Visualizing and Quantifying Endonuclease-Based Site-Specific DNA Damage
Summary
This article introduces essential steps of immunostaining and chromatin immunoprecipitation. These protocols are commonly used to study DNA damage-related cellular processes and to visualize and quantify the recruitment of proteins implicated in DNA repair.
Abstract
Cells are continuously exposed to various DNA damaging agents, inducing different cellular responses. Applying biochemical and genetic approaches is essential in revealing cellular events associated with the recruitment and assembly of DNA repair complexes at the site of DNA damage. In the last few years, several powerful tools have been developed to induce site-specific DNA damage. Moreover, novel seminal techniques allow us to study these processes at the single-cell resolution level using both fixed and living cells. Although these techniques have been used to study various biological processes, herein we present the most widely used protocols in the field of DNA repair, Fluorescence Immunostaining (IF) and Chromatin Immunoprecipitation (ChIP), which in combination with endonuclease-based site-specific DNA damage make it possible to visualize and quantify the genomic occupancy of DNA repair factors in a directed and regulated fashion, respectively. These techniques provide powerful tools for the researchers to identify novel proteins bound to the damaged genomic locus as well as their post-translational modifications necessary for their fine-tune regulation during DNA repair.
Introduction
Our genome is constantly being challenged by various DNA damaging agents. These assaults can derive from environmental sources, such as UV light or irradiation, as well as from endogenous sources, such as metabolic by-products caused by oxidative stress or replication errors1,2. These lesions can affect the integrity of either one or both DNA strands, and if the generated errors become persistent, it frequently leads to translocations and genome instability, which may result in tumorigenesis3,4. To maintain genome integrity, multiple repair systems have been developed during evolution. According to the chemical and physical properties of specific types of DNA damage, multiple repair mechanisms can be activated. Mismatches, abasic sites, single-strand breaks, and 8-oxoguanine (8-oxoG) can be removed either by mismatch repair or base-excision repair pathway5,6. Lesions caused by UV-induced photoproducts and bulky adducts can be repaired either by nucleotide-excision repair (NER) or DNA double-strand break repair (DSBR) process7,8. NER consists of two main sub-pathways: transcription-coupled NER (TC-NER) and global genomic NER (GG-NER). Regarding the cell cycle phase, following DNA double-strand break induction, two sub-pathways can be activated: non-homologous end joining (NHEJ) and homologous recombination (HR)1,9. NHEJ, which is the dominant pathway in resting cells, can be activated in all cell cycle phases, representing a faster but error-prone pathway10. On the other hand, HR is an error-free pathway, in which the DSBs are repaired based on sequence-homology search of the sister chromatids, therefore it is mainly present in S and G2 cell cycle phases11. Furthermore, microhomology-mediated end joining (MMEJ) is another DSB repair mechanism, distinct from the aforementioned ones, based on a KU70/80- and RAD51-independent way of re-ligation of previously resected microhomologous sequences flanking the broken DNA ends. Therefore, MMEJ is considered to be error-prone and highly mutagenic12. During DNA repair, DSBs can induce the DNA damage response (DDR), which results in the activation of checkpoint kinases that halt the cell cycle during repair13,14,15. The DDR is activated as a response to the recruitment and extensive spreading of initiator key players of the repair process around the lesions, contributing to the formation of a repair focus. In this early signaling cascade, the ATM (Ataxia Telangiectasia Mutated) kinase plays a pivotal role by catalyzing the phosphorylation of the histone variant H2AX at Ser139 (referred to as γH2AX) around the lesion16. This early event is responsible for the recruitment of additional repair factors and the initiation of downstream repair processes. Although the exact function of the recruited proteins at the repair focus has not yet been fully characterized, the formation and the dynamics of repair foci have been investigated by several laboratories. These markers are extensively used to follow the repair kinetics, but their precise role during the repair process remains elusive. Due to the great importance yet poor understanding of DNA repair-related cellular processes, several methods have been developed so far to induce and visualize the DDR.
Various methods and systems have been established to induce the desired type of DNA damage. For instance, some agents [such as neocarzinostatin (NCS), phleomycin, bleomycin, γ-irradiation, UV] can induce large numbers of random DNA breaks at non-predictive genomic positions, while others (endonucleases, such as AsiSI, I-PpoI or I-SceI, as well as laser striping) can induce DNA breaks at known genomic loci17,18,19,20,21. Here, we focus on the endonuclease-based techniques currently used to study the DDR in mammalian and yeast cells. Aside from highlighting the principles of these techniques, we emphasize both their advantages and disadvantages.
Protocol
1. Immunodetection of specific proteins
- Preparation of cell culture and experimental setup
- Maintain U2OS cells in monolayers in DMEM culture medium supplemented with 10% fetal calf serum, 2 mM glutamine, and 1% antibiotic-antimycotic solution.
NOTE: For endonuclease-based DNA damage induction, use charcoal-treated or steroid-free medium to avoid system leakiness. - Grow cells in a humidified 5% CO2 environment at 37 °C until 80% confluency, renewing medium every 2-3 days.
- Aspirate the medium and wash the cells with 1x PBS. Detach cells with Trypsin-EDTA solution. When the cells detach, stop the trypsin activity by adding culture medium to the cells, yielding a cell suspension.
- Count the cells using a cell counting chamber. Plate 2 x 104 cells/mL/well on a 24-well plate, with sterile 12 mm round coverslips in each well.
- Incubate cells for 24 h at 37 °C in a humidified 5% CO2 atmosphere to allow attachment onto the coverslips.
- Treat the cells with 10 ng/mL of neocarzinostatin (NCS) by directly pipetting the damaging agent to the cultured medium. Incubate the cells with the NCS-containing medium for 15 min, then wash them with 1x PBS and add fresh, supplemented culture medium to the cells. Otherwise, use appropriate agent (i.e., 4-OHT) to induce DSBs via endonuclease-based systems without refreshing the medium22.
NOTE: Alternatively, use irradiation to induce DNA damage, ranging from 30 min up to 8 h of recovery time by using neutron flux between 2-20 Gy23. - Incubate cells for 1-8 h at 37 °C in a humidified 5% CO2 atmosphere to follow the kinetics of DNA repair.
- Maintain U2OS cells in monolayers in DMEM culture medium supplemented with 10% fetal calf serum, 2 mM glutamine, and 1% antibiotic-antimycotic solution.
- Fixation of cells
NOTE: 300-500 µL of solutions/well should be used in the following steps (steps 1.2-1.5) to cover all cells adequately. Each incubation and washing step (except the antibody incubation) should be performed on an orbital shaker with gentle agitation.- Following DSB induction and incubation of cells, remove the medium from the attached cells and wash the cells once with 1x PBS.
- Fix cells with 4% formaldehyde-PBS solution for 20 min at 25 °C.
- Permeabilization of cells
- Remove the fixing solution and wash cells three times with 1x PBS for 5 min each.
- Remove the PBS and add 0.2% Triton X-100 dissolved in PBS. Incubate the samples for 20 min.
- Blocking of non-specific binding sites
- Wash the cells three times with 1x PBS.
- Block non-specific binding sites with 5% BSA (Bovine Serum Fraction V albumin) diluted in PBST (1x PBS supplemented with 0.1% Tween-20), and incubate the permeabilized samples for at least 20 min.
- Immunofluorescence staining
- Add the proper amount of primary antibody (i.e., anti-γH2AX, anti-DNA-PKcs) diluted in 1% BSA-PBST solution. Place each coverslip upside down onto a paraffin film over a 10 µL droplet of the diluted anti-γH2AX antibody.
NOTE: In case of co-immunostaining dilute appropriately both antibodies in the same 1% BSA-PBST solution. - Incubate the samples in a humidity chamber for 1.5 h at 4 °C.
NOTE: Incubation can also be performed at 4 °C overnight. - Place the coverslips back being side up into the 24-well plate and wash three times for 5 min with 1x PBS.
- Add the proper amount of secondary antibody diluted in 1% BSA-PBST. Place each coverslip upside down onto a paraffin film over a 10 µL droplet of the diluted antibody.
- Incubate the samples in a humidity chamber at 4 °C for 1 h.
- Place the coverslips back being side up into the 24-well plate and wash three times for 5 min with 1x PBS.
- Before removing the last PBS washing solution, gently take out the coverslips using a tweezer and needle and then place them upside down onto glass slides with droplets of mounting medium (supplemented with DAPI).
NOTE: Avoid the formation of air bubbles. When the mounting medium dries, it is recommended to seal the edges of the coverslips with nail polish to prevent shriveling of the samples.
- Add the proper amount of primary antibody (i.e., anti-γH2AX, anti-DNA-PKcs) diluted in 1% BSA-PBST solution. Place each coverslip upside down onto a paraffin film over a 10 µL droplet of the diluted anti-γH2AX antibody.
2. Chromatin immunoprecipitation
- Cell collection, crosslinking, cell and nuclear lysis, and DNA fragmentation
- Culture approximately 5 x 106 cells/mL in a 150 mm dish for each sample.
- Remove the culture medium and wash the cells twice with ice-cold 1x PBS.
- Fix the cells with 1% formaldehyde-PBS solution, place the plates on an orbital shaker, and agitate gently for 20 min.
NOTE: Formaldehyde is volatile; always prepare a fresh working solution. In some cases, the formaldehyde solution contains methanol to stabilize it, but it is better to use a methanol-free solution to avoid interference with downstream reactions. - Stop the fixation with 125 mM glycine and incubate on an orbital shaker with gentle agitation for 5 min at 25 °C.
- Place the plates on ice and wash twice with ice-cold 1x PBS.
- Scrape the cells in ice-cold 1x PBS and transfer them into 15 mL conical tubes.
- Centrifuge the cells at 2,500 x g for 5 min at 4 °C.
- Carefully aspirate the supernatant and resuspend the pellet in 2 mL of cell lysis buffer [5 mM PIPES pH 8.0, 85 mM KCl, 0.5% NP-40, 1x PIC (protease inhibitor cocktail)] and incubate on ice for 10 min.
- Centrifuge the cell suspension at 2,500 x g for 5 min at 4 °C.
- Carefully discard the supernatant and resuspend the pellet in 500-1,500 µL of nuclear lysis buffer (50 mM Tris-HCl pH 8.0, 10 mM EDTA pH 8.0, 0.8% SDS, 1x PIC) and incubate on ice for 30-60 min. Transfer the lysate into a polystyrene conical tube suitable for sonication.
NOTE: Since the nuclear lysis buffer contains SDS, it will precipitate on ice, and the solution will turn white. The solution should turn transparent following sonication. - Sonicate the lysate to shear DNA to an average fragment size of 300-1,000 bp.
NOTE: The appropriate sonication cycles and conditions should be set according to the cell type and the sonication equipment. Fragments smaller than 200 bp are not suitable for ChIP, because the nucleosome-DNA interactions can be disrupted.
- Reversal of crosslinking, determination of sonicated fragment sizes
- Take out 100 µL of the sonicated sample to verify the fragment size of the sonicated chromatin. The remaining chromatin should be stored at −80 °C.
- Add 0.5 mg/mL RNase A to each 100 µL of the sample and incubate them at 37 °C for 20 min to activate the RNase.
- Incubate the samples at 65 °C overnight.
- The next day, add 500 µg/mL of Proteinase K and 0.5% SDS, and incubate the samples at 50 °C for 3 h.
- Add 0.5 volume of phenol and 0.5 volume chloroform-isoamyl alcohol mix (24:1) to each sample.
- Vortex for 1 min.
- Centrifuge at 13,000 x g for 10 min.
- Transfer the upper aqueous phase to a new microcentrifuge tube.
- Add 1 volume chloroform-isoamyl alcohol mix (24:1) to each sample.
- Vortex for 1 min.
- Centrifuge at 13,000 x g for 10 min.
- Transfer the upper aqueous phase to a new microcentrifuge tube.
- Add 2.5 volumes of 96% ethanol and 0.1 volume of 3 M Na-Acetate pH 5.2.
- Incubate for at least 20 min at −80 °C.
- Centrifuge the samples at 13,000 x g for 10 min at 4 °C.
- Remove the ethanol and wash the pellet with 400 µL of 70% ethanol.
- Centrifuge the samples at 13,000 x g for 10 min at 4 °C.
- Remove the ethanol and air dry the pellet.
- Resuspend the pellet in 10 µL of TE.
- Run the samples on a 0.8% agarose gel. The sonicated chromatin size should be around 500 bp.
NOTE: Use bromophenol blue-free loading buffer because the size of this dye is approximately 500 bp, which can disturb the proper detection of chromatin fragments. Instead, it is recommended to use loading buffer complemented with xylene-cyanole, which is approximately 3,000 bp. - If the chromatin size is acceptable, dilute the frozen chromatin samples from step 2.1. in 3 volumes of dilution buffer (10 mM Tris-HCl pH 8.0, 0.5 mM EGTA pH 8.0, 1% Triton X-100, 140 mM NaCl, 1x PIC) and mix the samples via rotation for 10 min at 4 °C.
NOTE: This step is necessary to dilute the SDS present in the nuclear lysis buffer to avoid interference with downstream reactions, including the measurement of chromatin concentration. - Measure the DNA concentration of the chromatin samples at 260/280 nm using a spectrophotometer.
- Preparation of beads, pre-clearing, and immunoprecipitation
- Prepare beads (sheep anti-rabbit or mouse IgG) for the pre-clearing and immunoprecipitation steps. Wash beads twice for 10 min at 4 °C with RIPA buffer (50 mM Tris-HCl pH 8.0, 1 mM EDTA pH 8.0, 1% Triton X-100, 0.1% Na-DOC, 0.1% SDS, 150 mM NaCl and 1X PIC).
- Resuspend the beads in the same volume of RIPA buffer as in step 2.3.1.
- Pre-clear 25-30 µg chromatin of each sample with 4 µL of the beads via rotation for 1-2 h at 4 °C.
NOTE: Add RIPA buffer to each chromatin sample up to 500 µL of final volume to let the samples mix properly under rotation. Do not forget to take out chromatin for NAC (No Antibody Control) and TIC (Total Input Control) in the case of each sample set. TICs only require a final volume of up to 200 µL. - Precipitate the beads with a magnet and transfer the supernatant to a new microcentrifuge tube.
- Add the appropriate amount of antibody to each chromatin sample (except NAC and TIC) and rotate overnight at 4 °C.
- Next day, add 40 µL of washed beads to each sample (except TIC) and incubate them overnight, rotating at 4 °C.
- Washing
- Wash once with 300 µL of Low Salt buffer (20 mM Tris-HCl pH 8.0, 150 mM NaCl, 2 mM EDTA pH 8.0, 1% Triton X-100, 0.1% SDS, 1x PIC) for 10 min via rotation at 4 °C.
- Wash once with 300 µL of High Salt buffer (20 mM Tris-HCl pH 8.0, 300 mM NaCl, 2 mM EDTA pH 8.0, 1% Triton X-100, 0.1% SDS, 1x PIC) for 10 min via rotation at 4 °C.
- Wash once with 300 µL of LiCl buffer (250 mM LiCl, 1% NP-40, 1% Na-DOC, 1 mM EDTA pH 8.0, 10 mM Tris-HCl pH 8.0, 1x PIC) for 10 min via rotation at 4 °C.
- Wash twice with 300 µL of TE (10 mM Tris-HCl pH 8.0, 1 mM EDTA pH 8.0) for 10 min via rotation, for the first wash at 4 °C, and the second wash at 25 °C.
- Elution
- Add 200 µL of the elution buffer (1% SDS and 100 mM NaHCO3) to the beads and incubate at 65 °C in a thermo-shaker for 15 min with continuous shaking (approx. 400 RPM). Transfer the supernatant to a new tube and elute beads again in 200 µL of Elution buffer. Combine eluates (400 µL final volume).
- Add NaCl to a final concentration of 200 mM in each sample. Supplement the TIC samples with 200 µL of Elution buffer and add NaCl as well.
NOTE: From this step, TIC should be handled under the same conditions as the other samples. - Incubate the samples at 65 °C (without shaking) for at least 6 h.
- Add 1 mL of cold 100% ethanol to each sample, rotate the tubes twice to mix, and precipitate DNA overnight at −80 °C.
- The next day, centrifuge for 30 min at 13,000 x g at 4 °C.
- Discard the supernatant and wash the pellet with 70% EtOH.
- Centrifuge the samples at 13,000 x g for 10 min at 4 °C.
- Discard the supernatant and air dry the pellets.
- Resuspend the pellets in 100 µL of TE and add 0.5 mg/mL of RNase A to each sample. Incubate at 37 °C for 20 min to activate the RNase.
- Reversal of crosslinking
- Add 500 µg/mL of Proteinase K and 0.5% SDS then incubate the samples at 50 °C for 2 h.
NOTE: If proceeding to ChIP-seq, avoid phenol-chloroform extraction, as it inhibits the downstream NGS process. Instead, it is recommended to use a commercially available kit (see Table of Materials). - Add 0.5 volume of phenol and 0.5 volume of chloroform-isoamyl alcohol mix (24:1) to each sample.
- Vortex for 1 min.
- Centrifuge at 13,000 x g for 10 min.
- Transfer the upper aqueous phase to a new microcentrifuge tube.
- Add 1 volume of chloroform-isoamyl alcohol mix (24:1) to each sample.
- Vortex for 1 min.
- Centrifuge at 13,000 x g for 10 min.
- Transfer the upper aqueous phase to a new microcentrifuge tube.
- Add 500 µg/mL of Proteinase K and 0.5% SDS then incubate the samples at 50 °C for 2 h.
- DNA extraction
- Add 2.5 volumes of 96% ethanol and 0.1 volume of 3 M Na-Acetate pH 5.2.
- Incubate for at least 20 min at −80 °C.
- Centrifuge the samples at 13,000 x g for 10 min at 4 °C.
- Remove the ethanol and wash the pellet with 400 µL of 70% ethanol.
- Centrifuge the samples at 13,000 x g for 10 min at 4 °C.
- Remove the ethanol and air dry the pellet.
- Resuspend the pellet in 50 µL of TE.
Representative Results
Studying site-directed DSB-induced repair processes in cells can be achieved via either stable or transient transfection. However, it should be noted that stable transfection ensures a homogenous cell population, which gives a unified and thus more reliable cellular response. In the case of transient transfection, only a small proportion of the cell population takes up and maintains the plasmid, which introduces diversity into the experiment. Establishing ER-I-PpoI or ER-AsiSI endonuclease-based cell systems require a 50% confluent cell population, which is more effectively transfected with plasmids encoding the endonuclease. For transfection, commercially available transfection reagents or viral infection-based methods can also be used. If a microscopic visualization technique is to be applied and transient transfection is required, the directed DSBs can be induced by 4-OHT addition 24 h after transfection, which binds to the ER-fused endonucleases and allows the nuclear translocation and DSB induction. To determine the most appropriate time-points, immunofluorescence-based microscopy and western blot detection of γH2AX at different time-points following 4-OHT treatment can be performed. Under physiological conditions, a maximum of 10-15 γH2AX foci per cell can be detected, and the formation of strong repair foci can be triggered by endonucleases (or various other techniques not discussed here e.g., laser microirradiation). A typical I-PpoI endonuclease leads to the formation of elevated γH2AX signals around the nucleolus by inducing DSBs at ribosomal DNA (rDNA). If the breaks are repaired by NHEJ or HR, the number of repair foci decreases over time. For this reason, representative time-points at 0 h, 30 min, 1, 2, 4, and 8 h following 4-OHT treatments are recommended. To track DNA repair processes, the most commonly used cell line is U2OS, as all known repair pathways are fully functional in these cells. When investigating several proteins in the same cells, co-localization can be studied by combining antibodies conjugated with different fluorophores with different emission wavelengths raised in different animal species as shown in Figure 1. Therein, the induction of DSBs via an inducible stable cell line is represented which is based on the ER-AsiSI restriction endonuclease fused with hemagglutinin tag (HA). Doxycycline can induce the expression and sequestration of HA-ER-AsiSI in the cytoplasm which can be tracked using an antibody against HA (Figure 1. third column, second raw). Incubation for 4 h with 4-OHT, 24 h after doxycycline addition, can induce high number of DSBs since the endonuclease has been translocated to the nucleus (Figure 1. third column, third raw and second column, third raw). DSBs can be visualized by using an antibody recognizing γH2AX.
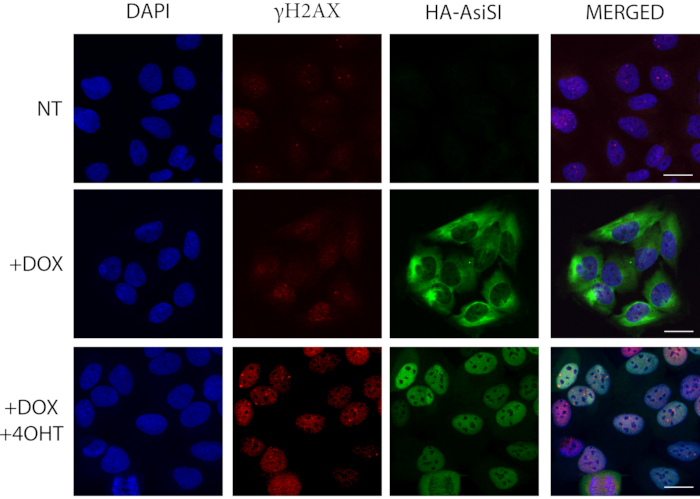
Figure 1: Immunofluorescence microscopy is used to detect γH2AX in cultured cells expressing HA-ER-AsiSI. DOX (doxycycline) addition activates the cytoplasmic expression of HA-ER-AsiSI and 4-OHT (4-hydroxytamoxifen) (4 h) induces the nuclear translocation of the fusion protein, which leads to the induction of DSBs at known genomic positions. HA (hemagglutinin) staining (anti-HA antibody) represents the HA-ER-AsiSI fusion protein in green, and the induction of breaks is verified by γH2AX staining (anti-γH2AX antibody) in red. Scale bars represent 20 µm. Please click here to view a larger version of this figure.
Co-localization of repair proteins at the damage site indicates that they are recruited to the same DNA lesion site, but they do not necessarily interact with each other. The resolution of the confocal microscopy is approximately 300 nm; to determine the binding pattern of specific repair proteins at the break site, super-resolution microscopy (STORM) is instead recommended 24. However, this method requires expensive microscopic equipment and an expert researcher. Alternatively, the binding pattern of the repair proteins can be examined by chromatin immunoprecipitation using DIvA or U2OS-pEP15 stable cell lines, which can express AsiSI and I-PpoI endonucleases in a regulated fashion, respectively17,21. Upon 4-OHT addition, both endonucleases can cut the DNA in a sequence specific manner which provides us the opportunity to design locus specific primers to the expected break sites and their surrounding genomic regions. By applying γH2AX antibody in the immunoprecipitation part of the ChIP, we can temporally follow the DNA repair kinetics upon different conditions (such as silencing or inhibition of certain repair factors of interest, i.e., DNA-PKcs). A typical experimental result obtained using ChIP-qPCR is represented in Figure 2. Therein, the temporal enrichment of γH2AX is demonstrated as a response to I-PpoI-induced DNA damage. On the left part of the image, the timely detected γH2AX signal is shown at the break site while on the right part, the γH2AX distribution is represented at a control gene region at which DSBs have not been induced.
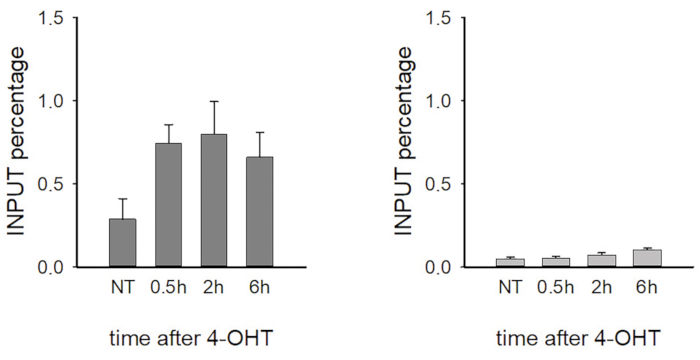
Figure 2: Temporal enrichment of γH2AX as determined by chromatin immunoprecipitation in response to I-PpoI-induced DNA damage. Left, γH2AX signal at the break site; right, γH2AX distribution at a control region where DSBs were not induced (anti-γH2AX antibody). The represented results are derived from one biological experiment, and error bars indicate the variations of the corresponding sample replicates. N=3. Please click here to view a larger version of this figure.
Discussion
Although DNA repair is a relatively recent research field, our knowledge is rapidly expanding with the help of various biochemical and microscopic methods. Preserving genetic information is crucial for cells since mutations occurring in genes involved in repair processes are among the leading causes of tumorigenesis and therefore elucidating the key steps of DNA repair pathways is essential.
Biochemical techniques (i.e., western blot, immunoprecipitation, mass-spectrometry, etc.) require large number of cells and the studied repair processes represent a snapshot of the desired cell population. Performing ChIP experiments is laboring, troublesome and many considerations have to be taken into account when designing specific experiment to study the process of DSB repair. The following steps are some examples: (I) cells should be properly lysed; it is highly recommended to apply a two-step lysation method by using separate cell- and nuclear lysis buffer to ensure higher accessibility to the chromatin fraction (II) chromatin should not be over- or undersonicated; the appropriate conditions of sonication should be optimized to each cell type beforehand (III) the appropriate amount of purified antibody should be determined since antibodies against the same protein from different companies exhibit non-identical characteristics (IV) for the efficient immunoprecipitation, 25-30 ug initial chromatin should be used in each condition (V) the appropriate time of fixation should be optimized, as overfixation can result in false positive results by cross-linking the distant protein complexes and underfixation can prevent the proper cross-linking between DNA and the desired protein (VI) based on the applied antibody, the type of the beads (protein A or G) should be carefully determined (VII) during washing steps, the order of the washing buffers should be thoroughly kept to avoid the release of the antibody from the beads (VIII) during DNA extraction, phenol traces should be properly eliminated to avoid decreasing the efficiency of the further downstream reactions. Since this method reduces the yield of recovered DNA, our personal advice is to rather use specific DNA-purification kit. When all critical points have been properly addressed, ChIP can provide valuable data for the occupancy of the desired protein at different genomic loci and can unravel critical steps in DNA repair processes.
However, ChIP combined with qPCR is an indirect approach to study the protein distribution at selected genomic regions and does not allow to specifically recognize the DNA binding site or directly examine the function of the protein. Mono- or polyclonal antibodies used to capture the protein-DNA complexes can also cross-react with other proteins leading to false-positive data and therefore, the antibodies used in this technique should be ChIP-grade and highly specific against the protein of interest. However, ChIP is a widely used technique and further approaches based on this have been developed, such as ChIP-on-chip and ChIP-seq. The former one relies on hybridizing the immunoprecipitated and purified DNA fragments on a microarray with a large variety of small random DNA sequences which further amplifies the annealed sequences providing valuable information about protein binding sites. However, ChIP-seq has emerged as an attractive alternative approach since it provides genome-wide mapping of protein-DNA complexes with higher resolution than ChIP-on-chip and high-throughput genome sequencing. ChIP-seq has revolutionized the field of DNA repair by disclosing DNA binding sites of various transcription factors providing insights into the gene regulation and unravelling chromatin landscapes in a genome-wide scale25. According to this, the field of DNA repair has tremendously profited from ChIP-seq since these data play a crucial role in various diseases and biological pathways, such as cancer progression. Nonetheless, various modifications of the ChIP method have been developed such as HaloChIP, which does not require specific antibody against the protein of interest but rather uses sequences encoding DNA-binding proteins fused with HaloTag, which are transfected to cells and subsequently to crosslinking, the desired protein-DNA complexes can be captured by using HaloLink Resin. However, this technique relies on overexpression and not the endogenous level of the desired proteins which can result in misinterpreted data26.
Furthermore, microscopic techniques provide valuable information about the spatiotemporal tracking of DNA damage repair, even in a single-cell level. The rapid improvement of antibodies raised against specific repair proteins has led to a deeper understanding of the mechanism of NER and DSBR sub-pathways, as well as their post-translational regulation. The microscopic field has been revolutionized by high-resolution techniques, such as super-resolution microscopy, which allows the visualization of DNA damage-induced cellular processes at the nucleosomal level, as well as ensuring the accurate mapping of protein co-localization24. However, it should be noted that the variance in cell lineages must be considered during the experiment as the rate of the repair can diverge, which can make the results difficult to interpret. Considering the rapid evolution of fluorescence imaging methodology and the deliberate design of the experimental setup, a precious opportunity to investigate the DNA damage-induced cellular and molecular responses at a single protein resolution in a single-cell level is on its way to perfection.
In conclusion, the combination of super-resolution microscopy and single-cell sequencing methodology can significantly improve our understanding of the DNA repair field.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This research was funded by the National Research, Development and Innovation Office grant GINOP-2.3.2-15-2016-00020, GINOP-2.3.2-15-2016-00036, GINOP-2.2.1-15-2017-00052, EFOP 3.6.3-VEKOP-16-2017-00009, NKFI-FK 132080, the János Bolyai Research Scholarship of the Hungarian Academy of Sciences BO/27/20, ÚNKP-20-5-SZTE-265, EMBO short-term fellowship 8513, and the Tempus Foundation.
Materials
| 4-OHT | Sigma Aldrich | H7904 | |
| Agarose | Lonza | 50004 | |
| Antibiotic-Antimycotic Solution (100×), Stabilized | Sigma Aldrich | A5955 | |
| Anti-gamma H2A.X (phospho S139) antibody | Abcam | ab26350 | |
| Bovine Serum Fraction V albumin | Biosera | PM-T1725 | |
| TrackIt™ Cyan/Yellow Loading Buffer | Thermo Fisher Scientific | 10482035 | |
| DMEM with 1.0 g/L Glucose, without L-Glutamine | Lonza | 12-707F | |
| Doxycycline | Sigma Aldrich | D9891 | |
| Dynabeads™ M-280 Sheep Anti-Mouse IgG | Invitrogen | 11202D | |
| Dynabeads™ M-280 Sheep Anti-Rabbit IgG | Invitrogen | 11204D | |
| EDTA | Sigma Aldrich | E6758 | |
| EGTA | Sigma Aldrich | E3889 | |
| Ethanol | Molar Chemicals | 02910-101-340 | |
| Fetal Bovine Serum (South America Origin), EU-approved | Gibco | ECS0180L | |
| Formaldehyde 37% solution free from acid | Sigma Aldrich | 1.03999 | |
| GlutaMAX™ Supplement | Thermo Fisher Scientific | 35050038 | |
| Glycine | Sigma Aldrich | 50046 | |
| IPure kit v2 | Diagenode | C03010015 | |
| Isoamyl alcohol | Sigma Aldrich | W205702 | |
| LiCl | Sigma Aldrich | L9650 | |
| NaCl | Sigma Aldrich | S5886 | |
| Na-DOC | Sigma Aldrich | D6750 | |
| NaHCO3 | Sigma Aldrich | S5761 | |
| Neocarzinostatin from Streptomyces carzinostaticus | Sigma Aldrich | N9162 | |
| NP-40 | Sigma Aldrich | I8896 | |
| PBS Powder without Ca2+, Mg2+ | Sigma Aldrich | L182-50-BC | |
| Phenol | Sigma Aldrich | P4557 | |
| PIPES | Sigma Aldrich | P1851 | |
| Polysorbate 20 (Tween 20) | Molar Chemicals | 09400-203-190 | |
| KCl | Sigma Aldrich | P5405 | |
| ProLong™ Gold Antifade Mountant with DAPI | Thermo Fisher Scientific | P36935 | |
| Protease Inhibitor Cocktail Set I | Roche | 11873580001 | |
| Proteinase K | Sigma Aldrich | P2308 | |
| P-S2056 DNAPKcs antibody | Abcam | ab18192 | |
| RNase A | Roche | 10109169001 | |
| CH3COONa | Sigma Aldrich | S2889 | |
| SDS | Sigma Aldrich | L3771 | |
| Tris Acetate-EDTA buffer | Sigma Aldrich | T6025 | |
| Tris-HCl | Sigma Aldrich | 91228 | |
| TRITON X-100 | Molar Chemicals | 09370-006-340 | |
| Trypsin from porcine pancreas | Sigma Aldrich | T4799 | |
| Trypsin-EDTA (0.5%), no phenol red | Gibco | 15400054 |
References
- Borsos, B. N., Majoros, H., Pankotai, T. Ubiquitylation-Mediated Fine-Tuning of DNA Double-Strand Break Repair. Cancers (Basel). 12 (6), (2020).
- Borsos, B. N., Majoros, H., Pankotai, T. Emerging Roles of Post-Translational Modifications in Nucleotide Excision Repair. Cells. 9 (6), (2020).
- Stephens, P. J., et al. The landscape of cancer genes and mutational processes in breast cancer. Nature. 486 (7403), 400-404 (2012).
- Turnbull, C., et al. Gene-gene interactions in breast cancer susceptibility. Human Molecular Genetics. 21 (4), 958-962 (2012).
- Saxowsky, T. T., Meadows, K. L., Klungland, A., Doetsch, P. W. 8-Oxoguanine-mediated transcriptional mutagenesis causes Ras activation in mammalian cells. Proceedings of the National Academy of Sciences of the United States of America. 105 (48), 18877-18882 (2008).
- Jiricny, J. The multifaceted mismatch-repair system. Nature Reviews Molecular Cell Biology. 7 (5), 335-346 (2006).
- Hanawalt, P. C., Spivak, G. Transcription-coupled DNA repair: two decades of progress and surprises. Nature Reviews Molecular Cell Biology. 9 (12), 958-970 (2008).
- Kanaar, R., Wyman, C., Rothstein, R. Quality control of DNA break metabolism: in the ‘end’, it’s a good thing. EMBO Journal. 27 (4), 581-588 (2008).
- Lemaitre, C., et al. Nuclear position dictates DNA repair pathway choice. Genes & Development. 28 (22), 2450-2463 (2014).
- Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annual Review of Biochemistry. 79, 181-211 (2010).
- Lambert, S., Lopez, B. S. Characterization of mammalian RAD51 double strand break repair using non-lethal dominant-negative forms. EMBO Journal. 19 (12), 3090-3099 (2000).
- Xu, S., et al. p300-mediated acetylation of histone demethylase JMJD1A prevents its degradation by ubiquitin ligase STUB1 and enhances its activity in prostate cancer. Cancer Research. , (2020).
- Kastan, M. B., Bartek, J. Cell-cycle checkpoints and cancer. Nature. 432 (7015), 316-323 (2004).
- Roy, R., Chun, J., Powell, S. N. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nature Reviews Cancer. 12 (1), 68-78 (2011).
- Krenning, L., vanden Berg, J., Medema, R. H. Life or Death after a Break: What Determines the Choice. Molecular Cell. 76 (2), 346-358 (2019).
- Rogakou, E. P., Pilch, D. R., Orr, A. H., Ivanova, V. S., Bonner, W. M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. Journal of Biological Chemistry. 273 (10), 5858-5868 (1998).
- Caron, P., et al. WWP2 ubiquitylates RNA polymerase II for DNA-PK-dependent transcription arrest and repair at DNA breaks. Genes & Development. 33 (11-12), 684-704 (2019).
- Caron, P., et al. Cohesin protects genes against gammaH2AX Induced by DNA double-strand breaks. PLoS Genetics. 8 (1), 1002460 (2012).
- Berkovich, E., Monnat, R. J., Kastan, M. B. Assessment of protein dynamics and DNA repair following generation of DNA double-strand breaks at defined genomic sites. Nature Protocols. 3 (5), 915-922 (2008).
- Paques, F., Duchateau, P. Meganucleases and DNA double-strand break-induced recombination: perspectives for gene therapy. Current Gene Therapy. 7 (1), 49-66 (2007).
- Pankotai, T., Bonhomme, C., Chen, D., Soutoglou, E. DNAPKcs-dependent arrest of RNA polymerase II transcription in the presence of DNA breaks. Nature Structural & Molecular Biology. 19 (3), 276-282 (2012).
- Iacovoni, J. S., et al. High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO Journal. 29 (8), 1446-1457 (2010).
- Poinsignon, C., et al. Phosphorylation of Artemis following irradiation-induced DNA damage. European Journal of Immunology. 34 (11), 3146-3155 (2004).
- Varga, D., Majoros, H., Ujfaludi, Z., Erdelyi, M., Pankotai, T. Quantification of DNA damage induced repair focus formation via super-resolution dSTORM localization microscopy. Nanoscale. 11 (30), 14226-14236 (2019).
- Kim, J. A., Kruhlak, M., Dotiwala, F., Nussenzweig, A., Haber, J. E. Heterochromatin is refractory to gamma-H2AX modification in yeast and mammals. Journal of Cell Biology. 178 (2), 209-218 (2007).
- Daniels, D. L., Urh, M. Isolation of intracellular protein–DNA complexes using HaloCHIP, an antibody-free alternative to chromatin immunoprecipitation. Methods in Molecular Biology. 977, 111-124 (2013).

