Tissue Collection and RNA Extraction from the Human Osteoarthritic Knee Joint
Summary
Primary tissues obtained from patients following total knee arthroplasty provide an experimental model for osteoarthritis research with maximal clinical translatability. This protocol describes how to identify, process, and isolate RNA from seven unique knee tissues to support mechanistic investigation in human osteoarthritis.
Abstract
Osteoarthritis (OA) is a chronic and degenerative joint disease most often affecting the knee. As there is currently no cure, total knee arthroplasty (TKA) is a common surgical intervention. Experiments using primary human OA tissues obtained from TKA provide the capability to investigate disease mechanisms ex vivo. While OA was previously thought to impact mainly the cartilage, it is now known to impact multiple tissues in the joint. This protocol describes patient selection, sample processing, tissue homogenization, RNA extraction, and quality control (based on RNA purity, integrity, and yield) from each of seven unique tissues to support disease mechanism investigation in the knee joint. With informed consent, samples were obtained from patients undergoing TKA for OA. Tissues were dissected, washed, and stored within 4 h of surgery by flash freezing for RNA or formalin fixation for histology. Collected tissues included articular cartilage, subchondral bone, meniscus, infrapatellar fat pad, anterior cruciate ligament, synovium, and vastus medialis oblique muscle. RNA extraction protocols were tested for each tissue type. The most significant modification involved the method of disintegration used for low-cell, high-matrix, hard tissues (considered as cartilage, bone, and meniscus) versus relatively high-cell, low-matrix, soft tissues (considered as fat pad, ligament, synovium, and muscle). It was found that pulverization was appropriate for hard tissues, and homogenization was appropriate for soft tissues. A proclivity for some subjects to yield higher RNA integrity number (RIN) values than other subjects consistently across multiple tissues was observed, suggesting that underlying factors such as disease severity may impact RNA quality. The ability to isolate high-quality RNA from primary human OA tissues provides a physiologically relevant model for sophisticated gene expression experiments, including sequencing, that can lead to clinical insights that are more readily translated to patients.
Introduction
The knee is the largest synovial joint in the human body, comprising the tibiofemoral joint between the tibia and the femur and the patellofemoral joint between the patella and the femur1. The bones in the knee are lined with articular cartilage and supported by various connective tissues, including menisci, fat, ligaments, and muscle, and a synovial membrane encapsulates the whole joint to create a synovial fluid-filled cavity1,2,3 (Figure 1). A healthy knee functions as a mobile hinge joint that allows frictionless motion in the frontal plane1,3. Under pathological conditions, movement can become restricted and painful. The most common degenerative knee joint disease is osteoarthritis (OA)4. A variety of risk factors are known to predispose to OA development, including older age, obesity, female sex, joint trauma, and genetics, among others5,6. There are currently an estimated 14 million people in the USA with symptomatic knee OA, with the prevalence increasing due to rising population age and rates of obesity7,8. Initially considered to be a disease of the cartilage, OA is now understood as a disease of the whole joint9. Commonly observed pathological changes in OA include articular cartilage erosion, osteophyte formation, subchondral bone thickening, and inflammation of the synovium9,10. Since there is no known cure for OA, treatments primarily focus on symptom (e.g., pain) management11,12, and once OA has progressed to end-stage, joint replacement surgery is often indicated13.
Joint replacement surgeries can either be partial or total knee replacements, with total knee arthroplasty (TKA) including replacing the entire tibiofemoral articulation and the patellofemoral joint. As of 2020, approximately 1 million TKAs are performed in the USA each year14. During TKA, an orthopedic surgeon resects the upper portion of the tibial plateau and the lower femoral condyles (Figure 2A, 2B) to be fitted with prosthetic implants. Sometimes misinterpreted by patients, in a TKA, only 8-10 mm is resected from the end of each bone, which is subsequently capped or resurfaced, with metal. An interposed polyethylene liner forms the bearing surface (i.e., padding) between the two metal implants. In addition, several soft tissue components of the joint are fully or partially excised to achieve proper joint balance. Among these tissues are the medial and lateral menisci (Figure 2C), infrapatellar fat pad (Figure 2D), anterior cruciate ligament (ACL; Figure 2E), synovium (Figure 2F), and vastus medialis oblique muscle (VMO; Figure 2G)15. Though TKAs are generally successful for OA treatment, around 20% of patients report reoccurrence of pain post-surgery16. Along with the high cost and relative invasiveness of the procedure, these limitations point to the need for further research to identify alternative treatments to mitigate the progression of OA.
To explore disease mechanisms in OA that may present new avenues for therapeutic intervention, experimental systems, including cells, tissue explants, and animal models can be used. Cells are typically cultured in monolayer and are derived from primary human or animal tissues (e.g., chondrocytes isolated from cartilage) or immortalized cells (e.g., ATDC517 and CHON-00118). While cells can be useful for manipulating experimental variables in a controlled culture environment, they do not capture conditions of the natural joint which are known to impact cell phenotypes19. To better recapitulate the complex cascade of chemical, mechanical, and cell-to-cell communication underlying OA, an alternative is found in primary human or animal tissue samples, whether used fresh or cultured ex vivo as explants, to preserve tissue structure and the cell microenvironment20. In order to study the joint in vivo, small (e.g., mouse21) and large (e.g., horse22) animal models for OA (e.g., through surgical induction, genetic alteration, or aging) are also useful. However, translation from these models to human disease can be limited by anatomical, physiological, and metabolic differences, among others23. Considering the advantages and disadvantages of experimental systems, the key strengths of being species-specific and maintaining the extracellular niche offered by the primary human OA tissues maximize the translational potential of research findings.
Primary human OA tissues can be readily obtained following TKA, making the high frequency of TKAs a valuable resource for research. Among potential experimental applications are gene expression and histological analyses. To realize the potential of primary human OA tissues for these research approaches and others, outlined are the following key considerations. First, the use of patient specimens is subject to ethical regulation, and protocols must meet Institutional Review Board (IRB) approvals24. Second, the inherent heterogeneity of human primary diseased tissues and the influence of variables such as age and sex, among others, create the need for careful patient selection (i.e., application of eligibility criteria) and data interpretation. Third, the unique biological properties of different tissues in the joint (e.g., low cellularity of cartilage and meniscus25) can present challenges during experiments (e.g., isolating high quality and quantity of RNA). This report addresses these considerations and presents a protocol for patient selection, sample processing, tissue homogenization, RNA extraction, and quality control (i.e., assessment of RNA purity and integrity; Figure 3) to encourage the use of primary human OA tissues in the research community.
Protocol
This study protocol was approved and followed institutional guidelines set by the Henry Ford Health System Institutional Review Board (IRB #13995).
1. Patient selection
- Identify the patients from among those scheduled to undergo TKA with an orthopedic surgeon.
- Select the patients based on the eligibility criteria defined by the study protocol. Examples of inclusion criteria include being 18 years of age or older and having a confirmed diagnosis of knee osteoarthritis. Examples of exclusion criteria include undergoing partial knee replacement or having a confirmed diagnosis of rheumatoid arthritis.
- Contact the patients to obtain informed consent prior to the surgery.
2. Sample processing (for RNA)
NOTE: Perform all tissue processing in a class II biosafety cabinet and follow sterile techniques. Always wear appropriate PPE (nitrile gloves, lab coat, safety goggles) when processing human samples. Several bone fragments are produced during TKA, a large amount of bone/cartilage will potentially be available for dissection. Due to disease progression, articular cartilage degeneration may be more severe on some bone portions, which can be factored into experimental design. Only tissues that mandate electrocautery for resection have thermal edge damage, and a concerted surgical effort is made to procure most tissues with a scalpel to minimize damage. Resected tissues must be kept hydrated at all times with sterile PBS.
- Disinfect all work surfaces and equipment with 70% ethanol, RNase decontaminant, DEPC-treated water, and again with 70% ethanol. Wipe away residual liquid with clean, lint-free tissues. Forceps, bone cutters, and scalpels are either autoclaved or soaked in 70% ethanol for at least 10 min prior to use. Keep equipment submerged in 70% ethanol when not in immediate use.
- Pre-label at least three cryovials with de-identified sample name and aliquot number for each tissue (e.g., TKA-1 Cartilage 1).
- Identify each tissue type from the specimen based on the differences in size, shape, color, and texture as shown and described in Figure 2. Identify the cartilage (Figure 2A arrow), bone (Figure 2B arrow), meniscus (Figure 2C), infrapatellar fat pad (Figure 2D), anterior cruciate ligament (Figure 2E), synovium (Figure 2F), and the vastus medialis oblique muscle (Figure 2G).
- Isolate the articular cartilage.
- Select a bone portion with minimal cartilage degradation.
- Using a No.10 scalpel, cut through the cartilage depth as far as possible to dissect the full thickness of the cartilage layer.
NOTE: A No.10 scalpel will only penetrate cartilage, not bone. - Using the full thickness of the cartilage, dissect three 5 mm cubes.
- Isolate the subchondral bone.
- Use the same bone section from which cartilage was collected.
- Using a No.10 scalpel, scrape any remaining cartilage and residual tissues from the bone surface.
- Hold the bone portion to be cut with forceps and use the bone cutters to cut three 5 mm cubes.
- Isolate the meniscus.
- Identify a relatively undamaged portion of either medial or lateral meniscus.
- Using a No.10 scalpel and forceps, dissect three 5 mm cubes.
- Isolate the infrapatellar fat pad.
- Using a No.10 scalpel and forceps, cut the yellow portion of the tissue into three equal-sized and homogenous portions (~500 mg each).
- Isolate the anterior cruciate ligament (ACL).
- Using a No.10 scalpel and forceps, dissect three 5 mm cubes.
- Isolate the synovium.
- Using a No.10 scalpel and forceps, isolate the pink cellular portion of the membrane as much as possible by scraping away fat tissue.
- Dissect three equal-sized and homogenous portions (~200 mg each).
- Isolate the vastus medialis oblique muscle (VMO).
- Using a No.10 scalpel and forceps, remove any fat tissue from the specimen, leaving only the red muscle tissue.
- Dissect three 5 mm cubes.
NOTE: The initial size of the tissue limits the size of the portions.
- Rinse the tissue portions with sterile PBS to remove any residue or debris.
- To perform histology, fix each tissue portion as described in Part 3.
- To perform RNA extraction, cut each of the three tissue portions into smaller pieces (~1-2 mm cubes).
- Transfer the smaller pieces to a 2 mL cryovial, secure caps tightly, flash freeze by submerging in liquid nitrogen for 30 s, and then transfer to a -80 °C freezer for long-term storage (up to 4 months tested in the current protocol). Repeat this for all the aliquots of all the tissues.
- Continue to Tissue homogenization as described in Part 4.
3. Sample processing (for histology)
CAUTION: Formalin is a hazardous chemical, only use in a chemical fume hood.
- Fill pre-labeled 15 mL conical tubes with 10% formalin solution.
- Using forceps, transfer the tissue section to the formalin-filled tubes.
- Fix the tissues in formalin for 1 week at room temperature while shaking/agitating if possible.
- After 1 week, discard the formalin into proper chemical waste disposal, rinse the tissues with PBS, and then transfer to a fresh 15 mL conical tube containing 70% ethanol for long-term storage at 4 °C until samples are embedded for sectioning.
NOTE: For bone/cartilage only, perform decalcification as follows. - After 1 week, discard the formalin into proper chemical waste disposal, and rinse the tissues with PBS.
- Transfer the tissue to a 50 mL conical tube with 45 mL of 10% EDTA solution (pH 7.4).
- If shaking/agitation is not possible, invert the tubes 10-15 times once daily.
- Discard and replace the EDTA solution once weekly.
- Twice weekly, test the texture with an instrument (i.e., spatula) or gloved finger to confirm decalcification by observing deformation when pressure is applied. Time required to decalcify bone tissue will vary between samples from 4-6 weeks.
- Once decalcified, rinse the tissue with PBS and transfer to a fresh 15 mL conical tube containing 70% ethanol for long-term storage at 4 °C until samples are embedded for sectioning.
4. Tissue homogenization
CAUTION: The protocol uses the hazardous chemical phenol. Work with phenol must be performed in a chemical fume hood.
NOTE: Thoroughly clean all equipment and surfaces to be used with 70% ethanol (soak for a minimum of 10 min), followed by RNase decontaminant (soak for a minimum of 10 min), rinse with DEPC-treated water, wipe with a clean, lint-free paper towel, and then respray or soak with 70% ethanol.
- Hard tissue homogenization (articular cartilage, subchondral bone, meniscus)
- Before homogenization, chill the mortar, pestle, and spatula using liquid nitrogen. These must be kept as cold as possible to prevent sample thaw.
- Process the samples one at a time, keeping the other samples at -80 °C until used.
- Transfer the tissue sample to mortar using a chilled spatula; pour additional liquid nitrogen on top of the tissue and allow it to evaporate. Crush the tissue using the pestle. Repeatedly add more liquid nitrogen to the tissue sample, allow it to evaporate, and then continue grinding with the pestle to make a fine powder.
- After powdering the tissue as much as possible, transfer to a pre-chilled 1.5 mL microcentrifuge tube.
- Pre-chill tubes by submerging in liquid nitrogen for 30 s prior to tissue transfer.
- Add 1 mL of the acid-guanidinium-phenol solution to each tube and keep it on ice.
- Repeat steps 4.1.3-4.1.5 for each hard tissue sample.
- After each tissue, clean the mortar, pestle, and spatula with 70% ethanol, RNase decontaminant, DEPC-treated water, and an additional soak in 70% ethanol. Wipe any residual liquid with a clean, lint-free tissue.
- Incubate the samples on ice for an additional 20 min.
- Soft tissue homogenization (infrapatellar fat pad, ACL, synovium, VMO)
- Disinfect the homogenizer by running tubes of 70% ethanol, RNase decontaminant, DEPC-treated water, and an additional 70% ethanol wash, each for 30 s. Wipe any residual liquid with a clean, lint-free tissue. Repeat this between each sample.
- Pre-label 5 mL round bottom tubes for each sample. Add 1 mL of the acid-guanidinium-phenol solution to each tube.
- Work on one sample at a time, keeping other samples to be processed at -80 °C until use.
- Transfer the tissue to a pre-labeled 5 mL tube with acid-guanidinium-phenol.
- Homogenize the tissues in 30 s pulses, keeping on ice during and between pulses. Repeat until the tissue is visually dissolved or for a maximum of five 30 s pulses.
NOTE: Some fibrous tissues (i.e., muscle) may not thoroughly homogenize. - Incubate the dissolved tissue on ice and move to the next sample.
- Clean the homogenizer as described in step 4.2.1. Ensure tissue chunks do not remain in the teeth of the probe; remove with sterile forceps if needed.
- After homogenization of all the samples, incubate on ice in acid-guanidinium-phenol for an additional 20 min.
- Transfer the samples from round bottom tubes to pre-labeled and pre-chilled 1.5 mL microcentrifuge tubes.
5. RNA extraction from tissues
CAUTION: This protocol uses hazardous chemicals such as phenol, chloroform, and isopropanol. Perform all the work in a chemical fume hood.
NOTE: Equipment and reagents are reserved for RNA work only and must be of proper chemical grade for molecular applications (i.e., sterile, nuclease-free). This protocol succeeds both Parts 4.1 and 4.2 tissue homogenization protocols. Thoroughly clean all equipment and surfaces to be used with 70% ethanol (soak for a minimum of 10 min), followed by RNase decontaminant (soak for a minimum of 10 min). Rinse with DEPC-treated water, wipe the residual liquid with a clean, lint-free paper towel, and then respray or soak with 70% ethanol.
- Centrifuge the microcentrifuge tubes at 10000 x g for 10 min at 4 °C to pellet the debris.
- Transfer the supernatant to a fresh 1.5 mL microcentrifuge tube.
NOTE: For fatty tissues, a lipid layer will sometimes be present on the top. Avoid transferring this by piercing the layer on the side of the tube with a pipette tip. - Add 200 µL of chloroform per 1 mL of the acid-guanidinium-phenol solution to each sample. Shake the tubes vigorously by hand for 30 s to mix. Then, incubate on ice for 2 min.
- Centrifuge samples at 10000 x g for 12 min at 4 °C.
NOTE: After centrifugation, three layers will have formed: an aqueous phase containing RNA, a white DNA interphase, and a pink protein phase at the bottom. - Transfer ~500 µL of the upper, aqueous phase to a fresh 1.5 mL microcentrifuge tube.
NOTE: Do not disturb the interphase and the protein fractions. Store these phases at -80 °C for future DNA or protein isolation. - Add an equal volume of acid-guanidinium-phenol solution to the transferred aqueous phase, mix by inverting the tube 8-10 times. Incubate the tube on ice for 20 min.
- Add 200 µL of chloroform per 1 mL of the acid-guanidinium-phenol solution to each sample. Shake vigorously by hand for 30 s to mix and then incubate on ice for 2 min.
- Centrifuge samples at 10000 x g for 12 min at 4 °C.
- Transfer <500 µL of the aqueous phase to a fresh microcentrifuge tube, careful not to contaminate the sample with the other phases.
CAUTION: Discard the remaining phases with appropriate hazardous material disposal methods. - Add an equal volume (as aqueous phase) of 100% isopropanol to each sample. Mix by inverting the tube 8-10 times. Incubate on ice for 5 min.
- Add 1 µL of glycogen coprecipitant to each sample to aid in locating the RNA pellet post-centrifugation.
- Centrifuge the samples at 12000 x g for 25 min at 4 °C.
- Locate the pellet in the tube (if using coprecipitant, it will appear blue). Carefully pour off the supernatant.
- Wash the pellet by adding 1 mL of ice-cold 75% ethanol to each sample, vortex to dislodge the pellet from the bottom of the tube.
NOTE: Prepare 75% ethanol using molecular grade pure ethanol and nuclease-free water. - Centrifuge at 7000 x g for 5 min at 4 °C. Carefully pour off the supernatant.
- Repeat steps 5.13 and 5.14 two more times.
- Quick spin (<2000 x g for 5 s) to bring any residual liquid to the bottom of the tube. Use a P20 pipette to remove any residual ethanol from the bottom of the tubes.
- Avoid touching the RNA pellet with a pipette tip. Change tips between samples.
- With tube caps open, air dry samples at room temperature for 10 min.
NOTE: Pellet may become translucent as it dries. Ensuring all residual ethanol has evaporated will improve RNA purity. - Add 25 µL of nuclease-free water to each tube to dissolve the pellet.
- Incubate at room temperature for 5 min.
- Gently pipette up and down to mix the RNA.
- Aliquot 5 µL of the sample into a fresh tube for quality control analysis (Part 6). Store the remaining 20 µL at -80 °C for gene expression assays.
6. Quality control
- Determine the concentration and purity of RNA using a spectrophotometer according to the manufacturer's instructions.
- Determine the RNA integrity using an electrophoresis device according to the manufacturer's instructions.
NOTE: Dilute the RNA to be within the range of the chip detection limits.
Representative Results
Seven unique human knee joint tissues are available for collection from patients undergoing TKA for OA (Figure 1). In this protocol, each of these tissues were identified and processed within 4 h of surgical removal (Figure 2). Following the steps outlined in Figure 3, portions of each tissue were formalin-fixed for histological assessment (Figure 4), while other portions were flash-frozen for RNA isolation. Separating hard tissues from soft tissues by method of disintegration (pulverization versus homogenization, respectively), RNA of high integrity and purity was extracted from each tissue type, with representative results shown in Table 1 (High quality columns). Notably, some subjects yielded lower quality RNA across multiple tissues (Table 1, Low quality columns), suggesting that despite an optimized method, external factors (e.g., disease severity) may be impacting RNA quality across tissue types.
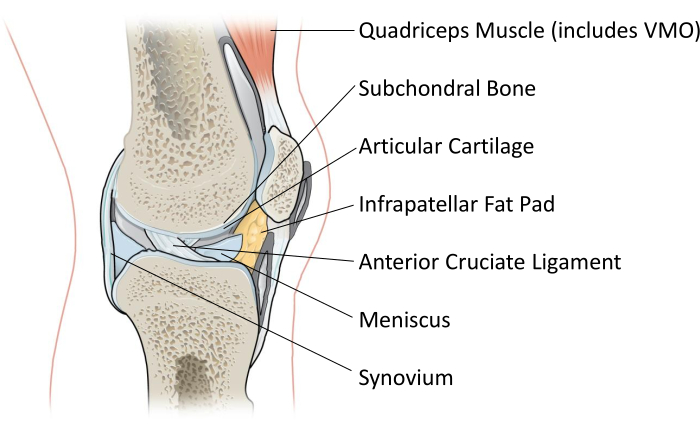
Figure 1: Schematic of the human knee joint showing a lateral cross-section. Each of the seven labeled tissues are collected during TKA and used for research purposes as described in this protocol. VMO = vastus medialis oblique muscle. Image accessed and modified from OpenStax College under a creative commons license26. Please click here to view a larger version of this figure.
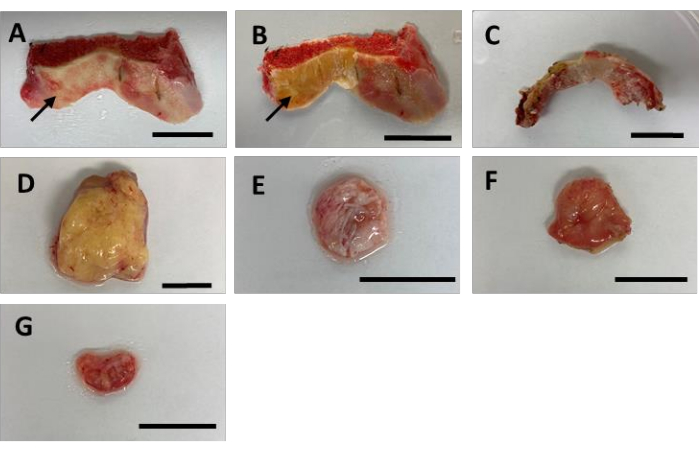
Figure 2: Representative gross images for each of the seven tissues obtained from patients undergoing TKA. (A) Anterior femoral bone cut with arrow pointing to articular cartilage to be collected. Cartilage is identified by a whitish layer found on the surface of the bone. (B) Anterior femoral bone cut with arrow pointing to subchondral bone to be collected. (C) Meniscus. Avoid collecting burnt sections caused by electrocauterization during TKA. (D) Infrapatellar fat pad (yellow in color). (E) Anterior cruciate ligament (white, fibrous, spongy tissue). (F) Synovium. One side will appear light colored and fibrous, often containing fat tissue, while the opposite side will appear pinkish and less fibrous. The pink side of the membrane contains the synovial lining. (G) Vastus medialis oblique muscle (red). This may often be the smallest tissue portion and may contain some fat tissue. Scale bar = 2 cm. Please click here to view a larger version of this figure.
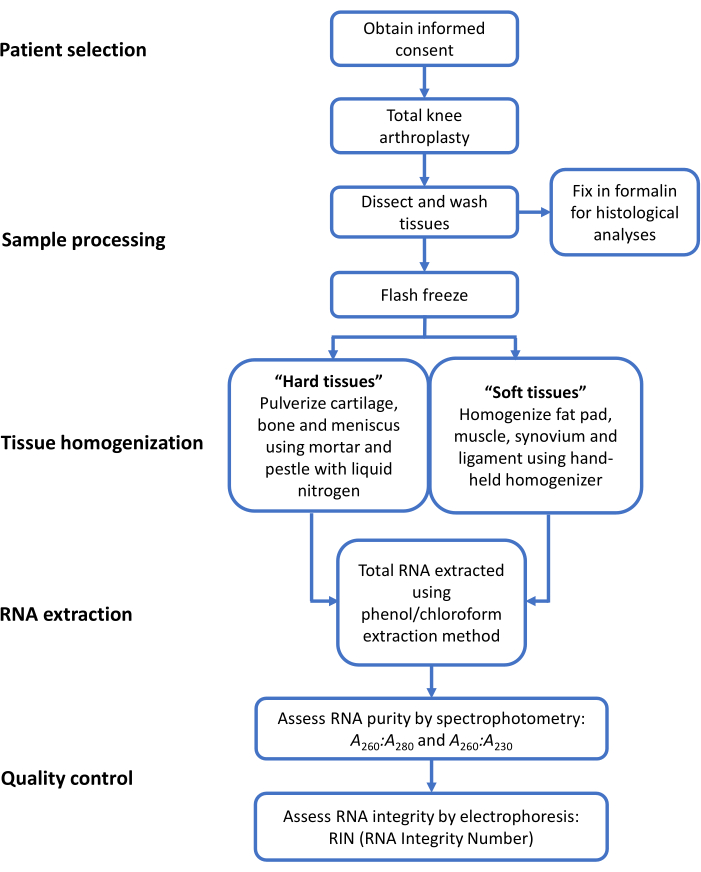
Figure 3: Overview of steps taken to collect primary human OA tissues for histology and RNA. This protocol describes patient selection, sample processing for histology and RNA, tissue homogenization for hard tissues and soft tissues, RNA extraction, and quality control for seven primary human knee OA tissues. Please click here to view a larger version of this figure.
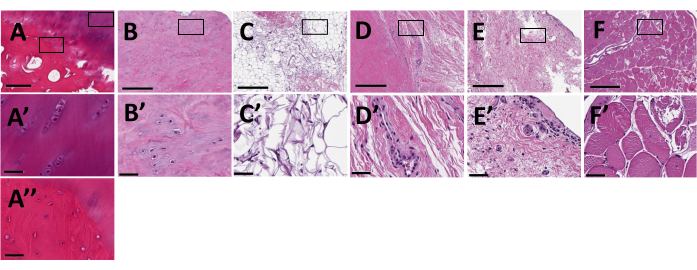
Figure 4: Histological sections for each of the seven tissues obtained from patients undergoing TKA. Hematoxylin and eosin-stained sections are shown at 6x magnification in panels A–F with inset magnified to 40x in panels A'–F' and A". (A, A') Articular cartilage; (A, A") subchondral bone; (B, B') meniscus; (C, C') infrapatellar fat pad; (D, D') anterior cruciate ligament; (E, E') synovium; (F, F') vastus medialis oblique muscle. Scale bars = 400 µm for A-F and 50 µm for A'-F' and A". Please click here to view a larger version of this figure.
| Tissue | N | RIN | [RNA] (ng/µL) in 25 µL | A260:A280 | A260:A230 | ||||||||
| High quality | Low quality | High quality | Low quality | High quality | Low quality | High quality | Low quality | High quality | Low quality | ||||
| "Hard tissues" | |||||||||||||
| Articular Cartilage | 10 | 4 | 7.0 ± 0.8 | 1.3 ± 1.2 | 135 (36-243) |
46 (26-78) |
1.80 ± 0.09 | 1.47 ± 0.34 | 1.40 ± 0.36 | 0.45 ± 0.26 | |||
| Subchondral Bone | 10 | 4 | 7.8 ± 0.6 | 3.6 ± 1.1 | 514 (181-1586) |
342 (122-769) |
1.96 ± 0.05 | 1.90 ± 0.17 | 1.72 ± 0.27 | 1.12 ± 0.62 | |||
| Meniscus | 10 | 4 | 7.5 ± 0.6 | 2.4 ± 0.3 | 242 (38-1629) |
95 (60-318) |
1.91 ± 0.05 | 1.71 ± 0.15 | 1.41 ± 0.47 | 0.77 ± 0.59 | |||
| "Soft tissues" | |||||||||||||
| Infrapatellar Fat Pad | 10 | 3 | 8.5 ± 0.6 | 6.1 ± 1.4 | 1905 (668-5100) |
1151 (381-2306) |
1.99 ± 0.01 | 2.02 ± 0.03 | 2.09 ± 0.17 | 1.47 ± 0.71 | |||
| Anterior Cruciate Ligament | 8 | 3 | 7.4 ± 0.4 | 5.2 ± 1.9 | 1836 (613-8456) |
727 (97-1479) |
1.97 ± 0.03 | 1.91 ± 0.18 | 1.88 ± 0.20 | 1.35 ± 0.70 | |||
| Synovium | 9 | 3 | 8.5 ± 0.6 | 6.2 ± 3.1 | 2239 (401-4100) |
1897 (902-3366) |
1.99 ± 0.04 | 2.00 ± 0.03 | 2.05 ± 0.11 | 1.91 ± 0.20 | |||
| Vastus Medialis Oblique | 9 | 3 | 8.5 ± 0.6 | 8.4 ± 0.7 | 1002 (377-1715) |
1097 (308-2138) |
1.96 ± 0.03 | 1.99 ± 0.03 | 1.82 ± 0.11 | 1.62 ± 0.27 | |||
Table 1. Quality and quantity of RNA isolated from OA tissues collected from TKA patients. RIN = RNA Integrity Number. Data presented as mean ± standard deviation for RIN, A260:A280 and A260:A230 ratios, and mean (range) for RNA concentration values. High quality samples consist of patients from which all tissue types yielded RNA with RIN > 6 (n = 8-10). Low quality samples consist of patients from which multiple tissue types yielded RNA with RIN < 6 (n = 3-4).
Discussion
The protocol presented has proved successful for collecting seven primary human OA tissues for RNA extraction (Table 1) and histological processing (Figure 4). Prior to collecting patient samples, it is necessary to establish an IRB-approved protocol, ideally in collaboration with a surgeon or surgical team. Applying a standardized protocol for specimen collection (e.g., resection from consistent in situ locations) is essential for maximizing experimental reproducibility. Tissue samples should be transported to the lab in sterile containers and processed within 4 h of the surgery to avoid degradation. During tissue dissection and processing, all tissues are kept hydrated in sterile PBS and are rinsed in fresh, sterile PBS to remove potential surface contaminants such as biofluids and other unwanted debris prior to being flash-frozen for RNA extraction or formalin-fixed for histology. A useful application of histological analysis is confirmation of the tissue types and the disease severity since these can be distinguished by cell number, distribution, and morphology, among other factors observable by standard stains such as hematoxylin and eosin (Figure 4).
Primary human OA tissues can present challenges for extracting RNA of sufficient quantity and quality, as defined by purity and integrity27. RNA quantity is a function of the overall cellularity of the tissue, and in the knee joint, there are low-cell, high-matrix tissues such as the cartilage, bone, and meniscus, and relatively high-cell, low-matrix tissues such as the fat pad, ACL, synovium, and VMO. For example, both articular hyaline cartilage and meniscal fibrocartilage28 are characterized by low cellularity, with the extracellular matrix containing varying quantities of collagens, proteoglycans, and other glycoproteins28,29. Having fewer cells results in less RNA per volume of tissue (reducing quantity) and having more protein results in co-purification with RNA (reducing purity)25,30. RNA purity can be determined by spectrophotometry where A260:A280 and A260:A230 values of <1.5 reflect the presence of organic contaminants (e.g., protein) and values of ~2.0 reflect pure RNA31. RNA integrity reflects the level of degradation, whether caused by experimental conditions (i.e., shearing forces) or by enzymatic digestion (e.g., nucleases), and is often determined by electrophoretic analysis. An RNA Integrity Number (RIN) of 1 reflects degraded RNA and a RIN of 10 reflects intact RNA31,32. For RNA-sequencing, a minimum RIN of 7 is often recommended33,34,35. Data presented in Table 1 reveals that these A260:A280, A260:A230, and RIN thresholds were met across all tissues from the patient samples in the High quality RNA group compared to patient samples in the Low quality RNA group, except for some A260:A230 values, which may reflect protein contamination of RNA in the low-cell, high-matrix tissues. While there are many factors that could be contributing to the quality of RNA isolated from a given patient sample, among them may be the level of disease severity. The diseased nature of OA tissues suggests degradative processes are occurring through increased levels of enzymes that can digest tissues, but also RNA, thereby reducing quality.
This protocol for RNA extraction aims to maximize RNA quantity and quality from primary human OA tissues. The most critical step pertained to whether the tissues were disintegrated by being pulverized or homogenized, and this was found to correlate with the tissue cellularity and matrix composition. Initially, all seven tissues were subjected to the same protocol where tissues were first pulverized by mortar and pestle using liquid nitrogen, then transferred to acid-guanidinium-phenol solution, and further homogenized using a hand-held tissue homogenizer. This method produced favorable RNA yield, purity, and integrity for the fat pad, ACL, synovium, and VMO (collectively soft tissues; also, relatively high-cell, low-matrix), but unfavorable results for cartilage, bone, and meniscus (collectively hard tissues; also, low-cell, high-matrix). Based on these observations, the seven tissues were divided into two groups for further protocol refinement. It was observed that the additional homogenization had minimal effect on further disintegrating the hard tissues after they had been pulverized into a fine powder. Conversely, dissociation of the soft tissues was successfully achieved with homogenization alone and did not require pulverization. Therefore, the homogenization of the hard tissues and pulverization of the soft tissues was eliminated. This was beneficial for minimizing shearing forces, processing time, and temperature fluctuation, all of which can improve RNA integrity. Two rounds of phenol/chloroform phase separation for all seven tissues were performed, as this has been reported to improve RNA purity without reducing yield31.
A potential limitation of this protocol is the batch effect that may arise from separating the tissues into two groups if the experimental design requires comparison among all tissues. The use of pulverization versus homogenization methods may alter technical (e.g., processing time) and environmental (e.g., temperature fluctuations) conditions that can introduce variability36. A second limitation is the potential inconsistency in identifying, dissecting, and orienting (for histological sectioning) the tissues within and across subjects. A third limitation is our inability to confirm potential correlations between patient disease severity and RNA quality in the current report. A fourth limitation is the lack of availability of healthy control tissues for comparison. Though control samples may be available from cadavers, these are less readily available than OA tissues from TKA. An experimental strategy to circumvent this is to use each subject as their own control, whether making comparisons across tissues or within tissues comparing treatment to control or lesioned to preserved areas. Finally, using tissue explants for RNA extraction does not permit gene expression analysis of the individual cell types that comprise the tissues (e.g., synovial fibroblasts versus synovial macrophages37).
Despite the noted limitations, primary human OA tissues are a valuable resource for research, offering advantages over other experimental systems for OA, including preserving the cell niche23. However, primary human OA tissues may be underutilized in research due to logistic or technical challenges. This protocol describes patient selection, sample processing, tissue homogenization, RNA extraction, and quality control to support the use of the samples obtained from TKA. Following sample processing, several experimental approaches can be pursued, including gene expression and histology, among others. Most relevant to the rapidly evolving omics field is the ability to isolate sufficient quantities of high-quality RNA for applications such as RNA-sequencing38,39. Molecular profiles can be compared within and across tissues from subjects with specific disease phenotypes (e.g., based on age, sex, and other OA risk factors). Insights gained may inform new therapeutic avenues that can be more readily translated back to the OA patient population.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors thank the study participants who made this research possible and dedicate this report to new scientists in the osteoarthritis field.
Materials
| 1.5 mL microcentrifuge tubes | Eppendorf | 05 402 | Sterile, nuclease-free. Reserved for RNA work only. |
| 10% Formalin | Cardinal Health | C4320-101 | Store in chemical cabinet when not in use. |
| 100% Chloroform (Molecular Biology Grade) | Fisher Scientific | ICN19400290 | Sterile, nuclease-free. Reserved for RNA work only, store in chemical cabinet when not in use. |
| 100% Ethanol (Molecular Biology Grade) | Fisher Scientific | BP2818500 | Sterile, nuclease-free. Reserved for RNA work only, when diluting use DEPC/nuclease-free water. |
| 100% Isopropanol (Molecular Biology Grade) | Fisher Scientific | AC327272500 | Sterile, nuclease-free. Reserved for RNA work only, store in chemical cabinet when not in use. |
| 100% Reagent Alcohol | Cardinal Health | C4305 | Diluted to 70% with dH2O for cleaning purposes. |
| 15 cm sterile culture dishes | Thermo Scientific | 12-556-003 | Sterile, nuclease-free. |
| 15 mL polypropylene (Falcon) tubes | Fisher Scientific | 14 959 53A | Sterile, nuclease-free. |
| 2 mL cryovials (externally threaded) | Fisher Scientific | 10 500 26 | Sterile, nuclease-free. |
| 5 mL round-bottom tubes | Corning | 352052 | Sterile, nuclease-free. Reserved for RNA work only. |
| 50 mL polypropylene (Falcon) tubes | Fisher Scientific | 12 565 271 | Sterile, nuclease-free. |
| Bioanalyzer | Agilent | G2939BA | For RNA integrity measurement. |
| Biosafety Cabinet | General lab equipment | ||
| Bone Cutters | Fisher Scientific | 08 990 | Sterilized with 70% EtOH. |
| Chemical Fume Hood | General lab equipment | ||
| Disposable Scalpels (No.10) | Thermo Scientific | 3120032 | Sterile, nuclease-free. |
| EDTA | Life Technologies | 15-576-028 | 10% solution with dH2O. |
| Forceps | Any vendor | Sterilized with 70% EtOH. | |
| Glycoblue Coprecipitant | Fisher Scientific | AM9516 | Reserved for RNA work only, store at -20 °C. |
| Kimwipes | Fisher Scientific | 06-666 | |
| Liquid Nitrogen | Any vendor | ||
| Liquid Nitrogen Dewar | General lab equipment | ||
| Mortar and Pestle | Any vendor | Reserved for RNA work only, sterilzed per protocol. | |
| Nanodrop Spectrophotometer | Thermo Scientific | ND-2000 | For RNA purity and yield measurements. |
| Nuclease-free/DEPC-treated water | Fisher Scientific | Sterile, nuclease-free. Reserved for RNA work only. | |
| PBS (Sterile) | Gibco | 20 012 050 | Sterile, nuclease-free. |
| Pipettes (2 µL, 20 µL, 200 µL, 1000 µL) & tips | Any vendor | Sterile, nuclease-free. | |
| Plasma/Serum Advanced miRNA kit | Qiagen | 217204 | |
| Refrigerated Centrifuge 5810R | Eppendorf | 22625101 | |
| RNAlater | Thermo Scientific | 50 197 8158 | Sterile, nuclease-free. |
| RNAse Away/RNAseZap | Fisher Scientific | 7002 |
|
| Spatula (semimicro size) | Any vendor | Reserved for RNA work only. | |
| Tissue homogenizer | Pro Scientific | 01-01200 | Reserved for RNA work only, sterilzed per protocol. |
| TRIzol Reagent | Fisher Scientific | 15 596 026 | Sterile, nuclease-free. Reserved for RNA work only. |
References
- Gupton, M., Imonugo, O., Terreberry, R. R. . Anatomy, Bony Pelvis, and Lover Limb, Knee. , (2020).
- Pacifici, M., Koyama, E., Iwamoto, M. Mechanisms of Synovial joint and articular cartilage formation: recent advances, but many lingering mysteries. Birth Defects Research Part C: Embryo Today: Reviews. 75 (3), 237-248 (2005).
- Gupton, M., Munjal, A., Terreberry, R. R. . Anatomy, Hinge Joints. , (2020).
- Chen, D., et al. Osteoarthritis: Toward a comprehensive understanding of pathological mechanism. Bone Research. 5, 16044 (2017).
- Murphy, L., et al. Lifetime risk of symptomatic knee osteoarthritis. Arthritis & Rheumatism. 59 (9), 1207-1213 (2008).
- O’Neill, T. W., McCabe, P. S., McBeth, J. Update on the epidemiology, risk factors and disease outcomes of osteoarthritis. Best Practice & Research: Clinical Anaesthesiology. 32 (2), 312-326 (2018).
- Nguyen, U. S., et al. Increasing prevalence of knee pain and symptomatic knee osteoarthritis: survey and cohort data. Annals of Internal Medicine. 155 (11), 725-732 (2011).
- Deshpande, B. R., et al. Number of persons with symptomatic knee osteoarthritis in the us: impact of race and ethnicity, age, sex, and obesity. Arthritis Care & Research. 68 (12), 1743-1750 (2016).
- Loeser, R. F., Goldring, S. R., Scanzello, C. R., Goldring, M. B. Osteoarthritis: a disease of the joint as an organ. Arthritis & Rheumatism. 64 (6), 1697-1707 (2012).
- McGonagle, D., Tan, A. L., Carey, J., Benjamin, M. The anatomical basis for a novel classification of osteoarthritis and allied disorders. Journal of Anatomy. 216 (3), 279-291 (2010).
- Bannuru, R. R., et al. OARSI guidelines for the non-surgical management of knee, hip, and polyarticular osteoarthritis. Osteoarthritis Cartilage. 27 (11), 1578-1589 (2019).
- Kolasinski, S. L., et al. American College of Rheumatology/Arthritis Foundation Guideline for the Management of Osteoarthritis of the Hand, Hip, and knee. Arthritis Care & Research. 72 (2), 220-233 (2020).
- Michael, J. W., Schluter-Brust, K. U., Eysel, P. The epidemiology, etiology, diagnosis, and treatment of osteoarthritis of the knee. Deutsches Ärzteblatt International. 107 (9), 152-162 (2010).
- Singh, J. A., Yu, S., Chen, L., Cleveland, J. D. Rates of total joint replacement in the United States: Future projections to 2020-2040 using the national inpatient sample. Journal of Rheumatology. 46 (9), 1134-1140 (2019).
- Gemayel, A. C., Varacallo, M. Total Knee Replacement Techniques. StatPearls. , (2020).
- Shan, L., Shan, B., Suzuki, A., Nouh, F., Saxena, A. Intermediate and long-term quality of life after total knee replacement: a systematic review and meta-analysis. Journal of Bone and Joint Surgery (American Volume). 97 (2), 156-168 (2015).
- Newton, P. T., et al. Chondrogenic Atdc5 cells: an optimised model for rapid and physiological matrix mineralisation. International Journal of Molecular Medicine. 30 (5), 1187-1193 (2012).
- Chuang, Y. W., et al. Lysophosphatidic acid enhanced the angiogenic capability of human chondrocytes by regulating Gi/Nf-Kb-dependent angiogenic factor expression. PLoS One. 9 (5), 95180 (2014).
- Johnson, C. I., Argyle, D. J., Clements, D. N. In vitro models for the study of osteoarthritis. Veterinary Journal. 209, 40-49 (2016).
- Grivel, J. C., Margolis, L. Use of human tissue explants to study human infectious agents. Nature Protocols. 4 (2), 256-269 (2009).
- Glasson, S. S., Blanchet, T. J., Morris, E. A. The surgical destabilization of the medial meniscus (dmm) model of osteoarthritis in the 129/Svev mouse. Osteoarthritis Cartilage. 15 (9), 1061-1069 (2007).
- McIlwraith, C. W., Frisbie, D. D., Kawcak, C. E. The horse as a model of naturally occurring osteoarthritis. Bone & Joint Research. 1 (11), 297-309 (2012).
- Cope, P. J., Ourradi, K., Li, Y., Sharif, M. Models of Osteoarthritis: The good, the bad and the promising. Osteoarthritis Cartilage. 27 (2), 230-239 (2019).
- Goldenberg, A. J., et al. IRB practices and policies regarding the secondary research use of biospecimens. Bmc Medical Ethics. 16, 32 (2015).
- Ruettger, A., Neumann, S., Wiederanders, B., Huber, R. Comparison of different methods for preparation and characterization of total rna from cartilage samples to uncover osteoarthritis in vivo. BMC Research Notes. 3, 7 (2010).
- Reno, C., Marchuk, L., Sciore, P., Frank, C. B., Hart, D. A. Rapid isolation of total RNA from small samples of hypocellular, dense connective tissues. BioTechniques. 22 (6), 1082-1086 (1997).
- Fox, A. J., Bedi, A., Rodeo, S. A. The basic science of human knee menisci: structure, composition, and function. Sports Health. 4 (4), 340-351 (2012).
- Carballo, C. B., Nakagawa, Y., Sekiya, I., Rodeo, S. A. Basic science of articular cartilage. Clinics in Sports Medicine. 36 (3), 413-425 (2017).
- Le Bleu, H. K., et al. Extraction of high-quality RNA from human articular cartilage. Analytical Biochemistry. 518, 134-138 (2017).
- Ali, S. A., Alman, B. RNA extraction from human articular cartilage by chondrocyte isolation. Analytical Biochemistry. 429 (1), 39-41 (2012).
- Schroeder, A., et al. The Rin: An RNA integrity number for assigning integrity values to rna measurements. BMC Molecular Biology. 7, 3 (2006).
- Li, S., et al. Multi-platform assessment of transcriptome profiling using RNA-seq in the abrf next-generation sequencing study. Nature Biotechnology. 32 (9), 915-925 (2014).
- Nazarov, P. V., et al. RNA sequencing and transcriptome arrays analyses show opposing results for alternative splicing in patient derived samples. BMC Genomics. 18 (1), 443 (2017).
- Madissoon, E., et al. scRNA-seq assessment of the human lung, spleen, and esophagus tissue stability after cold preservation. Genome Biology. 21 (1), (2019).
- Scholes, A. N., Lewis, J. A. Comparison of RNA isolation methods on RNA-seq: implications for differential expression and meta-analyses. BMC Genomics. 21 (1), 249 (2020).
- Smith, M. D. The normal synovium. Open Rheumatology Journal. 5, 100-106 (2011).
- Kukurba, K. R., Montgomery, S. B. RNA sequencing and analysis. Cold Spring Harbor Protocols. 2015 (11), 951-969 (2015).
- Mobasheri, A., Kapoor, M., Ali, S. A., Lang, A., Madry, H. The future of deep phenotyping in osteoarthritis: how can high throughput omics technologies advance our understanding of the cellular and molecular taxonomy of the disease. Osteoarthritis and Cartilage Open. 3 (2), 100144 (2021).

