In Vitro Analysis of E3 Ubiquitin Ligase Function
Summary
The present study provides detailed in vitro ubiquitylation assay protocols for the analysis of E3 ubiquitin ligase catalytic activity. Recombinant proteins were expressed using prokaryotic systems such as Escherichia coli culture.
Abstract
The covalent attachment of ubiquitin (Ub) to internal lysine residue(s) of a substrate protein, a process termed ubiquitylation, represents one of the most important post-translational modifications in eukaryotic organisms. Ubiquitylation is mediated by a sequential cascade of three enzyme classes including ubiquitin-activating enzymes (E1 enzymes), ubiquitin-conjugating enzymes (E2 enzymes), and ubiquitin ligases (E3 enzymes), and sometimes, ubiquitin-chain elongation factors (E4 enzymes). Here, in vitro protocols for ubiquitylation assays are provided, which allow the assessment of E3 ubiquitin ligase activity, the cooperation between E2-E3 pairs, and substrate selection. Cooperating E2-E3 pairs can be screened by monitoring the generation of free poly-ubiquitin chains and/or auto-ubiquitylation of the E3 ligase. Substrate ubiquitylation is defined by selective binding of the E3 ligase and can be detected by western blotting of the in vitro reaction. Furthermore, an E2~Ub discharge assay is described, which is a useful tool for the direct assessment of functional E2-E3 cooperation. Here, the E3-dependent transfer of ubiquitin is followed from the corresponding E2 enzyme onto free lysine amino acids (mimicking substrate ubiquitylation) or internal lysines of the E3 ligase itself (auto-ubiquitylation). In conclusion, three different in vitro protocols are provided that are fast and easy to perform to address E3 ligase catalytic functionality.
Introduction
Ubiquitylation is the process by which Ub is covalently linked to a substrate protein1. The Ub modification is catalyzed by successive enzymatic reactions involving the action of three different enzyme classes, i.e., Ub-activating enzymes (E1s), Ub-conjugating enzymes (E2s), Ub ligases (E3s), and possibly Ub chain elongation factors (E4s)2,3,4,5. After adenosine triphosphate (ATP)- and magnesium (Mg2+)-dependent activation of Ub by E1, the active site cysteine of E1 attacks the C-terminal glycine of Ub, forming a thioester complex (Ub~E1). The energy drawn from ATP hydrolysis causes the Ub to attain a high energy transitional state, which is maintained throughout the following enzyme cascade. Next, the E2 enzyme transfers the activated Ub to its internal catalytic cysteine, thereby forming a transient Ub~E2 thioester bond. Subsequently, Ub is transferred to the substrate protein.
This can be done in two ways. Either the E3 ligase may first bind to E2, or the E3 ligase can directly bind Ub. The latter way results in the formation of an E3~Ub intermediate. In either case, Ub is linked to the substrate protein by formation of an isopeptide bond between the C-terminal carboxyl group of Ub and the lysine Ɛ-amino group of the substrate6. The human genome encodes two E1s, approximately 40 E2s, and more than 600 putative ubiquitin ligases7. Based on the Ub transfer mechanism of the E3, Ub ligases are divided into three categories involving Homologous to E6AP C-Terminus (HECT)-type, Really Interesting New Gene (RING)/U-box-type, and RING between RING (RBR)-type ligases8. In this study, the U-box containing ligase, Carboxyl Terminus of HSC70-interacting Protein (CHIP), is used as a representative E3 enzyme. In contrast to HECT-type E3 enzymes that form Ub~E3 thioesters, the U-box domain of CHIP binds E2~Ub and promotes the subsequent Ub/substrate transfer directly from the E2 enzyme8,9. Based on the importance of the U-box for enzymatic function, an inactive CHIP U-box mutant, CHIP(H260Q), is utilized as a control. CHIP(H260Q) fails to bind to its cognate E2s, thus losing its E3 ligase activity10.
Protein ubiquitylation plays a crucial role in regulating a multitude of cellular events in eukaryotic cells. The diversity of cellular outcomes that are promoted by the reversible attachment of Ub molecules to substrate proteins can be attributed to the molecular characteristics of Ub. As Ub itself contains seven lysine (K) residues for further ubiquitylation, there is rich variety of Ub chain-types with different sizes and/or topologies11. For example, substrates can be modified by a single Ub molecule at one (mono-ubiquitylation) or multiple lysines (multi mono-ubiquitylation), and even by Ub chains (poly-ubiquitylation)11. Ub chains are either formed homo- or heterotypically via the same or different lysine residues of Ub, which could even result in branched Ub chains9. Thus, protein ubiquitylation leads to diverse arrangements of Ub molecules that provide specific information, e.g., for degradation, activation, or localization of conjugated proteins12,13. These different Ub signals enable fast reprogramming of cellular signalling pathways, which is an important requirement for the cell's ability to respond to changing environmental needs.
A central aspect of ubiquitylation is related to protein quality control. Misfolded or irreversibly damaged proteins must be degraded and replaced by newly synthesized proteins to maintain protein homeostasis or proteostasis14. The quality control E3 ligase, CHIP, collaborates with molecular chaperones in the Ub-dependent degradation of damaged proteins9,15,16,17. Apart from that, CHIP regulates the stability of the myosin-directed chaperone, UNC-45B (Unc-45 Homolog B), which is tightly coordinated with muscle function and deviations from the optimal levels lead to human myopathy18,19,20,21. Degradation of UNC-45B by the 26S proteasome is mediated by attachment of a K48-linked poly-Ub chain9. In the absence of substrate proteins, CHIP performs auto-ubiquitylation10,22,23, which is characteristic of RING/U-box E3 ubiquitin ligases24,25 and considered to regulate ligase activity26. Application of the in vitro ubiquitylation assay methods described in this paper helped to systematically identify E2 enzymes that team up with CHIP to promote the formation of free poly-Ub chains and/or auto-ubiquitylation of CHIP (protocol section 2). Furthermore, CHIP-dependent ubiquitylation of UNC-45B was observed, which is a known substrate of the E3 ligase18,19 (protocol section 3). Ultimately, CHIP-dependent transfer of activated Ub from the Ub~E2 thioester was monitored (protocol section 4).
Protocol
1. Preparation of buffers and reagents
NOTE: Buffers and reagents that were manually prepared in the laboratory are listed below. All other buffers and reagents used in the protocols were purchased from different sources and used according to the manufacturers' instructions.
- Prepare 10x phosphate-buffered saline (10x PBS). For this purpose, mix 1.37 M sodium chloride (NaCl), 27 mM potassium chloride (KCl), 80 mM of disodium-hydrogen-phosphate dihydrate (Na2HPO4.2 H2O), and 20 mM of potassium-dihydrogen phosphate (KH2PO4) in 1 L of double-distilled H2O (ddH2O). Autoclave the 10x PBS, and prepare a 1x PBS solution in ddH2O.
- Autoclaving is carried out for 15 min at 121 °C and 98.9 kPa.
NOTE: Autoclaving is carried out under these conditions for all buffers and solutions. If not indicated otherwise, solutions are stored at room temperature (RT).
- Autoclaving is carried out for 15 min at 121 °C and 98.9 kPa.
- Prepare 1 L of PBS-Tween (PBS-T) solution by adding 0.1% Tween to 1 L of 1x PBS.
- Prepare 10 mL of a 0.5 M L-lysine stock solution by dissolving L-lysine in ddH2O. Aliquot L-lysine in 1 mL portions, and store them at -20 °C until further use.
NOTE: L-lysine can be frozen and thawed repeatedly. - Prepare a 0.25 M ethylene diamine tetra acetic acid (EDTA) solution by dissolving EDTA in 200 mL of ddH2O.
- Adjust the pH to 8.0 with sodium hydroxide (NaOH).
- Adjust the volume to 250 mL with ddH2O.
- Autoclave the solution.
- Prepare 10 mL of a 20 mg/mL bovine serum albumin (BSA) solution by dissolving BSA in ddH2O. Store at -20 °C in 200 µL aliquots.
NOTE: BSA can be frozen and thawed repeatedly. - Prepare 1 L of 2-(N-morpholino)ethane sulfonic acid (MES) running buffer by dissolving 50 mM MES, 50 mM Tris base, 3.47 mM sodium dodecyl sulfate (SDS), and 1.03 mM EDTA in 0.9 L of ddH2O. After mixing properly, adjust the volume to 1 L.
- The pH of the 1x buffer is 7.6. Do not adjust the pH with acid or base.
- Prepare 200 mL of blocking solution (5% milk) by dissolving 10 g milk powder in 1x PBS-T. Store the blocking solution at 4 °C for up to one week.
- Prepare 1 L of transfer buffer (see the Table of Materials) according to the manufacturers' instruction.
- Prepare 2x SDS sample buffer by dissolving 125 mM Tris base, 4% SDS, 4% glycerol, 0.03% bromophenol blue, and 50 µL/mL of β-mercaptoethanol in 50 mL ddH2O. Aliquot in 1 mL portions, and store at -20 °C until use.
2. In vitro auto-ubiquitylation assay
- Assay preparation and execution
- Calculate the volume of each reagent required per reaction based on the given molarities of the proteins and the protein concentrations of the protein solutions. Per auto-ubiquitylation reaction, use 100 nM E1, 1 µM E2, 1 µM E3, and 50 µM Ub. Adjust the volume of the reaction to 20 µL with ddH2O.
NOTE: ATP solution and ubiquitylation buffer were purchased as 10x stock solutions and used at 1x. - Prepare a pipetting scheme for all reactions. Test nine different E2 enzymes for their ability to function (I) with CHIP, (II) with a catalytically inactive CHIP(H260Q) mutant, and (III) without CHIP in individual ubiquitylation reactions.
- To avoid pipetting errors, prepare a master mix on ice that contains the volume required for all reactions plus one extra reaction. Calculate the amount of master mix required per reaction.
NOTE: Master mix components are reagents that are equally required for each reaction, i.e., E1, E3, Ub, ATP, and ubiquitylation buffer. - Add ddH2O and master mix to the polymerase chain reaction (PCR) tubes. Keep the tubes on ice.
- Before starting the ubiquitylation reactions by adding the E2 enzymes, set up a PCR thermal cycler with the following program: 2 h, 37 °C followed by 4 °C, infinitely.
- Add 1 µM of the E2 enzymes in the respective tubes, and incubate the samples in the PCR thermal cycler for the indicated period of time.
- After 2 h, add SDS sample buffer to each reaction, and mix by pipetting up and down several times.
NOTE: The required volume of the sample buffer depends on the stock concentration of the sample buffer. Here, 20 µL of 2x SDS sample buffer were added to each reaction. - Boil the samples immediately at 95 °C for 5 min, and continue with polyacrylamide gel electrophoresis (PAGE). Store the denatured proteins at -20 °C.
- Calculate the volume of each reagent required per reaction based on the given molarities of the proteins and the protein concentrations of the protein solutions. Per auto-ubiquitylation reaction, use 100 nM E1, 1 µM E2, 1 µM E3, and 50 µM Ub. Adjust the volume of the reaction to 20 µL with ddH2O.
- Gel electrophoresis and western blot
- After thawing, spin down the samples for approximately 10 s and load the SDS-PAGE gel. Use a protein ladder as size reference.
NOTE: Here, 4-12% Bis-Tris gradient gels and 3 µL of protein ladder were used. Divide the sample volumes and load two gels equally using 20 µL of each sample. - Run the gel at 160-200 V for approximately 30-45 min so that the sample front reaches the bottom of the gel.
- Transfer each gel to a plastic container filled with semidry blotting buffer, and incubate it for 2-5 min at RT. Remove the stacking gel.
- Prepare two pieces of gel-sized blotting paper and a gel-sized nitrocellulose membrane per SDS gel and pre-soak in semidry blotting buffer. Assemble the western blot (WB) sandwich in a WB chamber from bottom to top in the following order: blotting paper, nitrocellulose membrane, SDS-PAGE gel, blotting paper.
- Remove air bubbles that may have been trapped between the sandwich layers. For this purpose, use a roller to carefully roll over the sandwich a few times.
- Close the chamber, and allow excess blot buffer to drain out.
- Place the chamber in the respective blotting device, and use the following program for semidry blotting: 25 V, 1.0 A, 30 min.
- Transfer the blotted membrane to a plastic container filled with blocking solution, and incubate for 30 min at RT.
- Prepare the primary antibody solution by adding the antibody to 10 mL of PBS-T containing a blocking reagent (see the Table of Materials).
NOTE: The working concentration of the antibody is antibody-specific. In this case, a monoclonal mouse anti-ubiquitin antibody was used at a dilution of 1:5,000. For the second gel, a monoclonal rabbit anti-CHIP antibody was used at 1:5,000 in PBS-T/blocking reagent. - Replace the blocking solution with the primary antibody solution, and incubate overnight at 4 °C on a rocker.
- Wash the membrane three times for 10 min with PBS-T.
- Prepare the secondary antibody solution by adding the respective antibody to 10 mL of PBS-T.
NOTE: Here, goat anti-mouse and mouse anti-rabbit antibodies are used at a dilution of 1:10,000. - Incubate the membrane with the secondary antibody for 1 h at RT on a rocker.
- Wash the membrane three times for 5 min with PBS-T.
- After thawing, spin down the samples for approximately 10 s and load the SDS-PAGE gel. Use a protein ladder as size reference.
- Data analysis
- Add western blot detection reagents for horseradish peroxidase (HRP)-conjugated antibodies to the washed membrane according to the manufacturer's instructions, and capture the HRP signal by using X-ray films or a charge-coupled device camera (Figure 1).
3. In vitro substrate ubiquitylation assay
- Assay preparation and execution
- Calculate the volume of each reagent required per reaction, based on the given molarities of the proteins and the protein concentrations of the protein solutions. Per substrate ubiquitylation reaction, use 100 nM E1, 1 µM E2, 1 µM E3, 1 µM substrate, and 50 µM Ub. Use 10x ATP solution and 10x ubiquitylation buffer at 1x. Adjust the volume of the substrate ubiquitylation reaction to 20 µL with ddH2O.
- Prepare a pipetting scheme for all reactions. Include proper control reactions verifying that the substrate ubiquitylation is E3-specific. Use the protein UNC-45B as a representative substrate of CHIP and a catalytically inactive CHIP mutant (H260Q) as a control. Prepare the following reactions: E1, E2, Ub mix (including Ub, ATP, ubiquitylation buffer); E1, E2, Ub mix, UNC-45B; E1, E2, Ub mix, CHIP(H260Q), UNC-45B; and E1, E2, Ub mix, CHIP, UNC-45B.
- To avoid pipetting errors, prepare a master mix on ice that contains the volume required for all reactions plus one extra reaction. Calculate the volume of the master mix that is required per reaction.
NOTE: Master mix components for this assay include E1, E2, Ub, ATP, and ubiquitylation buffer. - Add reaction components in the following order: ddH2O, substrate, E3 ligase.
- Before starting the ubiquitylation reaction by addition of the master mix, set up a PCR thermal cycler with the following program: 2 h, 37 °C followed by 4 °C, infinitely.
- Add the master mix to each tube, and incubate the samples in the PCR thermal cycler for the indicated period of time.
- After 2 h, add 2x SDS sample buffer to each reaction, and mix by pipetting up and down several times.
- After the run, boil the samples immediately at 95 °C for 5 min, and continue with PAGE. Alternatively, store the denatured proteins at -20 °C.
- Gel electrophoresis and western blot
- Perform gel electrophoresis and western blot as described in section 2.2. Use specific antibodies for the detection of the substrate and the E3 ligase, respectively.
NOTE: Here, UNC-45B is fused to a polypeptide protein tag derived from the c-myc gene product allowing for the use of a monoclonal mouse anti-MYC antibody. Anti-MYC is used at 1:10,000.
- Perform gel electrophoresis and western blot as described in section 2.2. Use specific antibodies for the detection of the substrate and the E3 ligase, respectively.
- Data analysis
- Perform data analysis as described in section 2.3 (Figure 2).
4. Lysine discharge assay
- Assay preparation and execution
- Charging of the E2 enzyme with Ub by E1
- Calculate the volume of each reagent required per reaction based on the given molarities of the proteins and the protein concentrations of the protein solutions. Per charging reaction, use 2 µM E1, 4 µM E2, and 4 µM lysine-free ubiquitin (Ub K0). Use 10x ATP and 10x ubiquitylation buffer at 1x. Adjust the volume of the charging reaction to 20 µL with ddH20.
NOTE: The lysine-free Ub K0 mutant is used to enforce the exclusive production of mono-ubiquitylated E2. Wild-type Ub can also be used; however, this might lead to diverse E2~Ub modifications (e.g., poly-ubiquitylation of E2), which are more difficult to analyze. For first time use or when using a new E2 enzyme, determine the level of Ub charging on E2 that can be achieved by E1. To determine the yield of charging, visualize uncharged vs. charged E2 via Coomassie staining. Here, approximately half of Ub K0 was reliably converted to UBE2D2~Ub when using equimolar ratios of UBE2D2 and Ub K0 (Supplemental Figure S2A). - Prepare a pipetting scheme for the charging reaction. Adjust the volume of the charging reaction to the subsequent discharge reactions of choice, e.g., use CHIP and CHIP(H260Q) in individual discharge reactions. Thus, prepare double the volume of one charging reaction.
NOTE: For first time use, test different concentrations of the E3 ligase of choice to monitor optimal discharge conditions, and analyze them by western blotting. In an ideal condition, the discharge of Ub from E2 will increase over time until the respective E2~Ub protein band disappears. - Incubate the charging reaction for 15 min at 37 °C.
- Calculate the volume of each reagent required per reaction based on the given molarities of the proteins and the protein concentrations of the protein solutions. Per charging reaction, use 2 µM E1, 4 µM E2, and 4 µM lysine-free ubiquitin (Ub K0). Use 10x ATP and 10x ubiquitylation buffer at 1x. Adjust the volume of the charging reaction to 20 µL with ddH20.
- Termination of the charging reaction
- Stop the charging reaction by addition of apyrase at a final concentration of 1.8 U/mL. Carry out the incubation with apyrase for 5 min at RT, followed by addition of EDTA at a final concentration of 30 mM. Adjust the volume of the charging reaction to 30 µL with ddH2O.
NOTE: Apyrase is used to convert ATP molecules to ADP (adenosine diphosphate) molecules. As the activity of the E1 enzyme is ATP-dependent, E1 can no longer charge E2. EDTA is additionally used to ensure that E1 is inhibited by quenching Mg2+ ions that are necessary co-factors for E1. The volume of apyrase that is required to stop the reaction can vary among assay conditions. Although apyrase alone already quenches available ATP molecules, a combination of apyrase and EDTA was more effective at stopping the charging reaction for the E1-UBE2D2 pair. As stopping the reaction is a critical factor for assay reproducibility, efficient stopping should be tested for new E1-E2 pairs. For this purpose, set up a charging reaction, stop it, and collect samples at specific time points, e.g., after 2, 5, 10, and 15 min. Non-changing levels of E2~Ub indicate that the reaction has stopped, and that the thioester was stable during that period of time (Supplemental Figure S2B).
- Stop the charging reaction by addition of apyrase at a final concentration of 1.8 U/mL. Carry out the incubation with apyrase for 5 min at RT, followed by addition of EDTA at a final concentration of 30 mM. Adjust the volume of the charging reaction to 30 µL with ddH2O.
- Discharging of E2~Ub by E3
- Prepare four tubes corresponding to the different time points (t0, t1, t2, t3); add non-reducing sample buffer (6.7 µL of 4x lithium dodecyl sulfate (LDS) buffer) to each tube.
NOTE: Do not use reducing agent in the sample buffer. - Remove 6 µL from the stopped charging reaction for t0, and adjust the volume to 20 µL with ddH2O.
- Incubate the t0 sample for 10 min at 70 °C.
NOTE: Do not boil the sample by extending the incubation time as thioesters are labile at higher temperatures. - Calculate the volume of each reagent required per discharge reaction. Use the remaining 24 µL of the charged E2, and add 10 mM L-lysine, 1 mg/ml BSA, and 500 nM of the E3 ligase. Use the ubiquitylation buffer at 1x, and adjust the volume to 80 µL with ddH2O.
- Prepare a pipetting scheme for all discharge reactions, and use CHIP(H260Q) as control in addition to CHIP(WT).
- Set up discharge reactions on ice by adding the required components in the following order: ddH2O, ubiquitylation buffer, BSA, lysine, E3 ligase, and the charged E2 to start the reaction.
- Incubate the discharge reaction at RT.
- Take samples after 5, 30, and 60 min by transferring 20 µL of the discharge reaction into the respective sample tubes.
NOTE: Suitable time points that allow for proper monitoring of the discharge can vary between E2-E3 pairs. - Vortex the sample immediately, and incubate it at 70 °C for 10 min.
- Directly proceed with PAGE.
NOTE: E2~Ub thioesters can be labile even after denaturing in sample buffer. Thus, gel electrophoresis should be performed directly after assay execution.
- Prepare four tubes corresponding to the different time points (t0, t1, t2, t3); add non-reducing sample buffer (6.7 µL of 4x lithium dodecyl sulfate (LDS) buffer) to each tube.
- Charging of the E2 enzyme with Ub by E1
- Gel electrophoresis and western blot
- Perform the gel electrophoresis and the western blotting as described in section 2.2. Use a Ub-specific antibody and, if available, also use an E3-specific antibody that was raised in a different species, allowing for simultaneous detection of the Ub and E3 ligase signal.
- Data analysis
- Perform data analysis as described in section 2.3 (Figure 3).
Representative Results
To identify E2 enzymes that cooperate with the ubiquitin ligase CHIP, a set of E2 candidates was tested in individual in vitro ubiquitylation reactions. Cooperating E2-E3 pairs were monitored by the formation of E3-dependent ubiquitylation products, i.e., auto-ubiquitylation of the E3 ligase and the formation of free Ub polymers. The ubiquitylation products were analyzed by western blotting. Data interpretation is based on the size comparison of the resulting protein bands with molecular weight markers. Protein ubiquitylation leads to the formation of specific band patterns characterized by the appearance of double bands or multiple iterative bands with a respective size difference of 8.6 kDa (size of a single Ub molecule).
Here, the ability of nine E2s to promote the formation of ubiquitylation products was tested in the presence of wild-type CHIP (Figure 1A), the inactive U-box mutant CHIP(H260Q) (Figure 1B), or without CHIP (Supplemental Figure S1). E3-independent ubiquitin products were formed in the presence of inactive CHIP and in the absence of CHIP (Figure 1B and Supplemental Figure S1). Inactive CHIP was not auto-ubiquitylated (Figure 1B). In contrast, wild-type CHIP was auto-ubiquitylated when combined with members of the UBE2D family (D1-D3) and members of the UBE2E family (E1, E3), respectively (Figure 1A, lanes 3, 4, and 5). Whereas free poly-Ub chains were produced in cooperation with UBE2D1-3, this was not detected for UBE2E1 or UBE2E3, respectively (Figure 1A, lanes 6 and 7).
The ability of the UBE2D family to promote both the formation of free Ub polymers and the auto-ubiquitylation of CHIP has been attributed to the presence of a non-covalent ubiquitin binding site on the backside of the E227,28. Similarly, the exclusive formation of free Ub chains by UBE2N/V1 (Figure 1A, lane 9) has been attributed to the binding of Ub by a specific UBE2V1 subunit (Uev subunit), directing the formation of K63-linked Ub chains29. No ubiquitylation products were formed in the presence of UBE2C1 and UBE2H (Figure 1A, lanes 2 and 8). In conclusion, CHIP can collaborate with several E2 enzymes in vitro; however, its auto-ubiquitylation is E2 enzyme-specific.
Next, the UBE2D2-CHIP pair was used to investigate the ubiquitylation of the myosin-directed chaperone, UNC-45B, by CHIP ligase activity (Figure 2). Substrate ubiquitylation was analyzed via western blotting. Non-ubiquitylated UNC-45B has a molecular weight of 103 kDa (Figure 2, lane 4). The inactive U-box mutant of CHIP performed neither ubiquitylation of UNC-45B nor auto-ubiquitylation (Figure 2, lane 3). In contrast, ubiquitylation of UNC-45B and auto-ubiquitylation of CHIP was detected upon incubation with wild-type CHIP (Figure 2, lane 6).Thus, CHIP can ubiquitylate UNC-45B in vitro, suggesting that UNC-45B is a conserved substrate of CHIP18.
Ultimately, the catalytic activity of CHIP was analyzed in the absence of any substrate protein. To this end, a lysine discharge assay was performed in which free lysine amino acids can serve as Ub acceptor in the absence of E3 substrates (Figure 3, Supplemental Figure S2C). The discharge assay consists of a charging step in which Ub is loaded onto the E2 by E1, a stop step to prevent further loading of Ub onto E2, and a discharge step where Ub is transferred from the transient Ub~E2 thioester onto free lysine amino acids and/or onto lysine residues of the E3 ligase. Western blot analysis was performed to visualize both the Ub-modified proteins and the E3 ligase CHIP. The uncharged UBE2D2 enzyme has a molecular weight of 17 kDa (Supplemental Figure S2C).
When charged with a single Ub molecule-ensured by the use of lysine-free Ub (Ub K0)-the molecular weight of UBE2D2~Ub shifts upwards to approximately 26 kDa. The time point zero (t0) represents the overall yield of charged E2 (Figure 3). In the presence of inactive CHIP (35 kDa), a faint E3 ligase-independent discharge of UBE2D230,31, but no auto-ubiquitylation of CHIP, was detected. In contrast, in the presence of wild-type CHIP (35 kDa), a faster discharge of UBE2D2 was detected, yielding a complete discharge within 60 min. Simultaneously, auto-ubiquitylation of CHIP was observed, indicating that CHIP promoted the transfer of Ub onto its own lysine residues.
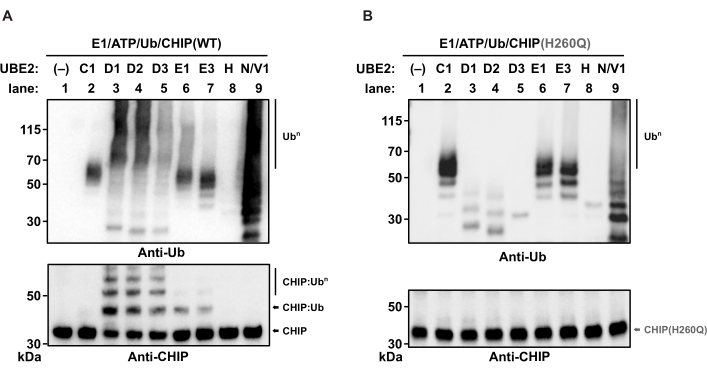
Figure 1: Western blot analysis for E2-E3 enzyme collaboration in vitro. In vitro auto-ubiquitylation reactions with different human E2 enzymes (from left to right: empty, UBE2-D1, -D2, -D3, -E1, -E3, -H, and -N/V1) were performed as indicated using (A) wild-type CHIP or (B) the inactive CHIP U-box mutant, CHIP(H260Q), as E3 ubiquitin ligase. The samples were divided equally, run on separate polyacrylamide gels, and immunoblotted with anti-CHIP and anti-Ub antibodies to visualize reaction products. Abbreviations: Ub = ubiquitin; CHIP = carboxyl terminus of HSC70-interacting protein; ATP = adenosine triphosphate. Please click here to view a larger version of this figure.
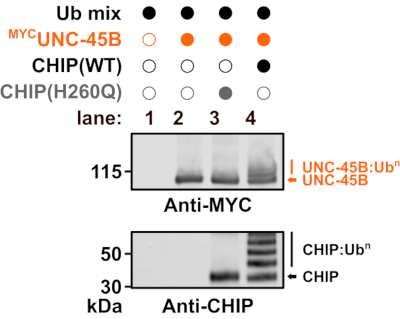
Figure 2: Western blot analysis monitoring substrate ubiquitylation. In vitro substrate ubiquitylation reactions were performed as indicated to detect ubiquitylation of human UNC-45B by CHIP wild-type or the inactive CHIP U-box mutant, CHIP(H260Q). Human UBE2D2 was used as E2 enzyme. Abbreviations: WT = wild-type; Ub = ubiquitin; CHIP = carboxyl terminus of HSC70-interacting protein. Please click here to view a larger version of this figure.
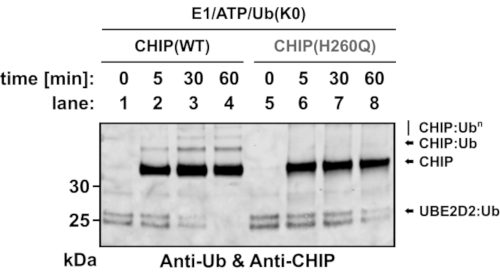
Figure 3: Western blot analysis monitoring the CHIP-dependent Ub transfer from UBE2D2~Ub thioester. Charging of human UBE2D2 by lysine-free Ub (Ub K0), stopping of the charging reaction, and discharging of UBE2D2~Ub was performed as described. For the discharge reactions, wild-type CHIP or the inactive CHIP U-box mutant, CHIP(H260Q), were used as ubiquitin ligases and 10 mM L-lysine was supplied as the potential Ub acceptor. Samples were collected after the indicated time periods, run on a polyacrylamide gel, and immunoblotted with an anti-Ub/anti-CHIP antibody mixture. Abbreviations: Ub = ubiquitin; CHIP = carboxyl terminus of HSC70-interacting protein; ATP = adenosine triphosphate. Please click here to view a larger version of this figure.
Supplemental Figure S1: Representative western blot monitoring E2 enzyme activities in the absence of E3. In vitro ubiquitylation reactions with different human E2 enzymes (from left to right: empty, UBE2-D1, -D2, -D3, -E1, -E3, -H, and -N/V1) were performed as indicated in the absence of an E3 ligase to screen E3-independent reaction products. The samples were run on a polyacrylamide gel and immunoblotted with anti-Ub. Abbreviations: Ub = ubiquitin; ATP = adenosine triphosphate. Please click here to download this File.
Supplemental Figure S2: Analysis of the charging yield and stability of the UBE2D2~Ub K0 thioester. (A) The yield of UBE2D2 charging achieved by E1 was tested under various conditions. The first charging reaction (I) consisted of 5 µM E1, 5 µM E2, and 50 µM Ub K0. The second charging reaction (II) consisted of 2 µM E1, 4 µM E2, and 4 µM Ub K0. The charging reactions were performed as described for 30 min at 37 °C. Samples were collected after 15 and 30 min. The reactions were stopped by addition of 4x LDS sample buffer and incubated at 70 °C for 10 min prior to gel electrophoresis and Coomassie staining. Uncharged UBE2D2 was used as control. Approximately half of the UBE2D2 was converted to UBE2D2~Ub. No additional charging was detected after 15 min. Moreover, neither the increase in E1 nor the increase in free ubiquitin altered the charging yield of UBE2D2~Ub. Thus, charging was subsequently performed for 15 min at 37 °C by using 2 µM E1, 4 µM E2, and 4 µM Ub K0. (B) After charging, the reaction was stopped by addition of 1.8 U/mL apyrase and 30 mM EDTA and incubated at RT. Samples were collected after 2, 5, 8, 10, and 15 min (lanes 2-6), and uncharged UBE2D2 was used as control (lane 1). 4x LDS sample buffer was added, and samples were incubated at 70 °C for 10 min prior to gel electrophoresis and Coomassie staining. Stopping was efficient as measured by the constant band intensity of the UBE2D2~Ub protein, also indicating that the thioester was stable during the indicated time period. (C) Charging of human UBE2D2 by lysine-free Ub (Ub K0), stopping of the charging reaction, and discharging of UBE2D2~Ub was performed as described. For the discharge reactions, wild-type CHIP or the inactive CHIP U-box mutant, CHIP(H260Q) were used as ubiquitin ligases, and 10 mM L-lysine was supplied as the potential Ub acceptor. Samples were collected after the indicated time periods, run on separate polyacrylamide gels, and Coomassie-stained. Please click here download this File.
Discussion
This paper describes basic in vitro ubiquitylation methods for the analysis of E3 ligase function. When performing in vitro ubiquitylation assays, it should be considered that some E2 enzymes can perform auto-ubiquitylation owing to the attack of their active cysteine on their own lysine residues that are located in close proximity to the active site30. To circumvent this problem, the use of an E2 mutant is recommended in which the respective lysine residue is exchanged for arginine, which results in a catalytically active E2 enzyme that is resistant to auto-ubiquitylation32. This is of particular importance when monitoring the Ub transfer via lysine discharge assays to ensure reproducibility of the assay. Another critical factor is the transient nature of the E2~Ub thioester bond. In case the thioester stability limits possible in vitro applications, the active cysteine residue of the E2 could be exchanged for a serine to generate a more stable E2~Ub oxyester33. For first time use or when using a new E2 enzyme, determine the level of Ub charging on E2 that can be achieved by E1. The charging efficiency can be affected by several factors including the species origin of E1, the temperature of the charging reaction, and the molar ratio of Ub to E2. To determine the yield of charging, visualize uncharged vs. charged E2 via Coomassie staining.
Altogether, these potential restrictions highlight the importance of the empirical optimization of in vitro assay conditions, such as E2-E3 pair, E2 charging and discharging kinetics, molarity and activity of the recombinant proteins, especially for the lysine discharge assay, to obtain reproducible results. Given the diverse cellular functions of the covalent attachment of Ub to substrate proteins, the analysis of protein ubiquitylation is a popular research field. However, the analysis of ubiquitylation events can be difficult, especially in vivo. This difficulty arises from multiple factors including the transient nature of the E3-substrate interaction34, redundancy of many E3 ligases, as well as promiscuity of E3 ligases for several substrates35. Furthermore, the characterization of specific ubiquitylation events in vivo is hindered by the existence of additional contributing factors such as ubiquitin chain elongation factors and deubiquitylation enzymes that modulate ubiquitin chain topology36. These constraints underline the importance of robust in vitro techniques that help to characterize enzymatic activities of E3 ubiquitin ligases in a defined recombinant system.
Apart from the described applications, in vitro ubiquitylation assays can also be used to detect the Ub-linkage type by using linkage-type specific antibodies. As an additional more detailed approach, the respective protein band of the Ub-modified substrate can be extracted from the polyacrylamide gel and analyzed via mass spectrometry. The identification of the linkage type supports the understanding of the physiological role of the E3-substrate interaction, which is of particular importance for the characterization of disease-associated substrate degradation pathways37,38. Similarly, the lysine discharge assay can be used as an effective tool to unravel catalytic differences between E3 ubiquitin ligases or E3 ligase families30. In conclusion, the in vitro ubiquitylation methods described herein are effective tools to analyze different aspects of E3 ligase function. Besides the relatively simple execution of the described methods, a major advantage is the general applicability as proteins from all eukaryotic sources could be recombinantly expressed and studied in vitro without the need for special equipment.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank the members of our laboratory for critical discussion and helpful advice on the manuscript. We apologize for not having cited valuable contributions due to size limitation. This work is supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – SFB 1218 – Projektnumber 269925409 and Cluster of Excellence EXC 229/ CECAD to TH. This work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany´s Excellence Strategy – EXC 2030 – 390661388 and – SFB 1218 – Projektnumber 269925409 to T.H. Diese Arbeit wurde von der Deutschen Forschungsgemeinschaft (DFG) im Rahmen der deutschen Exzellenzstrategie – EXC 2030 – 390661388 und – SFB 1218 – Projektnummer: 269925409 an T.H. gefördert.
Materials
| Amershan Protran 0.1 µm NC | GE Healthcare | 10600000 | nitrocellulose membrane |
| Anti-CHIP | Cell Signaling | 2080 | Monoclonal rabbit anti-CHIP antibody, clone C3B6 |
| Anti-MYC | Roche | OP10 | Monoclonal mouse anti-MYC antibody, clone 9E10 |
| Anti-ubiquitin | Upstate | 05-944 | Monoclonal mouse anti-Ub antibody, clone P4D1-A11 |
| Apyrase | Sigma | A6535-100UN | |
| ATP (10x) | Enzo | 12091903 | |
| BSA | Sigma | A6003-10G | |
| EDTA | Roth | 8043.2 | |
| KCl | Roth | 6781.1 | |
| K2HPO4 | Roth | P749.2 | |
| KH2PO4 | Roth | 3904.1 | |
| LDS sample buffer (4x) | novex | B0007 | |
| L-Lysine | Sigma | L5501-5G | |
| MES | Roth | 4256.4 | |
| MeOH | VWR Chemicals | 2,08,47,307 | 100% |
| Milchpulver | Roth | T145.3 | |
| NaCl | Roth | P029.3 | |
| NuPAGE Antioxidant | invitrogen | NP0005 | |
| NuPAGE Transfer buffer (20x) | novex | NP0006-1 | |
| Page ruler plus | Thermo Fisher | 26619 | Protein ladder |
| RotiBlock | Roth | A151.1 | Blocking reagent |
| SDS (20%) | Roth | 1057.1 | |
| S1000 Thermal Cycler | Bio Rad | 1852196 | |
| Trans-Blot Turbo | Bio Rad | 1704150EDU | Transfer system |
| Tris base | Roth | 4855.3 | |
| Tween 20 | Roth | 9127.2 | |
| UbcH Enzyme Set | BostonBiochem | K-980B | E2 enzymes |
| Ubiquitin | BostonBiochem | U-100H | |
| Ubiquitin-activating enzyme E1 | Enzo | BML-UW941U-0050 | |
| Ubiquitylation buffer (10x) | Enzo | BML-KW9885-001 | |
| Whatman blotting paper | Bio Rad | 1703969 | Extra thick filter paper |
References
- Kuhlbrodt, K., Mouysset, J., Hoppe, T. Orchestra for assembly and fate of polyubiquitin chains. Essays in Biochemistry. 41, 1-14 (2005).
- Pickart, C. M., Eddins, M. J. Ubiquitin: structures, functions, mechanisms. Biochimica et Biophysica Acta. 1695 (1-3), 55-72 (2004).
- Dye, B. T., Schulman, B. A. Structural mechanisms underlying posttranslational modification by ubiquitin-like proteins. Annual Review of Biophysics and Biomolecular Structure. 36, 131-150 (2007).
- Koegl, M., et al. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell. 96 (5), 635-644 (1999).
- Hoppe, T. Multiubiquitylation by E4 enzymes: ‘one size’ doesn’t fit all. Trends in Biochemical Sciences. 30 (4), 183-187 (2005).
- Stewart, M. D., Ritterhoff, T., Klevit, R. E., Brzovic, P. S. E2 enzymes: more than just the middle men. Cell Research. 26 (4), 423-440 (2016).
- Clague, M. J., Heride, C., Urbé, S. The demographics of the ubiquitin system. Trends in Cell Biology. 25 (7), 417-426 (2015).
- Buetow, L., Huang, D. T. Structural insights into the catalysis and regulation of E3 ubiquitin ligases. Nature Reviews Molecular Cell Biology. 17 (10), 626-642 (2016).
- French, M. E., Koehler, C. F., Hunter, T. Emerging functions of branched ubiquitin chains. Cell Discovery. 7, 6 (2021).
- Hatakeyama, S., Yada, M., Matsumoto, M., Ishida, N., Nakayama, K. I. U box proteins as a new family of ubiquitin-protein ligases. Journal of Biological Chemistry. 276 (35), 33111-33120 (2001).
- Herhaus, L., Dikic, I. Expanding the ubiquitin code through post-translational modification. EMBO Reports. 16 (9), 1071-1083 (2015).
- Deshaies, R. J., Joazeiro, C. A. RING domain E3 ubiquitin ligases. Annual Review of Biochemistry. 78, 399-434 (2009).
- Hochstrasser, M. Lingering mysteries of ubiquitin-chain assembly. Cell. 124 (1), 27-34 (2006).
- Hoppe, T., Cohen, E. Organismal protein homeostasis mechanisms. Genetics. 215 (4), 889-901 (2020).
- Okiyoneda, T., et al. Peripheral protein quality control removes unfolded CFTR from the plasma membrane. Science. 329 (5993), 805-810 (1997).
- Tawo, R., et al. The Ubiquitin ligase CHIP integrates proteostasis and aging by regulation of insulin receptor turnover. Cell. 169 (3), 470-482 (2017).
- Albert, M. C., et al. CHIP ubiquitylates NOXA and induces its lysosomal degradation in response to DNA damage. Cell Death and Disease. 11 (9), 740 (2020).
- Hoppe, T., et al. Regulation of the myosin-directed chaperone UNC-45 by a novel E3/E4-multiubiquitylation complex in C. elegans. Cell. 118 (3), 337-349 (2004).
- Kim, J., Löwe, T., Hoppe, T. Protein quality control gets muscle into shape. Trends in Cell Biology. 18 (6), 264-272 (2008).
- Janiesch, P. C., et al. The ubiquitin-selective chaperone CDC-48/p97 links myosin assembly to human myopathy. Nature Cell Biology. 9 (4), 379-390 (2007).
- Donkervoort, S., et al. Pathogenic variants in the myosin chaperone UNC-45B cause progressive myopathy with eccentric cores. The American Journal of Human Genetics. 107 (6), 1078-1095 (2020).
- Murata, S., Minami, Y., Minami, M., Chiba, T., Tanaka, K. CHIP is a chaperone-dependent E3 ligase that ubiquitylates unfolded protein. EMBO Reports. 2 (12), 1133-1138 (2001).
- Jiang, J., et al. CHIP is a U-box-dependent E3 ubiquitin ligase: identification of Hsc70 as a target for ubiquitylation. Journal of Biological Chemistry. 276 (64), 42938-42944 (2001).
- Yang, Y., Yu, X. Regulation of apoptosis: the ubiquitous way. The FASEB Journal. 17 (8), 790-799 (2003).
- Lamothe, B., et al. TRAF6 ubiquitin ligase is essential for RANKL signaling and osteoclast differentiation. Biochemical and Biophysical Research Communication. 359 (4), 1044-1049 (2007).
- Amemiya, Y., Azmi, P., Seth, A. Autoubiquitination of BCA2 RING E3 ligase regulates its own stability and affects cell migration. Molecular Cancer Research. 6 (9), 1385 (2008).
- Brzovic, P. S., Lissounov, A., Christensen, D. E., Hoyt, D. W., Klevit, R. E. A UbcH5/ubiquitin noncovalent complex is required for processive BRCA1-directed ubiquitination. Molecular Cell. 21 (6), 873-880 (2006).
- Sakata, E., et al. Crystal structure of UbcH5b~ubiquitin intermediate: Insight into the formation of the self-assembled E2~Ub conjugates. Structure. 18 (1), 138-147 (2010).
- Eddins, M. J., Carlile, C. M., Gomez, K. M., Pickart, C. M., Wolberger, C. Mms2-Ubc13 covalently bound to ubiquitin reveals the structural basis of linkage-specific polyubiquitin chain formation. Nature Structural and Molecular Biology. 13 (10), 915-920 (2006).
- Buetow, L., et al. Activation of a primed RING E3-E2 ubiquitin complex by non-covalent ubiquitin. Molecular Cell. 58 (2), 297-310 (2015).
- Dou, H., Buetow, L., Sibbet, G. J., Cameron, K., Huang, D. T. BIRC7-E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nature Structural and Molecular Biology. 19 (9), 876-883 (2012).
- McKenna, S., et al. Noncovalent interaction between ubiquitin and the human DNA repair protein Mms2 is required for Ubc13-mediated polyubiquitination. Journal of Biological Chemistry. 276 (43), 40120-40125 (2001).
- Plechanovová, A., et al. Mechanism of ubiquitylation by dimeric RING ligase RNF4. Nature Structural Biology. 18 (9), 1052-1059 (2011).
- Pierce, N. W., Kleiger, G., Shan, S. O., Deshaies, R. J. Detection of sequential polyubiquitylation on a millisecond timescale. Nature. 462 (7273), 615-619 (2009).
- Jain, A. K., Barton, M. C. Regulation of p53: TRIM24 enters the RING. Cell Cycle. 8 (22), 3668-3674 (2009).
- Swatek, K. N., Komander, D. Ubiquitin modifications. Cell Research. 26, 399-422 (2016).
- Yan, K., et al. The role of K63-linked polyubiquitination in cardiac hypertrophy. Journal of Cellular and Molecular Medicine. 22 (10), 4558-4567 (2018).
- Dammer, E. B., et al. Polyubiquitin linkage profiles in three models of proteolytic stress suggest the etiology of Alzheimer disease. Journal of Biological Chemistry. 286 (12), 10457-10465 (2011).

