Production of IgG Fusion Proteins Transiently Expressed in Nicotiana benthamiana
Summary
We describe here a simple method for expression, extraction, and purification of recombinant human IgG fused to GFP in Nicotiana benthamiana. This protocol can be extended to purification and visualization of numerous proteins that utilize column chromatography. Moreover, the protocol is adaptable to the in-person and virtual college teaching laboratory, providing project-based exploration.
Abstract
High demand for antibodies as therapeutic interventions for various infectious, metabolic, autoimmune, neoplastic, and other diseases creates a growing need in developing efficient methods for recombinant antibody production. As of 2019, there were more than 70 FDA-approved monoclonal antibodies, and there is exponential growth potential. Despite their promise, limiting factors for widespread use are manufacturing costs and complexity. Potentially, plants offer low-cost, safe, and easily scalable protein manufacturing strategies. Plants like Nicotiana benthamiana not only can correctly fold and assemble complex mammalian proteins but also can add critical post-translational modifications similar to those offered by mammalian cell cultures. In this work, by using native GFP and an acid-stable variant of green fluorescent protein (GFP) fused to human monoclonal antibodies, we were able to visualize the entire transient antibody expression and purification process from N. benthamiana plants. Depending on the experiment's purpose, native GFP fusion can ensure easier visualization during the expression phase in the plants, while acid-stable GFP fusion allows for visualization during downstream processing. This scalable and straightforward procedure can be performed by a single researcher to produce milligram quantities of highly pure antibody or antibody fusion proteins in a matter of days using only a few small plants. Such a technique can be extended to the visualization of any type of antibody purification process and potentially many other proteins, both in plant and other expression systems. Moreover, these techniques can benefit virtual instructions and be executed in a teaching laboratory by undergraduate students possessing minimal prior experience with molecular biology techniques, providing a foundation for project-based exploration with real-world applications.
Introduction
Industry reports indicate that thirteen out of the twenty most-highly grossing drugs in the United States were biologics (protein-based pharmaceuticals), of which nine were antibodies. As of 2019, there were over 570 antibody (Ab) therapeutics at various clinical development phases1,2,3. Current global Ab sales exceed 100 billion USD, and the monoclonal Ab (mAb) therapeutic market is expected to generate up to 300 billion USD by 20251,4. With such high demand and projected increases in revenue, researchers have been working to develop ways to produce Ab therapeutics on an ever-larger scale, with higher quality and lower-costs. Plant-based expression systems have several advantages over traditional mammalian cell lines for the affordable and large-scale manufacture of Ab therapeutics5,6. Production of protein therapeutics in plants ("molecular pharming") does not require expensive bioreactors or cell culture facilities as do traditional mammalian cell culture techniques7,8. Plants cannot contract human pathogens, minimizing potential contamination9. Both transient and transgenic plant-based protein expression can be utilized as lower-cost alternatives to mammalian or bacterial production systems10. Though transgenic plants are preferred for crop production, recombinant protein production using this method can require weeks to months. Advances in transient expression using viral vectors through either syringe or vacuum agroinfiltration allow for small- and large-scale production, respectively, of the desired protein in days11,12,13,14. Production of mAbs against Ebola, Dengue and, Zika, and numerous other recombinant proteins, have been produced and purified quickly and efficiently using transient expression in N. benthamiana plants15,16,17,18,19. These circumstances make transient plant-based expression an attractive option for developing multiple Ab therapeutics and the methods demonstrated in this protocol20.
First-generation mAbs were of murine derivation, which resulted in non-specific immunogenicity when used in human trials21. Over time, chimeric, humanized, and eventually, fully human Abs were produced to lessen immunogenicity induced by Ab therapeutics. Unfortunately, some of these Abs still cause host immunogenicity due to differences in glycosylation21. Developments in plant engineering have allowed for the modification of Ab glycans, which is essential since an Ab's stability and function can significantly be affected by its glycosylation state22. Advances have allowed production in plant systems of high-level expression of humanized mAbs, containing human glycans and resultantly the desired biological traits of a mass-produced human pharmaceutical19,21.
In addition to recombinant Abs, Ab fusion molecules (Ab fusions) have been explored for various purposes in recent decades. Ab fusions often consist of an Ab or Ab fragment fused to a molecule or protein and are designed to elicit responses from immune effector cells23. These molecules have been created as potential therapeutic interventions to treat various pathologies such as cancer and autoimmune diseases24,25,26,27. Recombinant immune complexes (RICs) are another class of Ab fusions that have been employed as vaccine candidates28. RICs take advantage of the immune system's ability to recognize Fc regions of Ab fusions and have been found to improve immunogenicity when combined with other vaccine platforms29,30,31.
Green Fluorescent Protein (GFP) is a bioluminescent protein derived from the jellyfish Aequorea Victoria, which emits green light when excited by ultraviolet light32,33. Over the years, GFP's use as a visual marker of gene expression has expanded from expression in Escherichia coli to numerous protein expression systems, including N. benthamiana plants34,35,36,37,38. Visible markers, such as GFP, have abundant implications in the teaching and learning of scientific concepts. Numerous entry-level students describe difficulties grasping scientific concepts when the idea being taught is not visible to the naked eye, such as the concepts of molecular biology and related fields39. Visual markers, like GFP, can thus contribute to the processing of information related to the scientific processes and could help lessen the difficulties students report in learning numerous scientific concepts.
Although GFP is often used as a marker to indicate gene and expression in vivo, it is difficult to visualize it in the downstream processes if using acidic conditions. This circumstance is primarily because GFP does not maintain its structure and resultant fluorescence at a low pH40. Temporary acidic environments are often required in various purification processes, such as protein G, protein A, and protein L chromatography, often utilized for Ab purification41,42,43,44. GFP mutants have been used to retain fluorescence under acidic conditions45,46.
Herein we describe a simple method for expression, extraction, and purification of recombinant IgG fusion proteins in N. benthamiana plants. We produced traditional GFP fused to the N-terminus of a humanized IgG heavy chain, creating a GFP-IgG fusion. Simultaneously, we developed the fusion of a plant codon-optimized sequence for an acid-stable GFP (asGFP) to the N-terminus of a humanized IgG heavy chain, creating an asGFP-IgG fusion. The advantages of producing GFP-IgG include the ability to visualize the presence of a target protein during expression, while asGFP-IgG allows seeing the presence of recombinant protein in not only the expression and extraction steps but also in the purification steps of the protein. This protocol can be adapted for the production, purification, and visualization of a range of GFP fusion proteins produced in N. benthamiana and purified using chromatography techniques that require low pH. The process can also be tailored to various amounts of leaf material. While Abs and fusion proteins tagged with GFP or asGFP are not intended to be used for therapies, these methods can be useful as controls during experiments and can also be further utilized as a teaching tool for molecular and cellular biology and biotechnology, both in-person and virtually.
Protocol
1. Cultivate N. benthamiana plants
- Place soil peat pellets on a tray and pour previously boiled, still hot (~40-45 °C), water over the peat pellets for full expansion. After pellets are fully expanded, place 2-3 N. benthamiana seeds on each peat pellet using tweezers.
- Pour about 0.5 in of water to cover the bottom of the tray. Label the tray with the seeding date. Continue to water the seedlings daily with appropriate amounts of fertilizer. Fertilizer (water-soluble all-purpose plant food) concentration is generally 2.5-2.8 g/L.
- Cover the tray with a humidome top when placed in the growth chamber. Keep the seeded peat pellets in the growth chamber at 23-25 °C, with a 16 h photoperiod and 60% relative humidity.
- After one week, remove extra plants leaving each pellet with only one seedling.
- When the plants are 2-3 weeks old, transfer each peat pellet to an individual pot containing moisture control soil. This demonstration used Miracle-Gro moisture control potting soil.
- Water daily with 1 g/L fertilizer. Never leave the soil completely dry. Plants are ready for infiltration when they are 5-6 weeks old.
2. Preparation of Agrobacterium tumefaciens for infiltration
NOTE: GFP-IgG fusion constructs can be obtained as described in this paper31. The asGFP gene was obtained and plant-optimized from this study45. The following steps must be done next to a Bunsen burner, and basic aseptic techniques should be applied to avoid contamination.
- Streak A. tumefaciens EHA105 harboring bean yellow dwarf virus (BeYDV)19 plant expression vector for each construct (asGFP-IgG, GFP-IgG, light chain) from a glycerol stock on LB agar (10 g/L Tryptone, 10 g/L NaCl, 5 g yeast extract, 15 g/L agar, 50 µg/mL kanamycin) plate.
- Grow for one day in a 30 °C standing incubator. Isolate a single colony for verification by standard colony screen PCR protocol.
- Use verified colony for each construct. Fill conical tube with 10 mL of LB media (10 g/L tryptone, 10 g/L NaCl, 5 g of yeast extract, 50 µg/mL). Next, add 10 µL of 100 µg/mL kanamycin. Add 10 µL of 2.5 µg/mL rifampicin to prevent E. coli contamination. Incubate at 30°C and 120-150 rpm overnight.
- The next day, if the Agrobacterium culture is grown to OD600 = 0.6-0.9, it can be used for infiltration. If it is overgrown (OD600 > 1), 1-2 mL should be transferred to fresh LB with antibiotics and grown to the required OD600. Depending on the initial culture's concentration, it may potentially take two days to grow to OD600 = 0.6-0.9.
- Once at appropriate OD600, place the cultures in a centrifuge, and pellet the bacteria by centrifugation at 4,500 x g for 20 min, room temperature (RT).
- Decant supernatant from both samples, and then resuspend each pellet in 1x infiltration buffer (10 mM 2-(N-morpholino) ethanesulfonic acid, 10 mM magnesium sulfate, adjusted to pH 5.5 with KOH) to get final OD600 = 0.4. This should take approximately 15-45 mL of infiltration buffer, depending on the initial culture density. Combine equal volumes of each IgG fusion construct with the light chain construct to get final OD600 = 0.2 per construct in each tube.
3. Needle-less syringe agroinfiltration
- Take a straightened paper clip and 5-6-week-old N. benthamiana plants from step 1. Using the paper clip's sharp edge, make a small puncture in the first epidermal layer of the leaf on the adaxial surface. Avoid puncturing it all the way through.
NOTE: The lower leaves are easier for infiltration, whereas the leaves on the top of the plant are harder. Generally, the expression of recombinant proteins is highest in the leaves located in the middle of a plant, and these leaves also get less necrotic. - Fill a 1 mL syringe, without a needle attached, with the prepared Agrobacterium solution from step 2. Cover the hole made in the previous step with the end of the syringe and slowly push to inject the bacteria into the leaf while applying gentle counterpressure from behind the leaf. Watch the leaf darken as the solution is injected without applying too much pressure on the syringe.
- Try to infiltrate most of the leaf area at a maximum of 3-4 times – excessive leaf damage may hinder protein yield. The infiltrated plant leaf will appear mostly dark from the bottom view.
NOTE: This bacterial solution should be enough for at least 3-4 plants per construct. Autoclave any remaining bacterial solution before discarding.
4. Grow and observe the infiltrated N. benthamiana
- Place infiltrated plants back in the growth chamber and continue to water daily.
- Observe the leaves for chlorosis and necrosis in infiltrated areas. Observe plants for GFP fluorescence (if GFP is present) under a long and short-wave UV lamp.
- Day 4-5 shows the highest fluorescence of both GFP constructs in the leaves. Harvest all the leaves at 4-5 dpi (days post-infiltration) and weigh the total leaf material.
- Use it immediately for downstream processing or store at -80 °C until ready to use.
5. Protein extraction
- Keep buffers and blender cups on ice or at 4 °C before use.
- Prepare 2-3 mL of ice-cold extraction buffer (100 mM Tris-HCl, 50 mM NaCl, 2 mM EDTA, pH 8 with HCl) per 1 g of plant material. Add 2 mM phenylmethylsulfonyl fluoride (PMSF) from stock (100 mM) and 50 mM sodium ascorbate to the extraction buffer just before extraction.
- Place plant tissue from step 4 into the prechilled blender cup. Add a measured amount of chilled extraction buffer to the blender cup (as indicated in step 5.2). Place the blender cup on the blender. Take a pre-cut sheet of parafilm and stretch it over the top of the blender cup. Blend to homogeneity with 20-sec intervals, stirring well between blend cycles as needed.
- Transfer blended material to a beaker. Add a stir bar and stir at 4 °C for 30 min to enhance protein solubility and to allow precipitation of solids.
- Place 2 layers of Miracloth over a clean beaker on ice and pour the extract through it to remove large leaf debris. After all the extract is poured, fold the Miracloth to squeeze the residual leaf extract. The extract should appear dark green without visible particulates.
- Transfer 50 µL of this sample to a new 1.5 mL tube and label "total extract" for later analysis. Transfer the extract to centrifuge tubes. Centrifuge the remainder of plant extract at 16,000 x g for 20 min, 4 °C and transfer the supernatant to a conical tube.
- Filter the soluble extract using a 50 mL syringe and syringe glass fiber filter (0.75 µm).
- Collect 50 µL of a sample after centrifugation, label "soluble extract" for later analysis.
6. Protein G column chromatography procedure
NOTE: The protocol described here is for gravity-flow chromatography using Pierce Protein G agarose resin. If using a different resin, refer to the manufacturer's instructions for adjustments. Never let the resin run dry and prevent all liquid from draining out. Recap the outlet as needed.
- Set up a polypropylene column that holds 20 mL of sample.
- Estimate the amount of slurry needed depending on the target immunoglobulin type and its affinity to the resin. Generally, 3 mL of total slurry with 1.5 mL bed volume is sufficient for the purification of several milligrams of Ab.
- Carefully pour the required amount of resuspended slurry into the capped column. Open the column outlet from the bottom of the column and allow it to drain until most of the buffer is gone.
- Immediately pour 10 mL of wash buffer 1x PBS (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH 7.4 with HCl) on top. Let it drain and repeat this wash step 2x.
- Apply the filtered sample from step 5 to the column and collect the flowthrough—aliquot 50 µL of flowthrough for later analysis. Save the rest of the flowthrough in case the Ab did not bind to the resin.
NOTE: Re-applying flowthrough to a new column does not usually result in a good yield; hence it is advised to start with new leaf material. - Wash the resin twice with 10 mL of 1x PBS to reduce non-specific binding. If desired, aliquot 50 µL of wash as the buffer drains through the column to verify that the target Ab is not eluted with a wash buffer.
- Set up and label five tubes with 125 µL of sterile 1 M Tris-HCl at pH 8. This is to neutralize the Abs in the acidic elution buffer to avoid potential structural changes. Alternatively, add 30 µL of 2 M Tris base to get a less diluted sample.
CAUTION: During elution, UV light may be used for visualization. This does not need to be done for the duration of the elution. If UV is being used, be sure to wear appropriate PPE to avoid damage to eyes and skin. A UV light does not need to be used during the elution step. - Elute the Abs by applying 5 mL of elution buffer (100 mM glycine, pH 2.5 with HCl) to the column and collect 1 mL fractions to each designated tube from the previous step.
- Immediately regenerate the column by applying 20 mL of wash buffer, followed by 10 mL of wash buffer. Ensure that the resin is not left in an acidic environment for an extended time. Elutions should appear fluorescent, often the highest fluorescence is seen in the second elution but can vary from extraction to extraction.
- For storage, wash the resin with 10 mL of 20% ethanol in PBS and let it drain halfway. Recap the top, then the bottom of the column, and keep upright at 4 °C.
NOTE: Generally, protein G resins can be reused up to 10 times without significant loss of efficiency. Refer to the manufacturer's guidelines for specific details. - Determine Ab concentration using a spectrophotometer by measuring absorbance at 280 nm, using the elution buffer as a blank. Store the eluates in -80 °C and aliquot 50 µL of each fraction to a separate tube for further analysis.
7. SDS-PAGE for GFP-Ig fusion detection
- Prepare all samples before setting up the SDS-PAGE.
- Add 4 µL of sample buffer (6x reducing sample buffer: 3.0 mL of glycerol, 0.93 g of DTT, 1 g of SDS, 7 mL of 4x Tris (pH 6.8) 0.5 M, 1.2 mg of bromophenol blue); (6x non-reducing sample buffer: 3.0 mL of glycerol, 1 g of SDS, 7 mL of 4x Tris (pH 6.8) 0.5 M, 1.2 mg of bromophenol blue) to 20 µL of each sample (total extract, soluble extract, flowthrough, wash, all elution fractions) for analysis. Ensure that tube caps are securely fastened.
- Treat only reducing samples for 5 min in a boiling water bath, and then put samples for 5 min on ice. Spin samples in a microcentrifuge for ~5 s and load 20 µL of each sample in the order of collection into the gel wells. Load 3 µL of dual-color protein ladder in a separate well.
- Run the SDS-PAGE gel at a constant 100 V to desired protein band separation; it takes about 1.5 hours. Monitor the ladder as an indicator of protein separation.
- Visualize the gel under the UV to observe GFP fluorescence.
- If desired, stain the gel with Coomassie stain to assess total protein in each sample. Alternatively, perform western blot to evaluate target protein using specific Abs.
NOTE: Both Coomassie staining and western blot can be performed by following standard protocols47,48.
Representative Results
This study demonstrates an easy and fast method to produce recombinant proteins and visualize them throughout downstream processes. Using N. benthamiana and following the provided protocol, recombinant protein production described here can be achieved in less than a week. The overall workflow of plant expression, extraction, and purification is shown in Figure 1. The stages of plant growth from 2-week old seedlings, 4-week old plants, and 6-week old plants are displayed in Figure 1A (1-3), respectively, while Figure 1B depicts leaf morphological changes due to necrosis (Figure 1B-1) or chlorosis (Figure 1B-2). Necrosis may occur at the injection site between days 3-5 after infiltration. These changes often depend on the protein's properties being expressed and the infiltrated plants' health (further examined in discussion). Simultaneously, chlorosis can also rely on the health of plants being used (further examined in discussion). The process of Agrobacterium growth and preparation for infiltration is shown in Figure 1C. Figure 1C-1 displays isolated colonies of Agrobacterium. Figures 1C (2-5) display the media's expected appearance after it is inoculated with a single isolated colony. Refer to Figure 3 for more details on these steps. The plant infiltration process is shown in Figure 1D and begins with an un-infiltrated plant and is followed by the infiltration process. The expression, extraction, and clarification of plant proteins are displayed in Figure 1E. The leaf material is placed in a blender and is homogenized, shown in Figures 1E (1-3). A sample representing total homogenate is then taken. It is then filtered through a Miracloth (gauze or even a coffee filter can substitute for reduced expenses), and the clarified suspension is centrifuged. The centrifugation allows for the separation of the supernatant from the remaining materials, as shown in Figures 1E (4-6). The clarified supernatant is then loaded on a protein G affinity chromatography column, Figure 1F (1-3). After most of the protein is bound, Figure 1F-4, the proteins are eluted from the resin Figure 1F (5,6).
Table 1 displays the plant optimized nucleic acid sequences used to produce asGFP45 (upper row) in the pBY!KEAM-GFPasH vector used in this study to express asGFP-IgG fusion and GFP33 (lower row) in the PBYEAM-GFPHgp vector used in this study to express GFP-IgG fusion. Nucleic acid sequences were examined using the Expasy protein translate tool (https://web.expasy.org/translate/) to determine amino acid sequences.
A representative Agrobacterium plate prepared using this protocol is shown in Figure 2. Desired colonies should appear round and uniform in shape and color. Colonies closer to the center of the plate have a higher likelihood of expressing kanamycin resistance. The liquid cultures will be prepared from a single isolated colony.
The expected appearance of media containing cultures is shown in Figure 3. Upon initial inoculation of an isolated colony, LB media will appear light yellow and translucent, as shown in Figure 3A. After incubation of an isolated colony overnight at 30 °C, LB media will appear turbid. As shown in Figure 3B, objects can no longer be seen through the media when growth is present in the LB. Following centrifugation, a pellet should form at the bottom of the tube. The tube will have a clear separation of LB media above the pellet and will appear light yellow and translucent, as shown in Figure 3C. The LB media supernatant is disposed of, and the pellet is resuspended in the infiltration buffer. At an OD600 of 0.2, the media will appear turbid, as shown in Figure 3D. OD600 should be measured as described in the methods.
Figure 4 represents the process of leaf infiltration. A slight prod of the leaf with a paperclip should yield a break in the leaf epidermis that does not pass entirely through the leaf. The break should barely pierce the leaf so the infiltration buffer can be injected into the leaf, shown in Figure 4A-C. The suspension of Agrobacterium and infiltration buffer is injected directly into the break in the leaf and slightly alters the infiltrated leaf's color; see Figure 4D-F.
The appearance of leaves expressing IgG fusions is represented in Figure 5. It displays leaves that express asGFP-IgG fusions (Figure 5A) and GFP-IgG fusions (Figure 5C) under white light. If the constructs in this protocol are used, when infiltrated at a 0.2 OD600, leaves should appear healthy on days 1-5 for both leaves expressing asGFP-IgG fusions and leaves expressing GFP-IgG fusions. There may be a slight necrotic appearance at injection sites on day 5, which is usually apparent by the lightening of the plant tissue in those areas. Figure 5 also displays leaves expressing asGFP-IgG fusions (Figure 5B) and GFP-IgG fusions (Figure 5D), respectively, under long-wave UV light from the leaf's top view. Fluorescence increases in intensity as the days progress for both constructs expressed. Leaves expressing asGFP-IgG fusions tend to have slightly less intense fluorescence than leaves expressing GFP-IgG fusions on all days.
When the supernatant of the asGFP-IgG extract is added to the Protein G column, the resin will become slightly green under white light due to plant chlorophyll pigments (Figure 6A). The addition of supernatant under short-wave UV light results in the accumulation of fluorescence in the Protein G resin, as shown in Figure 6B. Note that the supernatant will also be fluorescent alone under UV light. Still, fluorescence is expected to be much more concentrated when the asGFP-IgG fusion begins to bind to the resin.
Following the passing of supernatant of asGFP-IgG through the protein G resin, the resin should illuminate under short wave UV light, as shown in Figure 7A. At this point, most of the IgG will be bound to the resin. Upon adding the elution buffer, the fluorescence contained in the protein G resin will still be visible under short-wave UV light and will begin to lose intensity as more elution buffer of low pH passes through the resin. Fluorescence will start to accumulate in the eluates (Figure 7B). Eluate fractions will vary in fluorescence. As seen in Figure 8, fluorescence is the lowest intensity in the first elution and highest intensity in the second and third eluates. Results may vary, as the fluorescence will depend on the protein's expression, harvested leaf material, and other conditions used in the experiment.
After finishing the purification process, samples are analyzed on the 10% SDS-PAGE gel under reducing conditions (sample buffer contains DTT and samples were boiled for 5 min) and non-reducing conditions (sample buffer does not contain DTT and samples were not boiled). As shown in Figure 9A, only non-reducing samples, such as in Elution 2 NR lane, will fluoresce when exposed to short wave UV light. This lane's first band is fluorescing at the full product's expected size ~200 kDa, indicating that the asGFP is still conformationally correct. The fluorescent bands near the bottom of the gel are dye from the reducing buffer. Note that asGFP loses fluorescence when exposed to temperatures at or above 95 °C for 5 minutes; this is different from eGFP (enhanced GFP), which would maintain some fluorescence under the same conditions49,50. Two bands of the ladder, 75 kDa and 25 kDa, also fluoresce. Figure 9B represents a Coomassie stain of the same gel in Figure 9A. Elutions in lanes 6-9 have been prepared under reducing conditions. When run on a gel and Coomassie-stained, these samples should display the asGFP-IgG fusion components separately. These components include the heavy chain fused to GFP (~75 kDa), the heavy chain alone (50 kDa), the light chain (25 kDa), and the asGFP itself (27 kDa). The non-reducing sample was included in the last lane of the gel for comparison purposes and should display a single large band (~200 kDa), which should be made up of two heavy chains fused to the asGFP and respective light chains. Additionally, smaller bands are likely caused by native proteases. This cleavage can be prevented with the addition of protease inhibitors and by keeping the protein extract cold at all times, including when performing the column purification. The IgG fusion protein's individual components will not be distinguishable in the non-reducing samples on the Coomassie gel.
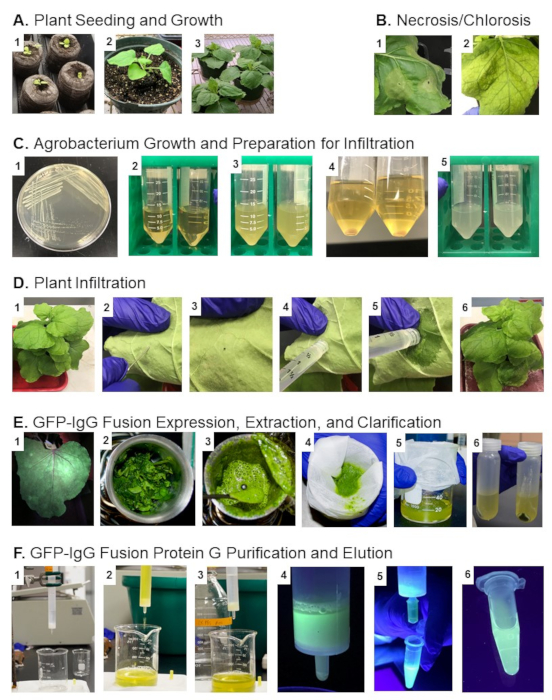

Figure 1: Workflow of plant expression, extraction, and purification processes. Please click here to view a larger version of this figure.
| Nucleotide Sequence Used | Amino Acid Sequence | ||
| Sequences used for asGFP in pBY!KEAM -GFPasH vector used in the expression of the asGFP-IgG fusion |
ATGGTGTCCAAGGGAGAGGAAGCTTCTGGAAGAGCCTTGTTC CAGTACCCTATGACTTCTAAAATCGAGTTGAATGGCGAGATCA ACGGAAAGAAGTTTAAGGTTGCTGGAGAGGGTTTCACCCCTTC ATCTGGAAGATTCAATATGCACGCTTACTGTACTACCGGAGAC TTGCCTATGTCCTGGGTTGTTATAGCTTCCCCGCTTCAGTACGG GTTTCACATGTTTGCCCACTACCCTGAGGATATCACTCACTTCTT CCAAGAATGTTTTCCTGGGTCTTATACTCTCGACAGAACTTTGA GGATGGAGGGAGACGGTACTCTTACTACTCACCACGAGTACTC CCTTGAGGACGGTTGCGTTACTTCCAAGACTACTTTGAACGCTT CTGGATTCGACCCCAAGGGAGCCACTATGACTAAGTCTTTCGT CAAACAGCTCCCAAACGAGGTCAAAATCACCCCACACGGGCCA AATGGTATTAGACTTACTTCCACTGTTCTCTACCTTAAGGAGGA CGGAACTATCCAGATCGGAACTCAAGACTGCATCGTTACCCCA GTTGGCGGCAGAAAAGTTACTCAGCCTAAGGCTCACTTTCTTC ATACTCAGATCATTCAGAAGAAGGACCCAAACGACACCAGAG ATCACATCGTTCAGACTGAGCTTGCCGTTGCTGGAAATCTTTG GCACGGCATGGATGAGCTTTACAAGA |
MVSKGEEASGRALF QYPMTSKIELNGEI NGKKFKVAGEGFTP SSGRFNMHAYCTT GDLPMSWVVIASPL QYGFHMFAHYPEDI THFFQECFPGSYTL DRTLRMEGDGTLTT HHEYSLEDGCVTSK TTLNASGFDPKGAT MTKSFVKQLPNEVK ITPHGPNGIRLTSTV LYLKEDGTIQIGTQD CIVTPVGGRKVTQP KAHFLHTQIIQKKDP NDTRDHIVQTELAV AGNLWHGMDELY K |
|
| Sequences used for GFP in pBYEAMGFPHgp vector used in the expression of GFP-IgG fusion |
ATGGCTAACAAGCACCTCTCATTGTCTCTCTTCCTTGTGCTCCTT GGTCTTTCTGCTTCTCTTGCTTCTGGTATGGTGAGCAAGGGCG AGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGG ACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGG GCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCA TCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGT GACCACCTTCAGCTACGGCGTGCAGTGCTTCAGCCGCTACCCC GACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCG AAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGG CAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACAC CCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGA GGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAA CAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGG CATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGC AGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCG GCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCAC CCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCA CATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCAC GGCATGGACGAGCTGTACAAGA |
MANKHLSLSLFLVLL GLSASLASGMVSKG EELFTGVVPILVELD GDVNGHKFSVSGE GEGDATYGKLTLKFI CTTGKLPVPWPTLV TTFSYGVQCFSRYP DHMKQHDFFKSA MPEGYVQERTIFFK DDGNYKTRAEVKFE GDTLVNRIELKGIDF KEDGNILGHKLEYN YNSHNVYIMADKQ KNGIKVNFKIRHNIE DGSVQLADHYQQN TPIGDGPVLLPDNH YLSTQSALSKDPNEK RDHMVLLEFVTAA GITH |
|
Table 1: Sequences used to create asGFP and GFP
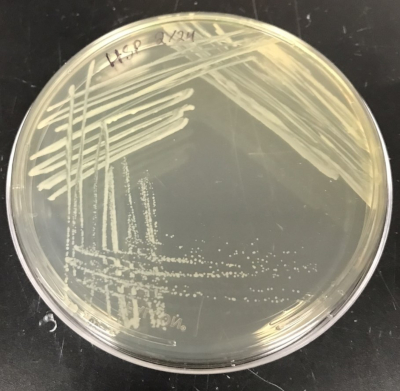
Figure 2: Agrobacterium colonies grown on LB plate containing kanamycin. Please click here to view a larger version of this figure.
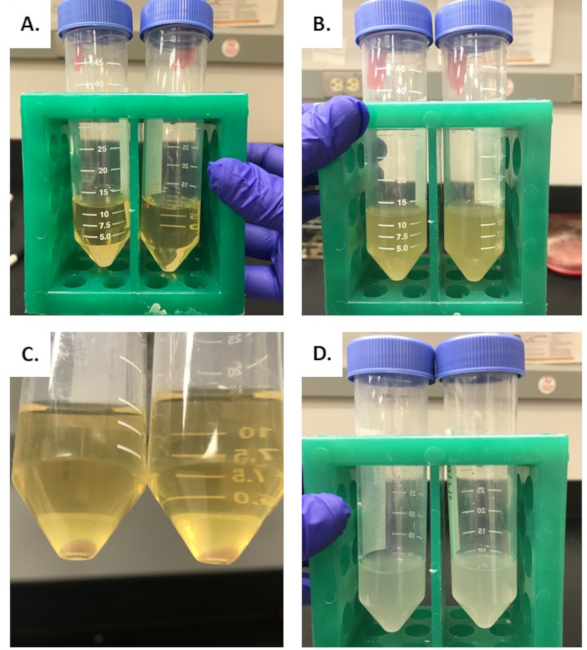

Figure 3: Appearance of media throughout the growth and processing of Agrobacterium. (A) The appearance of LB media immediately following inoculation of isolated Agrobacterium colony. (B) The presence of media after overnight incubation of an isolated colony at 30°C. (C) The appearance of media after cultures are spun down for 20 min at 4,500 x g. (D) Pellet resuspended in the infiltration buffer. Please click here to view a larger version of this figure.
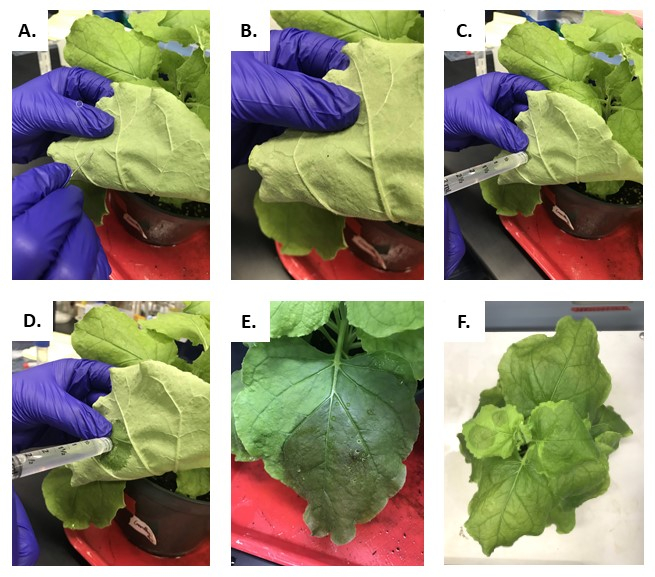

Figure 4: Process of infiltration of N. benthamiana plants. (A-B) Slightly poking the leaf results in a subtle hole at the top of the leaf. (C-D) Injection of Agrobacterium suspension into leaf. (E) Infiltrated Plant leaf from the top view. (F) Plant with multiple leaves infiltrated from the top view. Please click here to view a larger version of this figure.
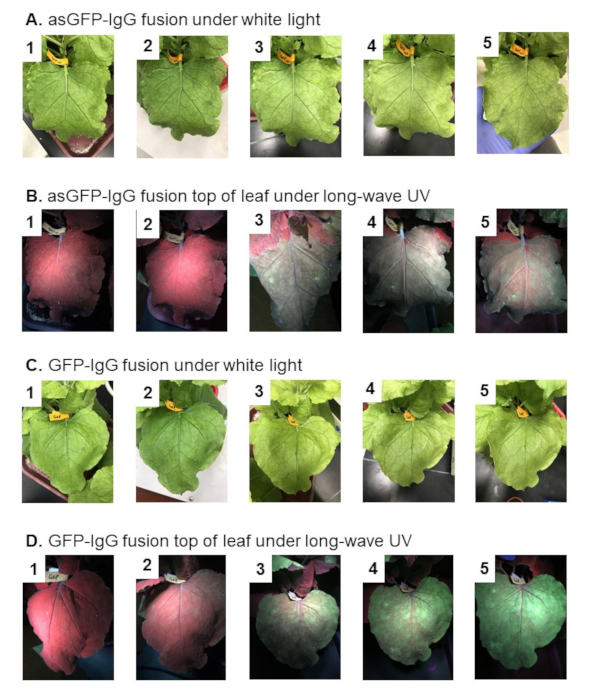
Figure 5: Visualization of leaves containing asGFP-IgG fusion and GFP-IgG fusion begin on day 1 post infiltration (dpi) in the first row, leading up to day 5 dpi in the last row for all conditions. A) asGFP-IgG fusion under white light. B) asGFP-IgG fusion under long-wave UV. C) GFP-IgG fusion under white light. D) GFP-IgG fusion under long-wave UV. Please click here to view a larger version of this figure.
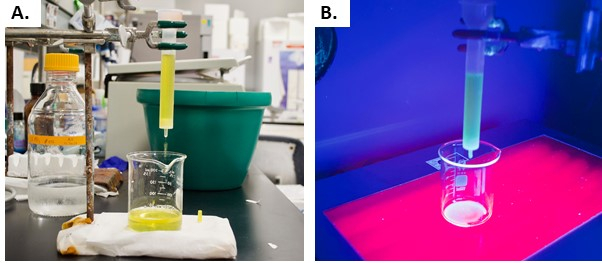

Figure 6: Supernatant of the asGFP-IgG extract being added to the Protein G column. A) Addition under white light. B) Addition under short wave UV light. Please click here to view a larger version of this figure.
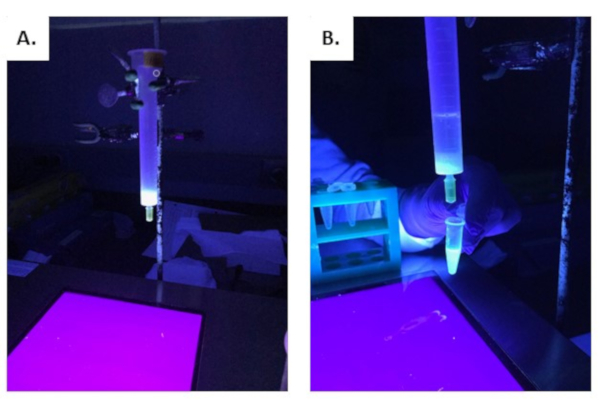

Figure 7: Protein G resin under short-wave UV light after supernatant of the asGFP-IgG extract was run through the column. A) Protein G resin under short-wave UV light. B) Protein G resin upon elution of proteins under low PH conditions. Please click here to view a larger version of this figure.
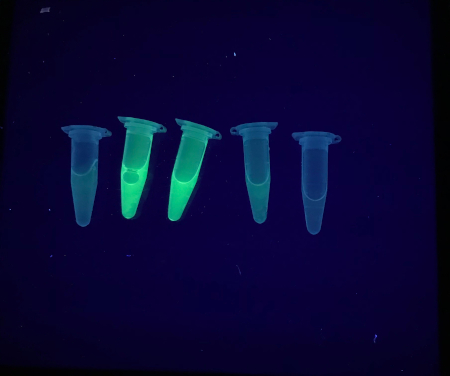
Figure 8: Purified elutions of asGFP-IgG obtained after exposure to low pH conditions through purification. Please click here to view a larger version of this figure.
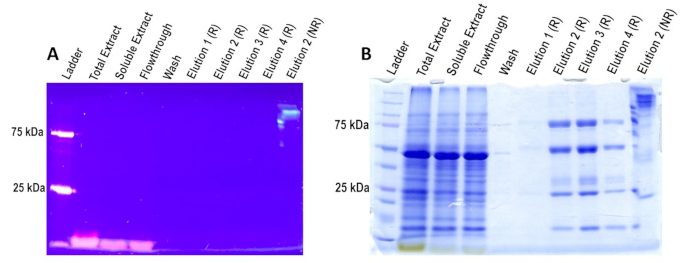
Figure 9: SDS-PAGE of Column Samples. Samples labeled with "R" are in reducing conditions, and samples labeled with "NR" are in non-reducing conditions. A) Under UV light, only non-reducing samples fluoresce in the 10% polyacrylamide gel, as seen in Elution 2 NR lane. The 75 kDa and 25 kDa ladder bands also fluoresce. B) Coomassie staining of the same gel reveals the presence of all proteins in the sample. In the reduced elutions, the IgG fusion without light chain, the heavy chain, light chain, and possibly degraded GFP are present at ~75 kDa, ~50 kDa, ~25 kDa, and ~10 kDa, respectively. In contrast, in the non-reduced elutions, one prominent band is present, consistent with the size of the entire asGFP-IgG fusion (~200 kDa). Please click here to view a larger version of this figure.
Discussion
This protocol can be utilized for the visual verification of any recombinant Ab or recombinant protein produced in N. benthamiana plants, including those that require temporary exposure to acidic environments for column purification purposes42,43,44. Furthermore, the fusion of asGFP to other proteins in different expression systems can be a useful tool for experimental visualization and education. The protocol herein can further be scaled to larger and smaller amounts of leaf material to produce the desired amount of recombinant protein. The described methods take advantage of previous studies that have identified and made versions of GFP that remain stable under acidic conditions46. Comprehensive prior studies have created immunoglobulin domains fused to a protein of interest, often termed IgG-fusions, which this protocol can also accommodate28. By creating and expressing an IgG-fusion composed of a humanized IgG1 fused to a GFP and asGFP, we were able to visualize the presence of the desired protein during the expression, extraction, and purification processes.
If a skilled researcher follows this protocol, leaves will begin to display necrosis signs at the infiltration sites between days 4-5. However, when using the described vectors, infiltrated areas of leaves should appear healthy up until day 5 with proper care. It is important to note that Agrobacterium infection on its own will eventually result in necrosis of the plant leaves due to the accumulation of reactive oxygen species (ROS) as part of the plant cell immune response51,52. This immune response and the resultant level of necrosis can vary based on several factors51, including the cellular targeting, the protein's location, the type of protein being produced, the expression vector, and the strain of Agrobacterium used53,54. Also, variations in the optical density (OD600) of the Agrobacterium used for infiltration can affect the levels of necrosis55. Many expression vectors utilize proteins that bind retinoblastoma protein to keep the plant cells in synthesis (S)-phase and increase cell division and transformation frequency56,57. Increases in protein production, such as those caused by binding to retinoblastoma protein, can lead to necrosis56,57. Advances in vector design, such as those used in modified versions of the geminivirus BeYDV replicons used in this protocol, have minimized necrosis while maintaining high protein expression levels58. Also, BeYDV replicons are non-competing, providing expression of multiple proteins on a single cassette without known size limitations53,59.
Several factors affect plant growth before and after infiltration, which might eventually lead to low protein yield. When seeding plants, too many seeds per plant pellet can result in many small plant sprouts leading to more modest plant growth. Hence, reducing the seed number per peat pellet and removing the extra sprouts after a week will result in better plant growth. Maintaining proper soil moisture is another factor affecting overall plant health. Overwatering, underwatering, adding too much or too little fertilizer might contribute to chlorosis and affect the plant health60,61,62. Necrosis and chlorosis can additionally be caused by the production of a protein that causes cell stress. This phenomenon has been seen many times with the expression of recombinant immune complexes (RIC)56. We have observed that changes in protein structure and fusion of proteins can help minimize necrosis; however, some proteins can remain toxic to plants even after various modifications. If using the expression vectors outlined herein, extraction of protein may be performed early and before the onset of significant necrosis, resulting in high protein yield56.
Different growth conditions can slow or even inhibit Agrobacterium growth. Agrobacterium grows optimally at 28 °C-30 °C and experiences a heat shock when incubated above 30 °C, producing cell division errors62. Growth can also be impeded by the addition of too much rifampicin, as different Agrobacterium strains are more or less naturally resistant to this antibiotic62. The bacterial culture prepared for infiltration with significantly higher OD600 than recommended will likely cause necrosis55. A slightly higher OD600 of the culture usually does not affect the yield, but if it is lower than 0.1, the protein yield might be considerably reduced. Accumulation of dead cells can occur under two circumstances; 1) the culture was overgrown, leading to a significant fraction of the OD being dead cells, and 2) damaging/killing the Agrobacterium after growth, such as with chemical residue or high centrifuge speeds. Infiltrations using an increased number of dead cells in the culture might reduce protein expression. Moreover, puncturing the leaves by applying too much pressure can damage the leaves and hence might lead to premature necrosis. Considering these possible factors when expressing recombinant proteins in Nicotiana benthamiana, can lead to enhanced protein production.
Obtaining low protein yield could be due to some issues in the extraction and purification steps. Firstly, the extraction buffer might need optimization depending on the protein of interest. During blending, plant material should be homogenous without any visible leaf pieces. Next, some proteins require detergents for solubilization in the extraction buffer, such as Tween-20 or Triton. Other proteins might need urea at high concentrations ~7.5 M for solubilization, while some can be extracted with PBS only. Degradation of protein can occur if buffers, plant tissue, centrifuges, etc. are not kept cool during the extraction process. Lack of protease inhibitors and sodium ascorbate or similar chemicals in the extraction buffer can also cause degradation or aggregation. Some protease inhibitors like PMSF degrade quickly, and sodium ascorbate takes some time to become aqueous. Overall, researchers should determine optimal conditions for their protein of interest.
The purification of IgG-fusions includes few steps that might need modifications if low protein yield is obtained. Analyzing the sample aliquots collected during the entire process by SDS-PAGE and Western will help to identify the fault of the methods. For example, if the flowthrough contains a substantial amount of protein, then the binding of the protein can be facilitated by changing the pH of the buffer. Using high concentrations of detergents during the extraction process might affect the binding property of the resin, especially if the resin is reused several times. Proper storage of the resin, as described in the methods, is vital for the resin's lifespan. Moreover, if the washing step removes the protein of interest from the resin, the buffers might need to be remade to solve this problem. Other issues with protein purification might be due to misfolded or degraded proteins, which might require further analysis of the overall protein design. Referencing the described troubleshooting may increase the efficiency of the purification using this protocol.
The described GFP-IgG fusion purification is helpful in a teaching environment. Visualization is fundamental to science education because it allows learners to comprehend concepts more easily39. Students often report misunderstandings in addition to difficulty understanding concepts at the molecular level39. In particular, the experiments that require an understanding of the specific protein location at each step can be modified by tagging protein of interest with fluorescent molecules. Therefore, GFP or asGFP, depending on the pH environment used, can be utilized to harness their fluorescence to facilitate elucidation of the protein purification technique for students.
In summary, we describe a simple method for expression and purification of a recombinant Ab fused to a GFP in N. benthamiana plants. This protocol can be used for the purification of an Ab fused to any desired target protein. The process can be edited to accommodate various amounts of leaf material and allows for visual determination of protein presence before, during, and after the conclusion of the protein extraction and purification process. These methods can be useful as controls and can be purposed for teaching techniques.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
We thank Maria Pia DiPalma for editing the video. We also thank the Office of Educational Outreach and Student Services at Arizona State University for their generous publication fee assistance. Research for this protocol was supported by the School of Life Sciences, Arizona State University.
Materials
| 5 mL syringe | any | N/A | |
| 50 mL syringe | any | N/A | |
| Agar | SIGMA-ALDRICH | A5306 | |
| Blender with cups | any | N/A | |
| Bromophenol blue | Bio-Rad | 1610404 | |
| DTT (DL-Dithiothreitol) | MP BIOMEDICALS | 219482101 | |
| EDTA (Ethylenedinitrilo)tetraacetic acid | SIGMA-ALDRICH | E-6760 | |
| Ethanol | any | N/A | |
| Glycerol | G-Biosciences | BTNM-0037 | |
| Glycine | SIGMA-ALDRICH | G7126-500G | |
| HCl (Hydrochloric acid) | EMD MILLIPORE CORPORATION | HX0603-4 | |
| Heating block | any reputable supplier | N/A | |
| Jiffy-7 727 w/hole peat pellets | Hummert International | 14237000 | |
| Kanamycin | Gold Biotechnology Inc | K-120-100 | |
| KCl (Potassium Chloride) | SIGMA-ALDRICH | P9541-500G | |
| KH2PO4 (Potassium Phosphate) | J.t.baker | 3248-05 | |
| KOH (Potassium Hydroxide) | VWR | BDH0262 | |
| Magnesium sulfate heptahydrate | SIGMA-ALDRICH | M2773 | |
| MES (2-(N-Morpholino)ethanesulfonic acid) | SIGMA-ALDRICH | M8250 | |
| Miracloth | Millipore | 4 75855-1R | |
| Moisture control potting mix | Miracle-Gro | 755783 | |
| Na2HPO4 (Sodium Phosphate) | J.t.baker | 3827-01 | |
| NaCl (Sodium Chloride) | Santa Cruz Biotechnology | sc-203274C | |
| Nicotiana benthamiana seeds | any reputable supplier | N/A | |
| PMSF (Phenylmethylsulfonyl Fluoride) | G-Biosciences | 786-787 | |
| Polypropylene Column | any | N/A | |
| Precision Plus Protein Dual Color Standards | Bio-Rad | 1610394 | |
| Protein G resin | Thermo Fisher Scientific | 20399 | |
| Rifampicin | Gold Biotechnology Inc | R-120-25 | |
| SDS (Sodium Dodecyl Sulfate) | G-Biosciences | DG093 | |
| Sodium Ascorbate | SIGMA-ALDRICH | A7631-500G | |
| Spectrophotometer | any reputable supplier | N/A | |
| Titan3 0.75 µm glass fiber filter | ThermoScientific | 40725-GM | |
| Tray for peat pellets with dome | any | N/A | |
| TRIS Base | J.t.baker | 4109-02 | |
| Tris-HCl | Amresco | M108-1KG | |
| Tryptone | SIGMA-ALDRICH | 17221 | |
| UV lamp | any | N/A | |
| Water Soluble All Purpose Plant Food | Miracle-Gro | 2000992 | |
| Yeast extract | SIGMA-ALDRICH | 9182 |
Referenzen
- Lu, R. M., et al. Development of therapeutic antibodies for the treatment of diseases. Journal of Biomedical Science. 27 (1), (2020).
- Kaplon, H., Reichert, J. M. Antibodies to watch in 2019. mAbs. 11 (2), 219-238 (2019).
- The top 20 drugs by U.S. sales. FiercePharma Available from: https://www.fiercepharma.com/special-report/top-20-drugs-by-2018-u-s-sales (2018)
- Grilo, A. L., Mantalaris, A. The Increasingly Human and Profitable Monoclonal Antibody Market. Trends in Biotechnology. 37 (1), 9-16 (2019).
- Kim, M. -. Y., et al. Novel vaccination approach for dengue infection based on recombinant immune complex universal platform. Vaccine. 33 (15), 1830-1838 (2015).
- Twyman, R. M., Schillberg, S., Fischer, R. Transgenic plants in the biopharmaceutical market. Expert Opinion on Emerging Drugs. 10 (1), 185-218 (2005).
- Stoger, E., Fischer, R., Moloney, M., Ma, J. K. -. C. Plant Molecular Pharming for the Treatment of Chronic and Infectious Diseases. Annual Review of Plant Biology. 65 (1), 743-768 (2014).
- Buyel, J. F., Fischer, R. Predictive models for transient protein expression in tobacco (Nicotiana tabacum L.) can optimize process time, yield, and downstream costs. Biotechnology and Bioengineering. 109 (10), 2575-2588 (2012).
- Yao, J., Weng, Y., Dickey, A., Wang, K. Y. Plants as factories for human pharmaceuticals: Applications and challenges. International Journal of Molecular Sciences. 16 (12), 28549-28565 (2015).
- Mor, T. S., Moon, Y. -. S., Palmer, K. E., Mason, H. S. Geminivirus vectors for high-level expression of foreign proteins in plant cells. Biotechnology and Bioengineering. 81 (4), 430-437 (2003).
- Hefferon, K. L. Plant virus expression vectors set the stage as production platforms for biopharmaceutical proteins. Virology. 433 (1), 1-6 (2012).
- Yamamoto, T., et al. Improvement of the transient expression system for production of recombinant proteins in plants. Scientific Reports. 8 (1), 4755 (2018).
- Zhong, G. Y., et al. Commercial production of aprotinin in transgenic maize seeds. Molecular Breeding. 5 (4), 345-356 (1999).
- Giddings, G., Allison, G., Brooks, D., Carter, A. Transgenic plants as factories for biopharmaceuticals. Nature Biotechnology. 18 (11), 1151-1155 (2000).
- Fulton, A., Lai, H., Chen, Q., Zhang, C. Purification of monoclonal antibody against Ebola GP1 protein expressed in Nicotiana benthamiana. Journal of Chromatography A. 1389, 128-132 (2015).
- Diamos, A., et al. A highly expressing, soluble, and stable plant-made IgG fusion carrying Zika virus envelope domain III elicits potent immunogenic responses in mice without adjuvant. Frontiers in immunology. 3140 (11), (2020).
- Hunter, J. G. L., et al. Evaluation of a toxoid fusion protein vaccine produced in plants to protect poultry against necrotic enteritis. PeerJ. 2019 (3), 6600 (2019).
- Dent, M., et al. Plant-produced anti-dengue virus monoclonal antibodies exhibit reduced antibody-dependent enhancement of infection activity. Journal of General Virology. 97 (12), 3280-3290 (2016).
- Diamos, A. G., et al. High Level Production of Monoclonal Antibodies Using an Optimized Plant Expression System. Frontiers in Bioengineering and Biotechnology. 7, 472 (2020).
- Olinger, G. G., et al. Delayed treatment of Ebola virus infection with plant-derived monoclonal antibodies provides protection in rhesus macaques. Proceedings of the National Academy of Sciences of the United States of America. 109 (44), 18030-18035 (2012).
- Mastrangeli, R., Palinsky, W., Bierau, H. Glycoengineered antibodies: towards the next-generation of immunotherapeutics. Glycobiology. 29 (3), 199-210 (2019).
- Montero-Morales, L., Steinkellner, H. Advanced Plant-Based Glycan Engineering. Frontiers in Bioengineering and Biotechnology. 6, 81 (2018).
- Joosten, V., Lokman, C., vanden Hondel, C. A. M. J. J., Punt, P. J. The production of antibody fragments and antibody fusion proteins by yeasts and filamentous fungi. Microbial Cell Factories. 2 (1), 1-15 (2003).
- Müller, D. Antibody fusions with immunomodulatory proteins for cancer therapy. Pharmacology and Therapeutics. 154, 57-66 (2015).
- Bootz, F., Neri, D. Immunocytokines: A novel class of products for the treatment of chronic inflammation and autoimmune conditions. Drug Discovery Today. 21 (1), 180-189 (2016).
- Jafari, R., Zolbanin, N. M., Rafatpanah, H., Majidi, J., Kazemi, T. Fc-fusion Proteins in Therapy: An Updated View. Current Medicinal Chemistry. 24 (12), (2017).
- Brinkmann, U., Kontermann, R. E. The making of bispecific antibodies. mAbs. 9 (2), 182-212 (2017).
- Mason, H. S. Recombinant immune complexes as versatile and potent vaccines. Human Vaccines & Immunotherapeutics. 12 (4), 988-989 (2016).
- Mason, H. S. Recombinant immune complexes as versatile and potent vaccines. Human Vaccines and Immunotherapeutics. 12 (4), 988-989 (2016).
- Diamos, A. G., et al. Vaccine synergy with virus-like particle and immune complex platforms for delivery of human papillomavirus L2 antigen. Vaccine. 37 (1), 137-144 (2019).
- Diamos, A. G., et al. Codelivery of improved immune complex and virus-like particle vaccines containing Zika virus envelope domain III synergistically enhances immunogenicity. Vaccine. 38 (18), 3455-3463 (2020).
- Prasher, D. C., Eckenrode, V. K., Ward, W. W., Prendergast, F. G., Cormier, M. J. Primary structure of the Aequorea victoria green-fluorescent protein. Gene. 111 (2), 229-233 (1992).
- Chalfie, M., Tu, Y., Euskirchen, G., Ward, W. W., Prasher, D. C. Green fluorescent protein as a marker for gene expression. Science. 263 (5148), 802-805 (1994).
- Palm, M., Baumstark-Khan, C., Horneck, G. Green Fluorescent Protein (GFP) Expression in Mammalian Cells After UV-Irradiation. Fundamentals for the Assessment of Risks from Environmental Radiation. , 311-316 (1999).
- Paemeleire, K., et al. Intercellular calcium waves in HeLa cells expressing GFP-labeled connexin 43, 32, or 26. Molecular Biology of the Cell. 11 (5), 1815-1827 (2000).
- Harper, B. K., Stewart, C. N. Patterns of Green Fluorescent Protein Expression in Transgenic Plants. Plant Molecular Biology Reporter. 18 (2), 141 (2000).
- Kaishima, M., Ishii, J., Matsuno, T., Fukuda, N., Kondo, A. Expression of varied GFPs in Saccharomyces cerevisiae: Codon optimization yields stronger than expected expression and fluorescence intensity. Scientific Reports. 6, (2016).
- . Glow in the Dark: Fluorescent Proteins as Cell and Tissue-Specific Markers in Plants Available from: https://pubmed.ncbi.nlm.nih.gov/21772029/ (2020)
- Mnguni, L. E. The theoretical cognitive process of visualization for science education. SpringerPlus. 3 (1), 1-9 (2014).
- Shaner, N. C., Patterson, G. H., Davidson, M. W. Advances in fluorescent protein technology. Journal of Cell Science. 120 (24), 4247-4260 (2007).
- Bjorck, L., Kronvall, G. Purification and some properties of streptococcal protein G, a novel IgG-Binding reagent. The Journal of Immunology. 133 (2), (1984).
- Andrew, S. M., Titus, J. A. Purification of Immunoglobulin G. Current Protocols in Immunology. 21 (1), 1-12 (2001).
- Hober, S., Nord, K., Linhult, M. Protein A chromatography for antibody purification. Journal of Chromatography B: Analytical Technologies in the Biomedical and Life Sciences. 848 (1), 40-47 (2007).
- Vola, R., Lombardi, A., Tarditi, L., Björck, L., Mariani, M. Recombinant proteins L and LG: efficient tools for purification of murine immunoglobulin G fragments. Journal of Chromatography B: Biomedical Sciences and Applications. 668 (2), 209-218 (1995).
- Shinoda, H., Ma, Y., Nakashima, R., Sakurai, K., Matsuda, T., Nagai, T. Acid-Tolerant Monomeric GFP from Olindias formosa. Cell Chemical Biology. 25 (3), 330-338 (2018).
- Roberts, T. M., et al. Identification and Characterisation of a pH-stable GFP. Scientific Reports. 6 (1), 1-9 (2016).
- Brunelle, J. L., Green, R. Coomassie blue staining. Methods in Enzymology. 541, 161-167 (2014).
- Mahmood, T., Western Yang, P. C. Western blot: Technique, theory, and trouble shooting. North American Journal of Medical Sciences. 4 (9), 429-434 (2012).
- Vessoni Penna, T. C., Ishii, M., Cholewa, O., de Souza, L. C. Thermal characteristics of recombinant green fluorescent protein (GFPuv) extracted from Escherichia coli. Letters in Applied Microbiology. 38 (2), 135-139 (2004).
- Kang, S. J., Park, E. A., Lee, D. H., Hong, K. W. Comparison of the stability of eGFP displayed on the Bacillus subtilis spore surface using CotB and C-terminally truncated CotB proteins as an anchoring motif under extreme conditions. Applied Biological Chemistry. 62 (1), 41 (2019).
- Kuta, D. D., Tripathi, L. Agrobacterium-induced hypersensitive necrotic reaction in plant cells: a resistance response against Agrobacterium-mediated DNA transfer. African Journal of Biotechnology. 4 (8), 752-757 (2005).
- Qiusheng, Z., Bao, J., Likun, L., Xianhua, X. Effects of antioxidants on the plant regeneration and GUS expressive frequency of peanut (Arachis hypogaea) explants by Agrobacterium tumefaciens. Plant Cell, Tissue and Organ Culture. 81 (1), 83-90 (2005).
- Diamos, A. G., Rosenthal, S. H., Mason, H. S. H.S. 5′ and 3′ Untranslated Regions Strongly Enhance Performance of Geminiviral Replicons in Nicotiana benthamiana Leaves. Frontiers in Plant Science. 7, 200 (2016).
- Huang, Z., Chen, Q., Hjelm, B., Arntzen, C., Mason, H. A DNA replicon system for rapid high-level production of virus-like particles in plants. Biotechnology and Bioengineering. 103 (4), 706-714 (2009).
- Diamos, A. G., Mason, H. S. Modifying the Replication of Geminiviral Vectors Reduces Cell Death and Enhances Expression of Biopharmaceutical Proteins in Nicotiana benthamiana Leaves. Frontiers in Plant Science. 9, 1974 (2019).
- Gordon-Kamm, W., et al. Stimulation of the cell cycle and maize transformation by disruption of the plant retinoblastoma pathway. Proceedings of the National Academy of Sciences of the United States of America. 99 (18), 11975-11980 (2002).
- Villemont, E., Dubois, F., Sangwan, R. S., Vasseur, G., Bourgeois, Y., Sangwan-Norreel, B. S. Role of the host cell cycle in the Agrobacterium-mediated genetic transformation of Petunia: Evidence of an S-phase control mechanism for T-DNA transfer. Planta. 201 (2), 160-172 (1997).
- Regnard, G. L., Halley-Stott, R. P., Tanzer, F. L., Hitzeroth, I. I., Rybicki, E. P. High level protein expression in plants through the use of a novel autonomously replicating geminivirus shuttle vector. Plant Biotechnology Journal. 8 (1), 38-46 (2010).
- Chen, Q., He, J., Phoolcharoen, W., Mason, H. S. Geminiviral vectors based on bean yellow dwarf virus for production of vaccine antigens and monoclonal antibodies in plants. Human Vaccines. 7 (3), 331-338 (2011).
- Morton, T. G., Gold, A. J., Sullivan, W. M. Influence of Overwatering and Fertilization on Nitrogen Losses from Home Lawns. Journal of Environmental Quality. 17 (1), 124-130 (1988).
- Brown, J. C. Iron Chlorosis in Plants. Advances in Agronomy. 13, 329-369 (1961).
- Morton, E. R., Fuqua, C. Laboratory Maintenance of Agrobacterium. Current Protocols in Microbiology. , (2012).

