2018).
- Debnath, J., Brugge, J. S. Modelling glandular epithelial cancers in three-dimensional cultures. Nature Reviews Cancer. 5 (9), 675-688 (2005).
- Lee, S. H., et al. Tumor Evolution and Drug Response in Patient-Derived Organoid Models of Bladder Cancer. Cell. 173 (2), 515-528 (2018).
- Puca, L., et al. Patient derived organoids to model rare prostate cancer phenotypes. Nature Communications. 9 (1), 2404 (2018).
- Murrow, L. M., Weber, R. J., Gartner, Z. J. Dissecting the stem cell niche with organoid models: an engineering-based approach. Development. 144 (6), 998-1007 (2017).
- Neal, J. T., et al. Organoid Modeling of the Tumor Immune Microenvironment. Cell. 175 (7), 1972-1988 (2018).
- Simmons, J. K., et al. Animal Models of Bone Metastasis. Veterinary Pathology. 52 (5), 827-841 (2015).
- Godebu, E., et al. PCSD1, a new patient-derived model of bone metastatic prostate cancer, is castrate-resistant in the bone-niche. Journal of Translational Medicine. 12, 275 (2014).
- . Keyence Fluorescence Microscope Available from: https://www.keyence.com/ss/products/microscope/bz-x/ (2019)

