CRISPR-Cas9-Mediated Precise Knock-In Edits in Zebrafish Hearts
Summary
This protocol describes an approach to facilitate precise knock-in edits in zebrafish embryos using CRISPR-Cas9 technology. A phenotyping pipeline is presented to demonstrate the applicability of these techniques to model a Long QT Syndrome-associated gene variant.
Abstract
Clustered regularly interspaced short palindromic repeats (CRISPR) in animal models enable precise genetic manipulation for the study of physiological phenomena. Zebrafish have been used as an effective genetic model to study numerous questions related to heritable disease, development, and toxicology at the whole-organ and -organism level. Due to the well-annotated and mapped zebrafish genome, numerous tools for gene editing have been developed. However, the efficacy of generating and ease of detecting precise knock-in edits using CRISPR is a limiting factor. Described here is a CRISPR-Cas9-based knock-in approach with the simple detection of precise edits in a gene responsible for cardiac repolarization and associated with the electrical disorder, Long QT Syndrome (LQTS). This two-single-guide RNA (sgRNA) approach excises and replaces the target sequence and links a genetically encoded reporter gene. The utility of this approach is demonstrated by describing non-invasive phenotypic measurements of cardiac electrical function in wild-type and gene-edited zebrafish larvae. This approach enables the efficient study of disease-associated variants in a whole organism. Furthermore, this strategy offers possibilities for the insertion of exogenous sequences of choice, such as reporter genes, orthologs, or gene editors.
Introduction
CRISPR-based gene editing strategies in animal models enable the study of genetically heritable disease, development, and toxicology at the whole-organism level1,2,3. Zebrafish provide a powerful model that is closer in numerous physiological aspects to humans than murine or human-derived cell models4. An extensive array of genetic tools and strategies have been used in zebrafish for both forward5 and reverse genetic screening6. Comprehensive genetic mapping and annotation in zebrafish have facilitated gene-editing approaches as a primary technique to engineer targeted gene knockouts (KOs) and precise knock-ins (KIs)7.
Despite this, generating precise KI edits in zebrafish is limited by low efficiencies and the difficulty of accurate detection. Although transcription factor-like effector nucleases (TALENs) have been successfully used and optimized for KIs8, CRISPR provides an improved gene-editing strategy with simpler sgRNA targeting. Numerous studies have used CRISPR to generate precise KIs in zebrafish9,10,11,12,13,14,15,16,17,18,19,20, although these edits generated through CRISPR-mediated homology-directed repair (HDR) tend to be inefficient with low intrinsic success rates that require genotyping as a primary screen9,10,14,21. This demonstrates the need for an efficient KI CRISPR system in zebrafish, as well as a reliable high-throughput system for detecting precise edits.
The goal of this study was to describe a platform for generating a precise cardiac gene KI in zebrafish hearts with simple and high-throughput detection of successful edits. A CRISPR-Cas9-based two-sgRNA exon replacement approach is described, which is based on a TALEN approach8. This approach involves excision of the target sequence using two-sgRNA guides and replacement with an exogenous template sequence that contains the KI of interest as well as a genetically encoded intronic reporter gene (Figure 1). The integration of a genetically encoded fluorescent reporter within the target gene intronic sequence enables the efficient detection of positive edits. A phenotyping platform is then described for assessing cardiac electrical function in zebrafish larvae for non-invasive characterization of the gene variants associated with inherited LQTS, a cardiac electrical disorder that predisposes individuals to sudden cardiac death.
These approaches will enhance the access to and use of zebrafish KI gene edits to model inherited diseases and address biological and physiological questions, such as mapping gene expression patterns, and developmental regulation. Since zebrafish hearts better parallel human cardiac electrophysiological characteristics than murine models, they may be particularly attractive as a genetically tractable system for cardiac disease modeling7,22,23.
Protocol
Studies using zebrafish were conducted in agreement with the policies and procedures of the Simon Fraser University Animal Care Committee and the Canadian Council of Animal Care and were completed under protocol # 1264K-18.
1. Design of CRISPR components for precise edits
- To design the two-sgRNA guides that will be used to excise the sequence containing the KI target site, first identify the zebrafish ortholog for the gene of interest.
NOTE: Figure 2 provides a summary overview of the steps to engineer precise edits using the two-sgRNA CRISPR-Cas9 approach. - Next, use a design software tool, such as CRISPOR24, which includes selection for Danio rerio as a species and of the Cas enzyme to be used.
NOTE: The gene of interest for this study was zkcnh6a (Ensembl Transcript ID: ENSDART00000090809.6; UniProt Protein ID: B3DJX4), and the target mutation was amino acid, R56Q.- For the two-sgRNA approach, choose one sgRNA location that precedes the target exon and a second sgRNA that is located within the immediate downstream intron.
- Ensure that the selected sgRNAs have high specificity and low predicted off-target binding. Use CRISPOR rankings to identify guides with minimal off-target binding. Do not consider guides with no mismatches in the seed sequence of potential off-targets.
- Identify the most likely potential off-target sites (based on CRISPOR scores, select the top three exon potential off-target sites) for PCR-based Sanger sequencing genotyping in step 6.1.3.
- Once the two-sgRNAs have been selected, obtain the reverse complement for each, such that there are four oligonucleotides to be used: two complementary oligos that precede the mutation and two complementary oligos that are downstream.
- On each of the four oligos, add compatible restriction sites for incorporation into a guide plasmid of choice; for integration into the DR274 plasmid, use a 5' BsaI restriction site to create an overhang. Ensure that the Bsa1 recognition site is engineered at the 5' end of the guide selected from CRISPOR and the Bsa1 recognition site is engineered at the 5' end of the complementary strands to ensure correct orientation of the guides in the DR274 plasmid (see Figure 3).
- To design the exogenous template used for HDR in zebrafish (Figure 1), choose two sequence fragments that will flank the mVenus YFP reporter gene housed in the pKHR5 plasmid.
NOTE: The intended modification/edit can be included in either the upstream or downstream fragment.- Using Ensembl, locate the target site within the gene sequence of interest, including approximately the 2 kb flanking sequence (homology arm), which will be used to make the template.
NOTE: Homology arms may be symmetrical or asymmetrical25,26 and approximately 1 kb each, upstream and downstream of the target site. - Divide the template into two segments that will be inserted either side of the mVenus YFP reporter gene (see Figure 4). Ensure that the split site is in an intron so that the coding sequence is not interrupted.
NOTE: If the gene is well characterized, check for functional roles in the intron, such as splicing sites or regulatory regions. Regions close to the 5' or 3' ends are more often involved in mRNA splicing. - Incorporate modifications into the template sequence that include i) silent mutations in the guide protospacer adjacent motif (PAM) or seed sequence (be aware of alternate PAM sites that the Cas enzyme might target) to prevent Cas enzyme recutting; ii) the modification of interest; iii) the creation of restriction endonuclease sites to facilitate cloning into the pKHR5 plasmid, which contains the mVenus YFP reporter gene (see Figure 3).
NOTE: In this study, the first template segment contained XhoI upstream of the R56Q mutation site and SalI downstream, while the second template segment had EcoRI upstream of the guide target sequence and BamHI downstream. If any of the selected restriction sites are present within the template sequence, mutations to silence these will be required, or alternate approaches such as Gibson Assembly can be used.
- Using Ensembl, locate the target site within the gene sequence of interest, including approximately the 2 kb flanking sequence (homology arm), which will be used to make the template.
2. Preparation of CRISPR components for embryo microinjection
- Prepare the Cas9 for microinjection 1 week prior to the microinjections. Use Cas9 protein or prepare Cas9 mRNA via in vitro transcription.
NOTE: In this study, Cas9 mRNA was used, since efficiencies tended to be higher.- Amplify bacterial cultures of commercially available XL1 Blue bacterial agar stab (containing Cas9 plasmid) using an appropriate antibiotic such as ampicillin. Use 675 µL of liquid culture (with 325 µL of glycerol) to create a backup glycerol stock for long-term storage at −80 °C.
- Use the remainder of the liquid culture for a miniprep purification, according to the protocol provided with the miniprep kit. Resuspend the final purified DNA in 50 µL of the provided elution buffer. Quantify the product via a spectrophotometer to examine the yield and purity.
- Linearize 2 µg of the purified DNA via restriction digest using an appropriate restriction enzyme and using the appropriate buffer and incubation time as listed for the enzyme of interest.
- Purify the linearized plasmid using a PCR Purification Kit, resuspending it in 30 µL of the provided elution buffer.
- After quantifying the product, use this as a template for in vitro transcription using the appropriate transcription kit for the promoter of interest. Follow the provided protocol and purify via lithium chloride precipitation27. Resuspend the purified RNA in 10 µL of nuclease-free H2O and quantify it before storing at −20 °C for use in the microinjection mix.
- Prepare the two sgRNA guides.
- Prepare the sgRNA plasmid with a scaffold by amplifying bacterial cultures from the commercially available XL1 Blue bacterial agar stab (see Table of Materials for details) in the same way as MLM3613 above (step 2.1.1), except use kanamycin instead of ampicillin.
- Anneal the two pairs of complementary single-stranded oligonucleotides (ssODNs) for the sgRNA guides designed above by first resuspending the ssODNs in 1x annealing buffer to a concentration of 100 µM.
- In separate reactions for each of the two sgRNAs, anneal the pair of complementary ssODNs using a thermal cycler. Mix 2 µg of each complementary ssODN pair with 50 µL of annealing buffer and incubate at 95 °C for 2 min, then cool to 25 °C over 45 min.
- Digest a commercially available plasmid that contains a gRNA scaffold. Digest 2 µg of the DR274 plasmid using 1 µL of BsaI, 2 µL of appropriate buffer, and ddH2O to 20 µL at 37 °C for 1 h. Confirm linearization (optional: purify using a PCR Purification Kit) using gel electrophoresis28.
- In two separate ligation reactions (one for each sgRNA), ligate the annealed ssODNs with the linearized DR274 plasmid. Use a molar insert:vector ratio of 3:1, calculating the appropriate mass through an online ligation calculator. Mix the required mass of the insert and vector with 1 µL of T4 DNA ligase, 2 µL of ligation buffer, and ddH2O to 12 µL and incubate at room temperature for 12 h.
- Transform 2 µL of the ligated product into appropriate competent cells (such as 10β cells) using standard approaches, and then amplify and purify the product using a commercially available Miniprep Kit. Optional: create a glycerol stock of this product.
- Transcribe the two sgRNAs by linearizing 2 µg of each guide using a 3' downstream restriction site that is as close to the end of the space sequence as possible. For the DR274 plasmid, linearize with HindIII, and then purify the RNA template using a PCR Purification Kit, resuspending in 30 µL of the elution buffer.
- Transcribe the two guides using an RNA transcription kit. Follow the manufacturer's protocol and purify via lithium chloride precipitation27. Resuspend the two purified sgRNA guides in 10 µL of nuclease-free H2O, quantify, and store at −20 °C for use in the microinjection mix.
NOTE: The RNA transcription kit cannot incorporate a 5' cap or poly-A tail.
- Prepare the double-stranded, exogenous HDR reporter template.
NOTE: The template is synthesized in two parts, one upstream and one downstream of the mVenus YFP reporter gene. These two segments are synthetic constructs ordered through a commercial provider and then ligated into the pKHR5 (which contains mVenus YFP) plasmid sequentially.- Prepare the pKHR5 plasmid by amplifying bacterial cultures from the commercially available DH5α bacterial strain (see Table of Materials for details) in the same way as MLM3613 above (step 2.1.1).
NOTE: The pKHR5 plasmid contains the mVenus YFP reporter gene sequence. - Resuspend the two template segments designed above to 100 µM in TE buffer and then transform into 10β cells.
- Digest the first template segment (the one upstream of the mVenus YFP reporter gene) and the pKHR5 plasmid using the restriction enzymes selected in step 1.3.3.
NOTE: Sequential digest of pKHR5 is necessary due to the close proximity of the selected restriction sites in the MCS. - To prepare the pKHR5 plasmid for the upstream template segment, digest 4 µg of pKHR5 (as per step 2.2.4) with SalI and then purify using a PCR purification kit. Resuspend in 30 µL of ddH2O and use this as a template for the second digestion reaction with XhoI. Purify the product using the PCR Purification Kit.
- Prepare the first template segment by digesting 2 µg of the template segment (step 2.3.2), 1 µL of XhoI, 1 µL of SalI, 2 µL of appropriate buffer, and ddH2O to 20 µL for 1 h at 37 °C and gel-purify the product.
- Ligate the upstream template segment from step 2.3.5 into the prepared pKHR5 using the reaction conditions described in step 2.2.5. Transform the ligated product into competent 10β cells, amplify, and purify using a miniprep (optional: create a glycerol stock of this product).
- Use the ligated product from step 2.3.6 (which contains the first template segment ligated into pKHR5 plasmid upstream of the mVenus YFP reporter gene) and digest to prepare for the second (downstream of the mVenus reporter) template segment. Digest 4 µg of the ligated product (from step 2.3.6, as in step 2.2.4) with BamHI, and then purify using a PCR purification kit. Resuspend in 30 µL of ddH2O and use this as the template for the second digest reaction with EcoRI. Purify the product using the PCR Purification Kit.
- Prepare the second template segment by digesting 2 µg of the template segment (step 2.3.2), 1 µL of BamHI, 1 µL of EcoRI, 2 µL of appropriate buffer, and ddH2O to 20 µL for 1 h at 37 °C and gel-purify the product.
- Ligate the downstream template segment into the prepared pKHR5 (from step 2.3.7) using the reaction conditions described in step 2.2.5. Transform the ligated product into competent cells, amplify, and purify using a miniprep kit. Create a glycerol stock of this final product, which contains both template segments ligated either side of the mVenus YFP reporter gene within pKHR5.
- Prepare the pKHR5 plasmid by amplifying bacterial cultures from the commercially available DH5α bacterial strain (see Table of Materials for details) in the same way as MLM3613 above (step 2.1.1).
- Prepare the microinjection mix using the Cas9 mRNA, two sgRNAs, and the exogenous HDR reporter template.
- Mix 200 ng/µL of Cas9 mRNA, 100 ng/µL of each sgRNA, and 200 ng/µL of exogenous HDR reporter template in 1x injection buffer to a final volume of 20 µL.
- Store the microinjection mix at −20 °C and discard the unused mix after three freeze-thaw cycles.
NOTE: Use 4 nL of this microinjection mix for microinjection into the yolk sac of each embryo.
3. Breeding of zebrafish and embryo microinjection
NOTE: Protocols for zebrafish breeding and the microinjection of single-cell embryos have been described previously29,30,31.
- For breeding, use zebrafish of the AB strain and that are 6-12 months of age. Inject the embryos in the one-cell stage at approximately 40 min post fertilization (see Figure 5).
NOTE: The biological sex of the injected embryos was not known; sexual dimorphism is not evident until approximately 3 months of age32.
4. Reporter gene screening of CRISPR-Cas9-edited larval zebrafish
- Visualize YFP integration in zebrafish larvae following the microinjection of CRISPR-Cas9 components to screen for successful HDR edits.
- In a 25 mm Petri dish, anesthetize 24 zebrafish larvae at 3 days post fertilization (dpf) in 0.3% tricaine methane sulfonate (MS-222, buffered to pH 7.0-7.4 with HEPES and sodium hydroxide) until they lose their self-righting reflex (typically 1-2 min). Once anesthetized, transfer each larva into an individual well of a 24-well plate.
- Using a microscope capable of detecting GFP/YFP, screen for reporter gene fluorescence in the eyes of each individual larva.
- Capture images of each larva and document the presence or absence of reporter gene expression.
5. Phenotyping of CRISPR-Cas9-edited larval zebrafish
- Following reporter gene screening, perform cardiac phenotyping (heart rate, pericardial dimensions, ECG) on each larva. Phenotype an equal number of reporter gene-positive and -negative larvae.
- Use a CCD camera (e.g., blackfly USB3) and video and image recording software (e.g., Micromanager for ImageJ) to measure heart rate and pericardial dimensions while the larvae are anesthetized.
- To measure the heart rate, using Micromanager, create a region of interest (ROI) so as to capture the heart and exclude other structures.
- Import the video into ImageJ as an image sequence, and ensure the correct number of frames is entered under number of images.
- After the file is open, use the rectangle selection tool to draw an ROI within the heart but excluding other moving elements, and save the ROI in the ROI manager (analyze | tools | ROI manager).
- Click plugins | install, and select the heart rate algorithm to install the code in the default folder, then select the plugin at the bottom of the plugins tab. Record the beats per minute (bpm) from the pop-up window.
NOTE: Image detection algorithms were custom-written to detect heart rate by measuring individual pixel density changes associated with ventricular systolic contraction. The code can be found at https://github.com/dpoburko/zFish_HR.
- Measure pericardial dimensions using a free tool such as ImageJ to free-draw ROIs around the pericardial sac and one of the eyes. Open the image in ImageJ and use the polygon selection tool to draw an ROI first around the pericardial sac, saving it in the ROI manager as in step 5.1.2, and repeat for the eye. Select these two ROIs in the ROI manager, then click measure. Record the area for each to later calculate the area of the pericardial sac normalized to the eye area in each larva.
- Following heart rate and pericardial measurements, record ECG from individual larvae.
NOTE: Protocols for recording zebrafish ECG have been described previously33,34,35,36.
6. Genotyping of CRISPR-Cas9-edited larval zebrafish
- Following phenotypic analyses, conduct on- and potential off-target genotyping to confirm accurate and precise HDR gene editing.
- Anesthetize each 3 dpf larva in 0.3% MS-222 and tail clip to isolate gDNA using the HOTShot method37. Incubate each excised tail clip in 15 µL of 25 mM NaOH at 95 °C for 20 min. Then, neutralize with 1.5 µL of Tris-HCl and centrifuge at 13,800 x g for 30 s. Retain the supernatant, which contains extracted gDNA.
- Recover the larvae in E3 media and return them to the housing system if further development or study is intended.
- Using the extracted gDNA as a template, perform PCR-based Sanger sequencing of on-target and potential off-target sites.
NOTE: Optional: a nested PCR approach may be beneficial for some gene regions. - Ensure that the on-target primer design captures the mutation site and the closest sgRNA binding site. Design a separate sequencing primer to detect the transition from the inserted homology arm and the target gene to confirm integration into the gene of interest. Design primers to sequence the top three potential off-target sites identified in step 1.2.3.
NOTE: Guide-design software programs often suggest primers to use, but customization may be necessary to achieve optimal results. - Compile on- and off-target genotyping, heart rate, pericardial dimensions, ECG phenotyping, and reporter gene data identifiable for each zebrafish.
Representative Results
The successful use of this two-sgRNA exon replacement CRISPR approach is highlighted by the introduction and simple detection of a precise edit to engineer the LQTS-associated variant, R56Q, in the zkcnh6a gene in zebrafish. Figure 6 shows a representative 3 dpf larvae injected at the one-cell embryo stage with CRISPR components as described above. Figure 6A shows the presence of the YFP mVenus reporter gene expression in the eye lens as a positive reporter of successful template integration. Figure 6B,C show Sanger sequencing chromatograms obtained from genomic DNA isolated from tail clip samples of wild-type and reporter gene-positive fish, respectively. Reporter gene-positive fish were found to have the precise edit, G to A, which introduces the R56Q variant into zkcnh6a. Genotyping showed a 100% correlation between YFP reporter gene expression and the presence of the precise R56Q gene edit, validating this fluorescence screening tool.
Phenotyping of gene-edited zebrafish larvae was conducted at 3 dpf. Figure 7 shows representative results from wild-type and R56Q gene-edited larvae. Heart rate was detected by video capture as described above. An example of the measurement of pericardial dimensions as a ratio of eye area is shown (Figure 7A). Figure 7B plots heart rate against normalized pericardial dimensions, highlighting a trend of bradycardia with increasing pericardial edema, which is associated with disorders of cardiac repolarization in zebrafish8,38,39,40. Figure 7C shows a representative example of ECG recordings from 3 dpf larvae. Standard intervals (QT, QRS) were measured from averaged ECG signals.
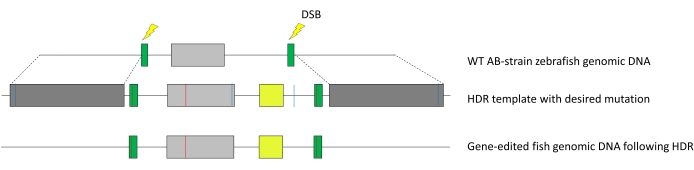
Figure 1: Integration of HDR template into the zebrafish genome. Dark grey, homology arms; green, sgRNA guide targets with silent mutation to prevent Cas9 recutting; light grey, target exon of interest; red line, point mutation; yellow, mVenus YFP reporter gene under an α-crystallin promoter; dashed lines indicate homology. Here, the targeted precise edit was R56Q in exon 2 of the zkcnh6a gene. Abbreviations: HDR = homology-directed repair; sgRNA = single-guide RNA; YFP = yellow fluorescent protein; DSB = double-stranded break; WT = wild type. Please click here to view a larger version of this figure.
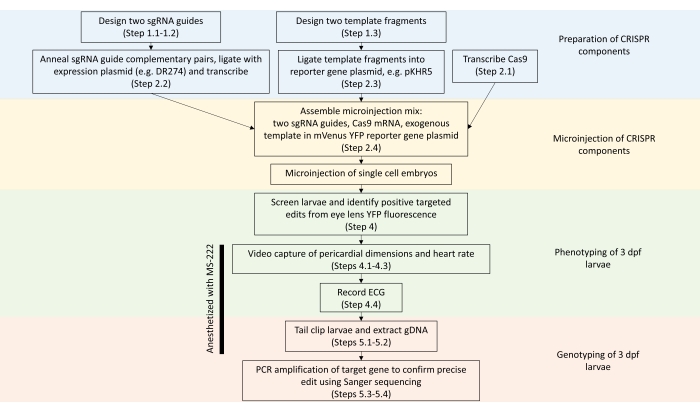
Figure 2: Summary of steps to engineer precise edits in zebrafish genes using the two-sgRNA CRISPR-Cas9 approach (related protocol step numbers are indicated in parentheses). Abbreviations: sgRNA = single-guide RNA; YFP = yellow fluorescent protein; gDNA = genomic DNA; ECG = electrocardiogram; dpf = days post fertilization; MS-222 = tricaine methane sulfonate. Please click here to view a larger version of this figure.
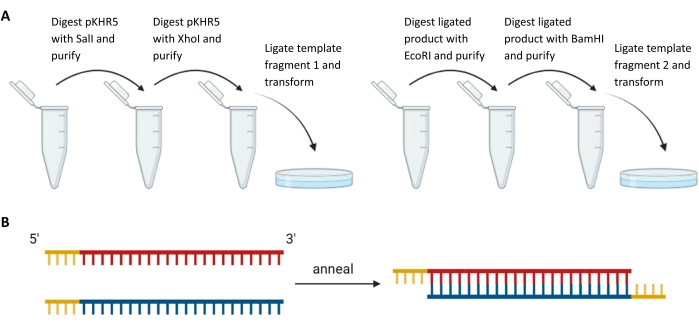
Figure 3: Preparation of exogenous template fragments and sgRNA guides. (A) Sequential digestion and ligation of template fragments upstream and downstream of the mVenus YFP reporter gene sequence in pKHR5. (B) Annealing of complementary sgRNA pairs with restriction overhang for ligation into DR274. Abbreviations: sgRNA = single-guide RNA; YFP = yellow fluorescent protein. Please click here to view a larger version of this figure.
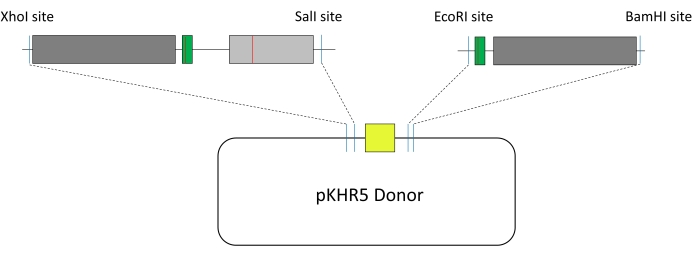
Figure 4: Construction of HDR template. Dark grey, homology arms; green, sgRNA guide targets with silent mutation to prevent Cas9 recutting; light grey, target exon of interest; red line, point mutation; yellow, mVenus YFP reporter gene under an α-crystallin promoter; dark blue line, added restriction sites. The two template fragments are integrated into the pKHR5 plasmid donor. Abbreviations: HDR = homology-directed repair; sgRNA = single-guide RNA. Please click here to view a larger version of this figure.
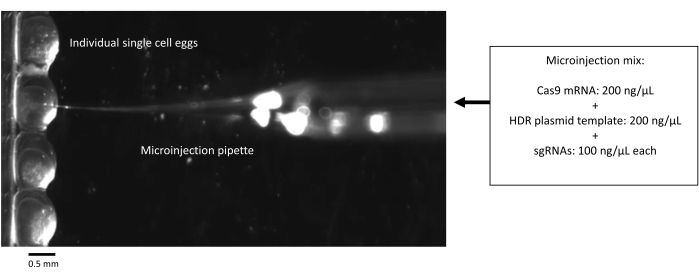

Figure 5: Microinjection of single-cell zebrafish embryos with CRISPR-Cas9 components. Scale bar = 0.5 mm. Abbreviations: HDR = homology-directed repair; sgRNA = single-guide RNA. Please click here to view a larger version of this figure.
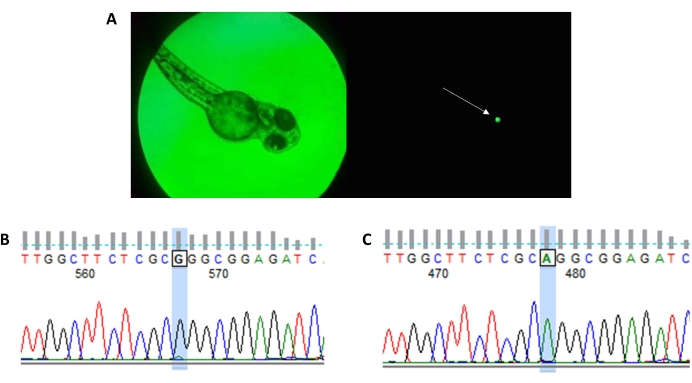
Figure 6: Easy detection of mVenus YFP reporter gene fluorescence indicates positive HDR exogenous template integration into the target gene. (A) Example of mVenus YFP expression in a zebrafish eye (arrow) in an edited zebrafish larva. (B) Successful edits are confirmed by sequencing chromatograms (left, WT; right, R56Q edit). Please click here to view a larger version of this figure.
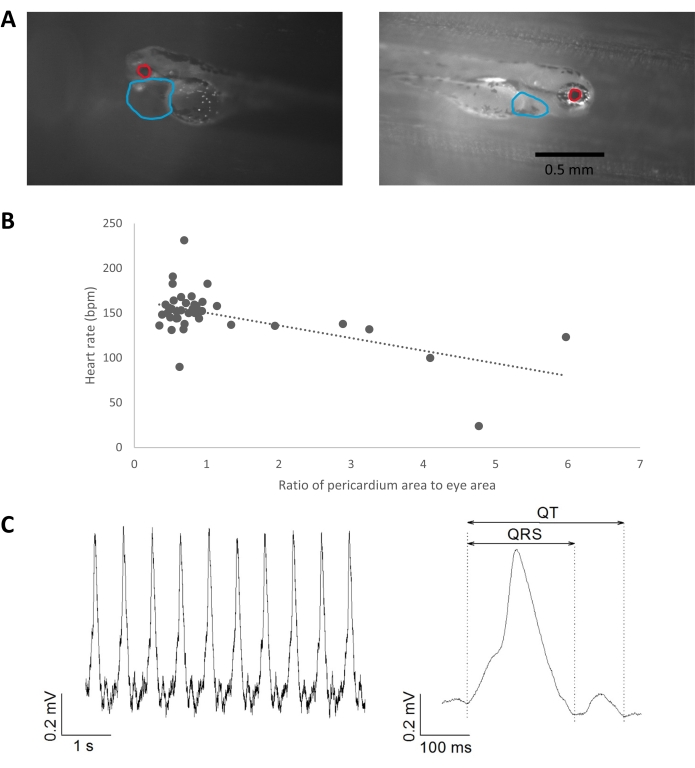
Figure 7: Phenotypic analysis of cardiac consequences in 3 dpf zebrafish following the precise R56Q edit in the zkcnh6a target gene. (A) Image detection of pericardial dimensions relative to eye size using the polygon tool in ImageJ. The boundaries of the pericardial sac were marked by the user from a single recording frame based on changes in translucency and pigmentation. Examples of normal pericardial dimensions and pericardial effusion are shown. Scale bar = 0.5 mm. (B) Correlation between pericardial dimensions (relative to eye dimension) and heart rate, R2 = 0.33. (C) Example of ECG recording from a 3 dpf zebrafish larva heart (left) and averaged complexes (right). Heart rate, 131 bpm; heart rate-corrected QTc interval, 460 ms. Abbreviations: dpf = days post fertilization; ECG = electrocardiogram. Please click here to view a larger version of this figure.
Discussion
The engineering of precise gene edits using CRISPR-Cas9 is challenged by the low efficiencies of HDR mechanisms and their efficient detection. Here, a CRISPR-Cas9-based two-sgRNA exon replacement approach is described that produces precise edits in zebrafish with straightforward visual detection of positive edits. The efficacy of this approach is demonstrated by generating precise edits in the zkcnh6a gene. This paper shows how cardiac function in gene-edited zebrafish larvae may be assessed using non-invasive phenotypic measures of heart rate, pericardial dimensions, and ECG morphology. This approach, from introducing a gene edit to phenotypic evaluation, can be completed from start to finish within approximately 1 week.
The benefits of the above editing and phenotyping approach are the ease of CRISPR modification design, the wide applicability in multiple physiological systems, the ability to insert large genes or gene fragments, and the ability to track variant effects longitudinally through development and generations. The success of precise edits in this approach may be related to the combination of the large template size (due to the reporter gene insert and long homology arms), which has been shown to increase the efficiency of edits in zebrafish14, and the two-sgRNA-guides strategy, which has been used effectively in zebrafish TALEN-induced edits8.
One particular strength of the described approach is the ability to insert large genes or gene fragments. This may be useful, for example, to insert human orthologs41, allowing for more clinically translatable characterization and comparison between orthologs. Alternatively, genes encoding Cas enzymes could also be inserted, allowing for a line of zebrafish with in vivo CRISPR editing mechanisms, providing an inducible system. Similarly, alternative CRISPR mechanisms, such as prime editing, could be integrated and result in a line of zebrafish that are readily edited precisely and efficiently.
Despite the advantages of this approach, there are some limitations. First, only a single gene and locus have been modified, and further testing at other sites or in other genes is necessary to evaluate how broadly applicable this approach is. Due to the long homology arms required, the template design costs are higher; however, this may be offset by efficient screening. Another limitation is that the screening approach requires fluorescence detection capability. However, optical requirements are relatively low and can be custom-built or commercially purchased at a reasonably low cost. Using a two-sgRNA approach increases the number of potential off-target events; however, this is likely mitigated by the lower probability that the two sgRNA guides will both anneal in a manner that facilitates the incorporation of the template to yield reporter gene expression. Finally, using Cas9 mRNA may lead to mosaicisms as the Cas9 is not active until later developmental stages. This could be accounted for by sequencing particular tissue types; however, given the size of the zebrafish larvae, this is technically challenging.
In summary, this CRISPR-Cas9 two-sgRNA precise editing approach in zebrafish enables the simple visual detection of positive edits and may be adapted to incorporate large genes of interest at any locus. Combined with phenotypic measures, this allows for a reliable and high throughput platform for studying clinically-relevant cardiac variants.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This research was supported by a Canadian Institutes of Health Research Project grant (T.W.C.) and Natural Sciences and Engineering Research Council of Canada Discovery grants (T.W.C.).
Materials
| Program | |||
| CRISPOR | TEFOR Infrastructure | ||
| ENSEMBL | European Bioinformatics Institute | ||
| ImageJ | National Institutes of Health (NIH) | ||
| Micro-Manager | Open Source (Github) | ||
| NEBiocalculator | New England Biolabs (NEB) | ||
| EQUIPMENT | |||
| 24-well Plate | VWR | ||
| 25 mm Petri Dish | VWR | ||
| Blackfly USB3 Camera | Teledyne FLIR | ||
| C1000 Thermal Cycler | Bio-Rad | ||
| Centrifuge 5415C | Eppendorf | ||
| EZNA Gel Extraction Kit | Omega Biotek | ||
| MAXIscript T7 Transcription Kit | Invitrogen | ||
| MaxQ 5000 Incubator | Barnstead Lab Line | ||
| Miniprep Kit | Qiagen | ||
| mMessage mMachine T7 Ultra Transcription Kit | Invitrogen | ||
| ND1000 Spectrophotometer | Nanodrop | ||
| PCR Purification Kit | Qiagen | ||
| PLI 100A Picoinjector | Harvard Apparatus | ||
| PowerPac Basic Power Supply | Bio-Rad | ||
| Stemi 305 Steroscope | Zeiss | ||
| Wide Mini Sub Cell GT Electrophoresis System | Bio-Rad | ||
| ZebTec Zebrafish Housing System | Tecniplast | ||
| SERVICES | |||
| Gene Synthesis | Genewiz | ||
| Sanger Sequencing | Genewiz | ||
| REAGENTS | |||
| 10β Competent Cells | NEB | ||
| 10X PCR Buffer | Qiagen | ||
| 100 mM Nucleotide Mixture | ABM | ||
| Ampicillin | Sigma | ||
| BamHI Endonuclease w/ buffer | NEB | ||
| BsaI Endonuclease w/ buffer | NEB | ||
| DR274 Plasmid (XL1 Blue bacterial agar stab) | Addgene | ||
| EcoRI Endonuclease w/ buffer | NEB | ||
| Glycerol | |||
| HEPES | Sigma | ||
| HindIII Endonuclease w/ buffer | NEB | ||
| Kanamycin | Sigma | ||
| Methylene Blue | Sigma | ||
| MLM3613 Plasmid (XL1 Blue bacterial agar stab) | Addgene | ||
| MS-222 (Tricaine) | Sigma | ||
| pKHR5 Plasmid (DH5α bacterial agar stab) | Addgene | ||
| PmeI Endonuclease w/ buffer | NEB | ||
| SalI Endonuclease w/ buffer | NEB | ||
| Sodium Hydroxide | Sigma | ||
| T4 Ligase w/ buffer | Sigma | ||
| Taq Polymerase | Qiagen | ||
| TE Buffer | Sigma | ||
| Tris Hydrochloride | Sigma | ||
| XhoI Endonuclease w/ buffer | NEB | ||
| RECIPES | |||
| Solution | Component | Supplier | |
| Annealing Buffer (pH 7.5-8.0) | 10 mM Tris | Sigma | |
| 50 mM NaCl | Sigma | ||
| 1 mM EDTA | Sigma | ||
| E3 Media (pH 7.2) | 5 mM NaCl | Sigma | |
| 0.17 mM KCl | Sigma | ||
| 0.33 mM CaCl2 | Sigma | ||
| 0.33 mM MgSO4 | Sigma | ||
| Injection Buffer (pH 7.5) | 20 mM HEPES | Sigma | |
| 150 mM KCl | Sigma |
References
- Zarei, A., Razban, V., Hosseini, S. E., Tabei, S. M. B. Creating cell and animal models of human disease by genome editing using CRISPR/Cas9. The Journal of Gene Medicine. 21 (4), 3082 (2019).
- Lee, H., Yoon, D. E., Kim, K. Genome editing methods in animal models. Animal Cells and Systems. 24 (1), 8-16 (2020).
- Li, Q., et al. Applications of genome editing technology in animal disease modeling and gene therapy. Computational and Structural Biotechnology Journal. 17, 689-698 (2019).
- Gut, P., Reischauer, S., Stainier, D. Y. R., Arnaout, R. Little fish, big data: Zebrafish as a model for cardiovascular and metabolic disease. Physiological Reviews. 97 (3), 889-938 (2017).
- Kegel, L., et al. Forward genetic screen using zebrafish to identify new genes involved in myelination. Oligodendrocytes: Methods and Protocols. 1936, 185-209 (2019).
- Shah, A. N., Davey, C. F., Whitebirch, A. C., Miller, A. C., Moens, C. B. Rapid reverse genetic screening using CRISPR in zebrafish. Nature Methods. 12 (6), 535-540 (2015).
- González-Rosa, J. M. Zebrafish models of cardiac disease: From fortuitous mutants to precision medicine. Circulation Research. 130 (12), 1803-1826 (2022).
- Hoshijima, K., Jurynec, M. J., Grunwald, D. J. Precise editing of the Zebrafish genome made simple and efficient. Developmental Cell. 36 (6), 654-667 (2016).
- Albadri, S., Del Bene, F., Revenu, C. Genome editing using CRISPR/Cas9-based knock-in approaches in zebrafish. Methods. 121-122, 77-85 (2017).
- Armstrong, G. A. B., et al. Homology directed knockin of point mutations in the zebrafish tardbp and fus genes in ALS using the CRISPR/Cas9 system. PLoS One. 11 (3), 0150188 (2016).
- Bai, H., et al. CRISPR/Cas9-mediated precise genome modification by a long ssDNA template in zebrafish. BMC Genomics. 21 (1), 67 (2020).
- de Vrieze, E., et al. Efficient generation of knock-in zebrafish models for inherited disorders using CRISPR-Cas9 ribonucleoprotein complexes. International Journal of Molecular Sciences. 22 (17), 9429 (2021).
- Eschstruth, A., Schneider-Maunoury, S., Giudicelli, F. Creation of zebrafish knock-in reporter lines in the nefma gene by Cas9-mediated homologous recombination. Genesis. 58 (1), 23340 (2020).
- Irion, U., Krauss, J., Nüsslein-Volhard, C. Precise and efficient genome editing in zebrafish using the CRISPR/Cas9 system. Development. 141 (24), 4827-4830 (2014).
- Kimura, Y., Hisano, Y., Kawahara, A., Higashijima, S. Efficient generation of knock-in transgenic zebrafish carrying reporter/driver genes by CRISPR/Cas9-mediated genome engineering. Scientific Reports. 4 (1), 6545 (2014).
- Levic, D. S., Yamaguchi, N., Wang, S., Knaut, H., Bagnat, M. Knock-in tagging in zebrafish facilitated by insertion into non-coding regions. Development. 148 (19), (2021).
- Prykhozhij, S. V., et al. Optimized knock-in of point mutations in zebrafish using CRISPR/Cas9. Nucleic Acids Research. 46 (17), 102 (2018).
- Wierson, W. A., et al. Efficient targeted integration directed by short homology in zebrafish and mammalian cells. eLife. 9, 53968 (2020).
- Boel, A., et al. CRISPR/Cas9-mediated homology-directed repair by ssODNs in zebrafish induces complex mutational patterns resulting from genomic integration of repair-template fragments. Disease Models & Mechanisms. 11 (10), (2018).
- Tessadori, F., et al. Effective CRISPR/Cas9-based nucleotide editing in zebrafish to model human genetic cardiovascular disorders. Disease Models & Mechanisms. 11 (10), (2018).
- Zhang, Y., Zhang, Z., Ge, W. An efficient platform for generating somatic point mutations with germline transmission in the zebrafish by CRISPR/Cas9-mediated gene editing. The Journal of Biological Chemistry. 293 (17), 6611-6622 (2018).
- Vornanen, M., Hassinen, M. Zebrafish heart as a model for human cardiac electrophysiology. Channels. 10 (2), 101-110 (2016).
- Nemtsas, P., Wettwer, E., Christ, T., Weidinger, G., Ravens, U. Adult zebrafish heart as a model for human heart? An electrophysiological study. Journal of Molecular and Cellular Cardiology. 48 (1), 161-171 (2010).
- Haeussler, M., et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biology. 17 (1), 148 (2016).
- Liang, X., Potter, J., Kumar, S., Ravinder, N., Chesnut, J. D. Enhanced CRISPR/Cas9-mediated precise genome editing by improved design and delivery of gRNA, Cas9 nuclease, and donor DNA. Journal of Biotechnology. 241, 136-146 (2017).
- Richardson, C. D., Ray, G. J., DeWitt, M. A., Curie, G. L., Corn, J. E. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nature Biotechnology. 34 (3), 339-344 (2016).
- Sambrook, J., Fritsch, E. F., Maniatis, T. Precipitation of Large RNAs with Lithium Chloride. Molecular Cloning: A Laboratory Manual, Book 3. E.15. , (1989).
- Sambrook, J., Fritsch, E. F., Maniatis, T. Agarose Gel Electrophoresis. Molecular Cloning: A Laboratory Manual, Book 1. , 3-20 (1989).
- Sorlien, E. L., Witucki, M. A., Ogas, J. Efficient production and identification of CRISPR/Cas9-generated gene knockouts in the model system Danio rerio. Journal of Visualized Experiments. (138), e56969 (2018).
- Avdesh, A., et al. Regular care and maintenance of a zebrafish (Danio rerio) laboratory: An introduction. Journal of Visualized Experiments. (69), e4196 (2012).
- Rosen, J. N., Sweeney, M. F., Mably, J. D. Microinjection of zebrafish embryos to analyze gene function. Journal of Visualized Experiments. (25), e1115 (2009).
- Kossack, M. E., Draper, B. W. Genetic regulation of sex determination and maintenance in zebrafish (Danio rerio). Current Topics in Developmental Biology. 134, 119-149 (2019).
- Tanaka, Y., et al. Functional analysis of KCNH2 gene mutations of type 2 long QT syndrome in larval zebrafish using microscopy and electrocardiography. Heart and Vessels. 34 (1), 159-166 (2019).
- Dhillon, S. S., et al. Optimisation of embryonic and larval ECG measurement in zebrafish for quantifying the effect of QT prolonging drugs. PLoS One. 8 (4), 60552 (2013).
- Yu, F., et al. Evolving cardiac conduction phenotypes in developing zebrafish larvae: Implications to drug sensitivity. Zebrafish. 7 (4), 325-331 (2010).
- Hurst, R. M. . Development and optimization of tools for embryonic electrocardiograph recording for heart dysfunction in zebrafish. , (2018).
- Meeker, N. D., Hutchinson, S. A., Ho, L., Trede, N. S. Method for isolation of PCR-ready genomic DNA from zebrafish tissues. BioTechniques. 43 (5), 610-614 (2007).
- Arnaout, R., et al. Zebrafish model for human long QT syndrome. Proceedings of the National Academy of Sciences of the United States of America. 104 (27), 11316-11321 (2007).
- Milan, D. J., Peterson, T. A., Ruskin, J. N., Peterson, R. T., MacRae, C. A. Drugs that induce repolarization abnormalities cause bradycardia in zebrafish. Circulation. 107 (10), 1355-1358 (2003).
- Langheinrich, U., Vacun, G., Wagner, T. Zebrafish embryos express an orthologue of HERG and are sensitive toward a range of QT-prolonging drugs inducing severe arrhythmia. Toxicology and Applied Pharmacology. 193 (3), 370-382 (2003).
- MacRae, C. A. Cardiac arrhythmia: In vivo screening in the zebrafish to overcome complexity in drug discovery. Expert Opinion on Drug Discovery. 5 (7), 619-632 (2010).

