Patterning Bioactive Proteins or Peptides on Hydrogel Using Photochemistry for Biological Applications
Summary
In this method, we use photopolymerization and click chemistry techniques to create protein or peptide patterns on the surface of polyethylene glycol (PEG) hydrogels, providing immobilized bioactive signals for the study of cellular responses in vitro.
Abstract
There are many biological stimuli that can influence cell behavior and stem cell differentiation. General cell culture approaches rely on soluble factors within the medium to control cell behavior. However, soluble additions cannot mimic certain signaling motifs, such as matrix-bound growth factors, cell-cell signaling, and spatial biochemical cues, which are common influences on cells. Furthermore, biophysical properties of the matrix, such as substrate stiffness, play important roles in cell fate, which is not easily manipulated using conventional cell culturing practices. In this method, we describe a straightforward protocol to provide patterned bioactive proteins on synthetic polyethylene glycol (PEG) hydrogels using photochemistry. This platform allows for the independent control of substrate stiffness and spatial biochemical cues. These hydrogels can achieve a large range of physiologically relevant stiffness values. Additionally, the surfaces of these hydrogels can be photopatterned with bioactive peptides or proteins via thiol-ene click chemistry reactions. These methods have been optimized to retain protein function after surface immobilization. This is a versatile protocol that can be applied to any protein or peptide of interest to create a variety of patterns. Finally, cells seeded onto the surfaces of these bioactive hydrogels can be monitored over time as they respond to spatially specific signals.
Introduction
There are many stimuli that influence cell behavior. Generally, typical cell culturing techniques rely on soluble factors to elicit cellular responses; however, there are limitations to this approach. These methods are unable to accurately display all signaling motifs commonly found in vivo. Such signaling mechanisms include sequestered growth factors, cell-cell signaling, and spatially-specific biochemical cues.Furthermore, substrate stiffness can play an important role in cell behavior and stem cell differentiation and is not easily manipulated using common cell culturing practices1,2. Biomaterial approaches offer a new platform to begin exploring these signaling mechanisms. In particular, hydrogels are excellent candidates for tuning substrate stiffness3,4, immobilizing proteins and peptides5,6, and creating spatially specific patterns7,8.
Hydrogels are commonly used as scaffolds in tissue engineering due to their biophysical and biochemical commonalities with the extracellular matrix (ECM)9,10. Natural polymers are common choices for scaffolds, as they are biocompatible and are found in many tissues of the body. The limitation of using natural polymers as substrates is that they lack easily manipulated chemical moieties for bioconjugation. On the other hand, synthetic hydrogels, as such as PEG, are excellent platforms for targeted chemistries11,12. Additionally, PEG hydrogels do not elicit a cellular response and therefore are used as inert backbones for creating bioactive scaffolds.
To create bioactive hydrogels, both photopolymerization and thiol-ene click chemistry reactions are employed. These photoreactions require a photoinitiator and a UV light source. When photoinitiators are introduced to UV light, bonds break to form radicals. Theses radicals are necessary for initiating the reaction but can negatively affect protein bioactivity12,13. Therefore, it is crucial to optimize photoinitiator and UV exposure times to maintain protein bioactivity.
In this method, hydrogels are synthesized through acrylate-acrylate chain growth photopolymerization. PEG-diacrylate (PEGDA) monomers react with each other to form branched polymer networks responsible for the structure of the hydrogel. The concentration of PEGDA monomers within the gel precursor solution will control the substrate stiffness. Due to the small pore size of the hydrogel, ECM proteins such as fibronectin can be easily incorporated within the hydrogel for the purpose of cell attachment. Finally, these hydrogels can be surface-patterned with bioactive peptides or proteins via thiol-ene click chemistry reactions. Here, unreacted free acrylates within the hydrogel system will react with free thiols located on the protein or peptide when exposed to UV light. After the proteins or peptides are immobilized on the hydrogel surface, the hydrogel can be stored at 4 °C for several weeks without losing bioactivity. This offers convenience, flexible experimental planning, and the possibility for collaboration between labs. Overall, this platform allows for biomechanical and spatial biochemical control, independent of each other, for the opportunity to influence cellular behavior.
Protocol
1. Preparation of Materials for Hydrogel Synthesis
- Prepare stock solutions of PEGDA, lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP), and fibronectin under sterile conditions and based on calculations (Table 1A).
- Weigh out and dissolve compounds in phosphate-buffered saline (PBS). Typically, maintain PEGDA working solution concentrations between 50 and 200 mg/mL (5-20% weight/volume). Pipette the PEGDA solution through a 0.22-µm syringe filter for sterilization.
NOTE: Keep the PEGDA solutions covered with a foil wrapping to protect them from light. Prepare a fresh solution of PEGDA (recommended) for each experiment. - Reconstitute fibronectin protein powder based on the manufacturer's recommendation. Maintain the fibronectin stock solution at 1 mg/mL, aliquot it into 60 to 100 µL aliquots, and store them at -20 °C until use. Defrost the aliquots at 4 °C for several hours or overnight before use. If protein stock is not already provided in sterile conditions, use a 0.22 µm syringe filter for sterilization.
NOTE: Fibronectin is used for cell attachment. Other ECM proteins can be substituted. - Weigh out LAP for the stock solution and dissolve it in PBS; a typical stock concentration is 25 mg/mL. Sonicate if there is any trouble with dissolving. Pass the solution through a 0.22 µm filter for sterilization. Cover the LAP with a foil wrapping and keep it at 4 °C for up to several months.
NOTE: LAP is the photoinitiator used for the photochemistry.
- Weigh out and dissolve compounds in phosphate-buffered saline (PBS). Typically, maintain PEGDA working solution concentrations between 50 and 200 mg/mL (5-20% weight/volume). Pipette the PEGDA solution through a 0.22-µm syringe filter for sterilization.
- Prepare sterile glass slide molds for hydrogel formation.
- Autoclave a polyester sheet; one is needed for each gel mold. Soak two glass slides and three plastic spacers (0.5 mm thick) for each gel mold in 70% ethanol for at least 2 h, but generally allow them to soak overnight. Soak five binder clips for each gel mold in 70% ethanol for 10 min prior to use.
- Place the glass slides, spacers, and binder clips onto a small autoclaved sheet in the cell culture hood to allow them to air dry for several hours.
- Prepare hydrogel molds by placing plastic spacers around the edge of a glass slide. Place the second glass slide on top. Secure the spacers tightly next to each other with binder clips, two on each long side and one on top.
NOTE: If this is not properly assembled, the gel solution will leak out. - Surface-sterilize the mold with cell culture hood UV light for 30 min prior to use. Halfway through, flip the mold to expose both surfaces.
2. Modifying Proteins with a Free Thiol
- Prepare stock solutions using calculations from the spreadsheet for converting amine into thiol on proteins (Table 1A).
- To make the reaction buffer, adjust the PBS to pH 8 and add 5 mM EDTA. Pipette reaction buffer into a 0.22 µm syringe filter for sterilization.
NOTE: EDTA is important, as it protects thiols from forming disulfide bonds. Thiol groups must remain in their reduced form for the reaction to occur. - Weigh out 2-iminothiolane (Traut's Reagent) and dissolve it in reaction buffer for a stock solution concentration of 2 mg/mL (14 mM). Pass the solution through a 0.22-µm syringe filter for sterilization. Store the stock solution at 4 °C for up to several months.
NOTE: 2-iminothiolane is the molecule that reacts with solvent-exposed free amines on the proteins and converts them into free thiols. - Reconstitute the protein in reaction buffer at a concentration between 0.1 and 1 mg/mL. Unless previously sterile, pipette this solution through a 0.22-µm syringe filter for sterilization.
NOTE: This protein is the biochemical signal that will be patterned onto the hydrogel surface. Also, while protein concentrations can vary, higher concentrations are ideal for stronger protein patterns.
- To make the reaction buffer, adjust the PBS to pH 8 and add 5 mM EDTA. Pipette reaction buffer into a 0.22 µm syringe filter for sterilization.
- React the protein with 2-iminothiolane by mixing both stock solutions together; use Table 1B to calculate the correct volume ratio. Typically, use a molar ratio of 8:1 2-iminothiolane to protein.
NOTE: For larger proteins or more dilute concentrations, use a higher molar ratio of 2-iminothiolane. Refer to the manufacturer's protocol15. - Incubate the reaction for 1 h at room temperature. Use a desalting spin micro-column to remove the thiolated protein product from the remaining reactants, following the manufacturer's protocol16.
NOTE: The exclusion limit of this resin is 5 KDa. - Use Ellman's Assay to quantitatively measure the number of free thiols per protein. Follow the manufacturer's protocol for the assay17.
3. Hydrogel Formation
- Create the gel precursor solution based on values calculated from Table 1C. Mix together the PEGDA, fibronectin, and LAP volumes. Pipette the solution vigorously to ensure a homogenous solution, but avoid creating bubbles. Keep the solution protected from light.
- Pipette the gel precursor solution carefully between the two glass slides of the gel mold. Typically, add a volume between 800 and 1,000 µL to the mold. Expose the gel solution in the mold to UV light (wavelength: 365 nm, power: 3-4 mW/cm2) for 1-2 min to form the hydrogel.
NOTE: Do not expose the hydrogel to prolonged UV light, as it will limit the surface protein patterning capabilities. - Take off the binder clips and gently remove top glass slide by applying opposite pressure to the two side spacers.
- Use an appropriately sized biopsy punch to cut out the hydrogel samples. Punch out multiple hydrogels from the gel rectangle to serve as replicates and control samples.
4. Hydrogel Stiffness Measurements
- Punch out hydrogels with an 8 mm-diameter biopsy punch for an 8 mm parallel plate rheometer. Load one hydrogel sample into the rheometer (see the Table of Materials).
- Lower the parallel plate geometry that is 8 mm in diameter until making contact with the surface of the gel. Keep a gap distance of 0.5 mm for a 0.5 mm-thick hydrogel sample.
- Perform time sweeps for 5 min at 0.1% strain, 0.1 Hz frequency, and 37 °C. Average the G' values across each time sweep. Run independent hydrogels for replicates of each composition.
NOTE: The G' values should be stable across time points; if they are not, percent strain and frequency sweeps should be performed to select the appropriate values14.
5. Protein Patterning
- Prepare the desired pattern for the photomask using a computer aided design (CAD) program. Print the photomask on a transparent sheet using a high-resolution printer. Soak the photomask in 70% ethanol for 10 min prior to use. Allow the photomask to air dry in a cell culture hood before use.
- Add the thiolated protein solution prepared in step 2 to the surface of the hydrogel for surface patterning. Pipette 1-2 µL/cm2 of thiolated protein solution to the surface of each cut-out gel.
NOTE: It is important to minimize the protein volume; only add enough to evenly cover the entire surface of the hydrogel sample. - Carefully place the photomask on the surface of the hydrogel. Do not allow air bubbles between the photomask and the hydrogel surface. Gently press down on the mask to remove any bubbles that form.
- Expose the hydrogel to a second round of UV light (wavelength: 365 nm, power: 3-4 mW/cm2) for 30-60 s.
NOTE: It is important to exposure the pattern to enough UV light to create a strong pattern without causing the loss of protein function. - Wash the hydrogels with PBS to remove unreacted species and place each hydrogel within the well plate. Be careful when placing the gels into each well; make sure that the patterned surface is face up. Wash the gels in PBS at 4 °C; gels are stable for at least two weeks at 4 °C.
6. Preparing Hydrogels for Cell Seeding
- Incubate the gels in basal medium for 5-10 min at 37 °C prior to seeding. For human umbilical vein endothelial cells (HUVECs), use EGM-2 medium without growth factors. Minimize the volume of medium added to each well to prevent hydrogel floating. For a 48-well plate, use a 250 µL volume of medium.
- Spin down the plate at 300 x g for 3 min to ensure that the hydrogels are located at the bottom of the well and are not floating. Do this immediately prior to cell seeding.
7. Cell Seeding on Hydrogels
- Use standard sterile mammalian cell culture procedures for cell seeding and experimental procedures. Remove HUVECs from tissue culture flasks using standard cell detachment protocols. Wash the tissue culture dish with sterile PBS and incubate with trypsin for 3-5 min at 37 °C.
- Quench the trypsinized cell suspension with excess cell medium or trypsin neutralizer solution.
- Spin down the cell suspension in the centrifuge at 300 x g for 5 min. Carefully remove the supernatant and keep the cell pellet.
- Resuspend the cell pellet in basal EGM-2 with no growth factors. Use a hemocytometer for cell counting.
- Add 75,000 cells/cm2 to each hydrogel surface by pipetting slowly into the center of each well so as not to disturb the gels. Place the well plate into a cell culture incubator at 37 °C. For several days, periodically remove the dish for observation.
8. Evaluation of Bioactivity
- Culture E. coli overnight in suspension in LB broth in an orbital suspension at 37 °C and 200 rpm. Spin down the E. coli culture, decant, and reconstitute the cell pellet in minimal volume of PBS. Weigh out the lysozyme and reconstitute at 1 mg/mL for the stock solution.
- Pipette a small volume (25 µL) of lysozyme stock solution into microcentrifuge tubes. Add varying levels of LAP photoinitiator (0-12 mM) and expose the samples to 1 min of UV light. In a separate group, add the same amount of LAP (2 mM) to lysozyme solution and expose samples to varying UV light times (0-4 min). Include replicates for each treatment.
- Add equal volume of concentrated E. coli solution to each lysozyme sample for a final lysozyme protein concentration of 0.5 mg/mL. Incubate the mixed solutions for 4 h at room temperature.
NOTE: This allows time for functional lysozyme to lyse the bacterial cell wall and release proteins into the solution. - Use untreated lysozyme incubated with E. coli for a positive control; consider this a 100% bioactive measurement. Use lysozyme solution incubated with PBS alone as a negative control.
- Spin down the samples to remove cell debris and keep the supernatant. Run a Bradford assay to quantify the total concentration of protein within the supernatant to measure the amount of bacterial lysate.
- Run a Bradford assay following the manufacturer's protocol18. Calculate the fold change in protein concentration compared to the negative control.
Representative Results
The protocol to create bioactive patterns on the surface of PEG hydrogels is illustrated in Figure 1. A spreadsheet was developed to calculate the volume and concentration for each stock solution (Table 1A). Proteins to be immobilized onto the surface of the hydrogel are modified with 2-iminothiolane (Figure 1B). This reaction is performed using the volumes from Table 1B. The precursor hydrogel solution is prepared with 10% weight/volume of PEGDA with LAP (Figure 1A). Various precursor PEGDA concentrations can be used to yield the desired substrate stiffness (Figure 2A). Fibronectin is included within this precursor solution for cell attachment purposes. After thorough mixing, this solution is pipetted into the prepared mold and exposed to UV light (Figure 1C). UV light exposure should be minimized; exposure should be just enough to produce a hydrogel. Hydrogel samples are punched out to the appropriate diameter for the desired well plate (Figure 1C). For surface patterning, modified protein solution is pipetted onto the surface of a hydrogel and spread evenly. Minimal volume should be used; protein volume should be just enough to cover the entire surface of the hydrogel. The predesigned photomask is placed directly onto the hydrogel surface; air bubbles between the mask and the hydrogel should be avoided. A second round of UV light is used to covalently conjugate UV-exposed proteins to the hydrogel. Hydrogel samples are rinsed to remove unreacted proteins and reveal the immobilized protein pattern (Figure 1D).
It is important to minimize photoinitiator concentration and UV exposure time when proteins are present. Using lysozyme bioactivity as an indicator, we found that the LAP photoinitiator concentration should be less than 2 mM (Figure 2B) and the UV exposure time should total less than 2 min (Figure 2C) to retain a protein bioactivity greater than 80%.
UV exposure time during hydrogel formation and protein patterning are both important parameters for developing a successful protocol (Figure 3). First of all, minimizing UV exposure during hydrogel formation is critical to maintaining free acrylate functional groups for subsequent protein immobilization reactions (Figure 3A). Hydrogels exposed to UV light for longer than 2 min are unable to create immobilized protein patterns. Additionally, as the UV exposure to the protein pattern increases, more proteins react to the surface (Figure 3B).
Finally, cells can be cultured onto these patterned hydrogel substrates to manipulate cell behavior. To show the potential of immobilized patterns on hydrogels, we patterned VEGF, a growth factor important for endothelial cells, and cultured HUVECs on the surface using basal EGM-2 medium (Figure 4). HUVECs were uniformly seeded onto the surface of VEGF-patterned PEG hydrogels (Figure 4A). Two days after seeding, HUVECs were observed to migrate towards the spatial regions of the hydrogel that contained immobilized VEGF (Figure 4B, C). This is one example of a bioactive protein pattern on PEG hydrogels being used to influence cell behavior.
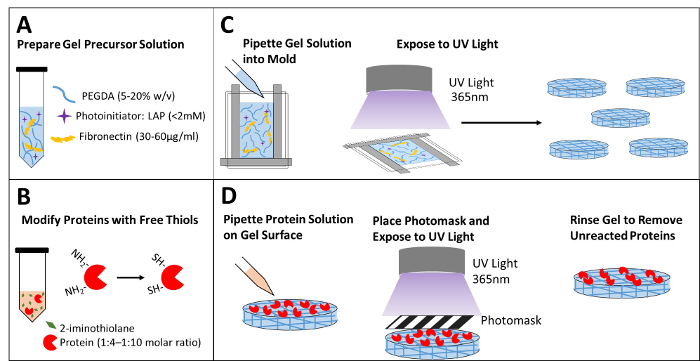
Figure 1: Schematic of hydrogel formation and protein patterning. (A) Prepare precursor hydrogel solution with PEGDA monomers, photoinitiator, and extracellular protein for cell attachment. (B) Modify the proteins with free thiol groups by reacting with 2-iminothiolane. (C) Pipette the precursor solution into the prepared mold and expose it to UV light to form the hydrogel. Punch out hydrogel samples of the desired size. (D) Pipette the modified protein solution onto the surface of the hydrogel, place a photomask on the surface, and expose it to a second round of UV light. Rinse the gel to remove unreacted species prior to imaging and cell seeding. Please click here to view a larger version of this figure.
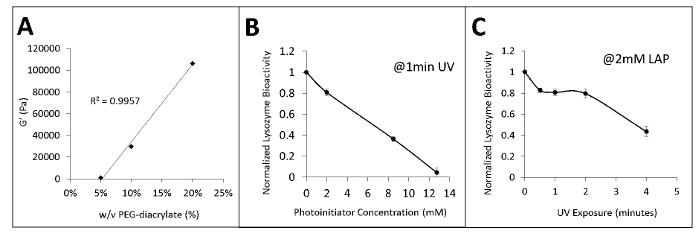
Figure 2: Modulating stiffness and protein bioactivity. (A) Changes in the concentration of PEGDA monomers within the precursor gel solution alter hydrogel stiffness. (B) Increasing the concentration of LAP photoinitiator lowers protein function after 1 min of UV exposure. (C) Increasing UV exposure time lowers protein function with 2 mM LAP. All error bars represent standard deviation of replicates. Please click here to view a larger version of this figure.
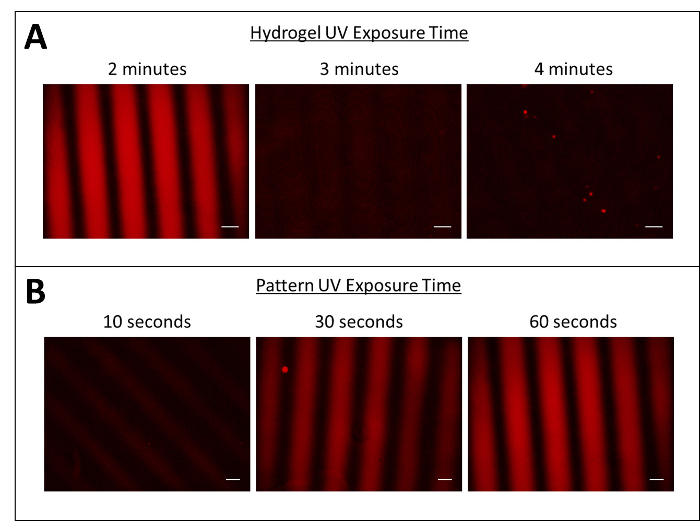

Figure 3: Pattern and hydrogel UV exposure times optimized for protein patterning. (A) Minimizing UV exposure times for hydrogel formation allows for surface patterning. (B) Increasing UV exposure times for surface patterning increases pattern strength. Scale bars = 200 µm. Please click here to view a larger version of this figure.
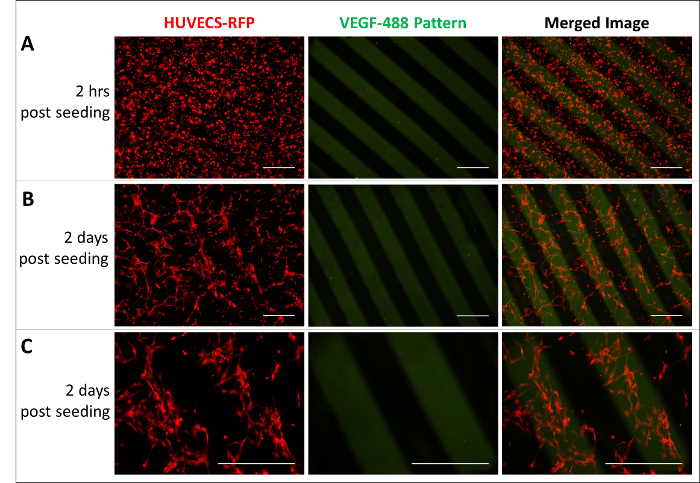
Figure 4: Endothelial cells responding to a VEGF pattern. (A) Uniform HUVECs seeded on hydrogels. (B and C) HUVECs sense the VEGF pattern and migrate towards immobilized VEGF. Images were taken two days after seeding in (B) 4X and (C) 10X magnification. Scale bars = 500 µm. HUVEC-RFPs (red) and VEGF-488 pattern (green) were captured with excitation and emission filters at 528/553 and 465/495, respectively. Please click here to view a larger version of this figure.
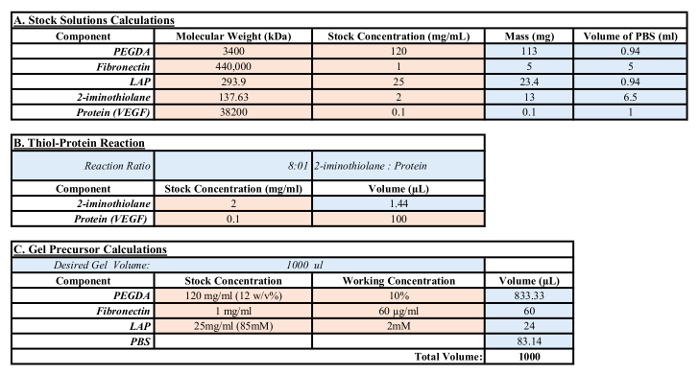
Table 1: Calculations for stock and gel precursor solutions. The red box indicates user-defined values, such as molecular weight, desired stock concentrations, and weighed-out masses. Blue boxes represent values that have been calculated based on user-defined values.
Discussion
This protocol provides a method for creating bioactive protein patterns for biological applications. There are several modifications that can be made to adapt this protocol for different experiments. First, cell attachment requirements will vary for different cell types. If poor cell attachment to the gels is initially observed, increasing the concentration of the ECM protein within the precursor solution is advised. Other ECM proteins can be used instead of fibronectin, including different types of collagen, laminin, or a combination thereof. For each new cell type, cell attachment should be optimized prior to hydrogel patterning. This protocol also allows for user-designed photomasks. Based on the desired application, photomasks can be produced with various feature sizes, shapes, and overall patterns. Uniform immobilization can be achieved in the absence of a photomask.
This protocol has certain limitations. As highlighted in the Representative Results, this method is sensitive to the amount of UV light at each step. Overexposure during the hydrogel formation step limits the available acrylate groups for subsequent surface bioconjugation. Therefore, a key step in this protocol is well-managed UV light exposure times for each step. Also, this protocol requires a high concentration protein stock for successful patterning. Low concentrations of protein will result in poor surface patterns. Additionally, photomasks are also limiting in that they can only produce discrete patterns. More complex patterns can be achieved with similar approaches but require a more advanced methodology.
This protocol is significant to existing methods as it provides a simple method for adjusting substrate stiffness and protein patterning. Using acrylate chemistry for the hydrogel formation allows for a magnitude range of substrate stiffness within the physiological range. Simply adjusting the concentration of PEGDA within the precursor solution gives control over the hydrogel stiffness. Additionally, the use of click chemistry for protein patterning allows for rapid conjugation between hydrogel substrate and thiolated modified protein. This is a key design feature, as it allows this protocol to be applicable to any protein or peptide of interest.
PEG hydrogels are promising biomaterials that can be used to explore new platforms for displaying biochemical cues to biological systems. Whether uniform surface immobilization or spatially specific patterns, these techniques provide novel ways to control cell behavior. Moving forward, the advancement of biomaterial technology will provide new insights into cell behavior and further our abilities to recapitulate in vivo signaling motifs within in vitro systems. This can be beneficial for stem cell differentiation and modeling developmental signaling within a controlled experimental system.
Declarações
The authors have nothing to disclose.
Acknowledgements
This study was mainly supported by grants from the American Heart Association Scientist Development Grant (12SDG12050083 to G.D.), the National Institutes of Health (R21HL102773, R01HL118245 to G.D.) and the National Science Foundation (CBET-1263455 and CBET-1350240 to G.D.).
Materials
| PEG-diacrylate (PEGDA) | Laysan Bio | ACRL-PEG-ACRL-3400 | Can also be synthesized or purchased through other venders. Different molecular weights can be used. |
| Lithium Phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) | Synthesized in lab | ||
| Fibronectin | Corning | 356008 | Other cell attachment proteins can be used, such as laminin, matrigel |
| Phosphate-buffered saline (PBS) | Sigma | D8537-500ML | |
| Photomask | FineLine Imaging | n/a | Custom prints on transparent sheets with high resolution DPI. |
| Binder Clips | Various Vendors | ||
| Compact UV Light Source (365nm) | UVP | UVP-21 | Other UV light sources can be used, calibration of power is required. |
| 2-iminothiolane (Pierce Traut’s Reagent) | Thermo Sci. | 26101 | |
| Ellman’s Reagent: DTNB; 5,5-dithio-bis(2-nitrobenzoic acid) | Thermo Sci. | 22582 | |
| human umbilical vein endothelial cells (HUVECs) | Lonza | passage number between 6- 10 | |
| EGM-2 Media | Lonza | CC31-56, CC-3162 | EGM-2 without growth factors was used in experiments. Full EGM-2 media was used for cell maintainance |
| 0.25% Trypsin EDTA | Life Tech | 25200-056 | |
| Trypsin Neutralizer | Life Tech | R-002-100 | |
| Centrifuge | Various Venders | ||
| Hemocytometer | Hausser Sci. Bright-line | ||
| Ethylenediaminetetraacetic acid (EDTA) | Sigma Aldrich | E6758 | |
| 0.22µm filter | Cell Treat | 229743 | |
| 1mL Syringe | |||
| Glass Microscope Slides | Fisher Sci. | 12-550C | |
| Plastic spacers | Various Venders | 0.5mm thickness | |
| 70% Ethanol | BICCA | 2546.70-1 | |
| Bio-shield | Bio-shield | 19-150-0010 | |
| Bradford Reagent | BIO-RAD | ||
| Desalting Resin – Sephadex G-25 | GE Healthcare | 95016-754 | |
| Microspin Columns | Thermo Sci. | PI69725 | |
| AR-G2 rehometer | TA Instruments |
Referências
- Yao, S., et al. Co-effects of matrix low elasticity and aligned topography on stem cell neurogenic differentiation and rapid neurite outgrowth. Nanoscale. 8 (19), 10252-10265 (2016).
- Evans, N. D., et al. Substrate stiffness affects early differentiation events in embryonic stem cells. Eur Cells Mater. 18, 1-13 (2009).
- Ye, K., et al. Matrix Stiffness and Nanoscale Spatial Organization of Cell-Adhesive Ligands Direct Stem Cell Fate. Nano Lett. 15 (7), 4720-4729 (2015).
- Discher, D. E., Janmey, P., Wang, Y. L. Tissue cells feel and respond to the stiffness of their substrate. Science. 310 (5751), 1139-1143 (2005).
- Sridhar, B. V., Doyle, N. R., Randolph, M. A., Anseth, K. S. Covalently tethered TGF-beta1 with encapsulated chondrocytes in a PEG hydrogel system enhances extracellular matrix production. J Biomed Mater Res A. 102 (12), 4464-4472 (2014).
- Salinas, C. N., Anseth, K. S. Decorin moieties tethered into PEG networks induce chondrogenesis of human mesenchymal stem cells. J Biomed Mater Res A. 90 (2), 456-464 (2009).
- Joddar, B., Guy, A. T., Kamiguchi, H., Ito, Y. Spatial gradients of chemotropic factors from immobilized patterns to guide axonal growth and regeneration. Biomaterials. 34 (37), 9593-9601 (2013).
- Wylie, R. G., Ahsan, S., Aizawa, Y., Maxwell, K. L., Morshead, C. M., Shoichet, M. S. Spatially controlled simultaneous patterning of multiple growth factors in three-dimensional hydrogels. Nat Mater. 10 (10), 799-806 (2011).
- Tibbitt, M. W., Anseth, K. S. Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnol Bioeng. 103 (4), 655-663 (2009).
- Lutolf, M. P., Hubbell, J. A. Synthetic biomaterials as instructive extracellular microenvironments for morphogenesis in tissue engineering. Nat Biotechnol. 23 (1), 47-55 (2005).
- Saik, J. E., Gould, D. J., Keswani, A. H., Dickinson, M. E., West, J. L. Biomimetic hydrogels with immobilized ephrinA1 for therapeutic angiogenesis. Biomacromolecules. 12 (7), 2715-2722 (2011).
- McCall, J. D., Anseth, K. S. Thiol-ene photopolymerizations provide a facile method to encapsulate proteins and maintain their bioactivity. Biomacromolecules. 13 (8), 2410-2417 (2012).
- Fairbanks, B. D., Schwartz, M. P., Bowman, C. N., Anseth, K. S. Photoinitiated polymerization of PEG-diacrylate with lithium phenyl-2,4,6-trimethylbenzoylphosphinate: polymerization rate and cytocompatibility. Biomaterials. 29 (6), 997-1003 (2009).
- Zuidema, J. M., Rivet, C. J., Gilbert, R. J., Morrison, F. A. A protocol for rheological characterization of hydrogels for tissue engineering strategies. J Biomed Mater Res B Appl Biomater. 102 (5), 1063-1073 (2014).
- Truat’s Reagent Instructions. Thermo Scientific Available from: https://tools.thermofisher.com/content/sfs/…/MAN0011238_Trauts_Reag_UG.pdf (2017)
- Ellman’s Reagent Instructions. Thermo Scientific Available from: https://tools.thermofisher.com/content/sfs/manuals/MAN0011216_Ellmans_Reag_UG.pdf (2017)
- Desalting Columns. GE Life Sciences Available from: https://www.gelifesciences.com/gehcls_images/GELS/Related%20Content/Files/1478781880316/litdoc52130800_2016111034421.pdf (2017)

