Bottom-up and Shotgun Proteomics to Identify a Comprehensive Cochlear Proteome
Summary
Proteome analysis of the cochlear sensory epithelium can be challenging due to its small size and because membrane proteins are difficult to isolate and identify. Both membrane and soluble proteins can be identified by combining multiple preparative methods and separation techniques along with high-resolution mass spectrometry.
Abstract
Proteomics is a commonly used approach that can provide insights into complex biological systems. The cochlear sensory epithelium contains receptors that transduce the mechanical energy of sound into an electro-chemical energy processed by the peripheral and central nervous systems. Several proteomic techniques have been developed to study the cochlear inner ear, such as two-dimensional difference gel electrophoresis (2D-DIGE), antibody microarray, and mass spectrometry (MS). MS is the most comprehensive and versatile tool in proteomics and in conjunction with separation methods can provide an in-depth proteome of biological samples. Separation methods combined with MS has the ability to enrich protein samples, detect low molecular weight and hydrophobic proteins, and identify low abundant proteins by reducing the proteome dynamic range. Different digestion strategies can be applied to whole lysate or to fractionated protein lysate to enhance peptide and protein sequence coverage. Utilization of different separation techniques, including strong cation exchange (SCX), reversed-phase (RP), and gel-eluted liquid fraction entrapment electrophoresis (GELFrEE) can be applied to reduce sample complexity prior to MS analysis for protein identification.
Introduction
Proteomics is the study of complex biological systems by analyzing protein expression, function, modifications, and interactions1. Several methods have been utilized for proteome analysis of the inner ear, including antibody microarray2, two-dimensional gel electrophoresis3-5, and DIGE6. However, only a limited number of proteins have been identified and characterized2,7-10, compared to the over 10,000 genes and expressed sequence tags (ESTs) identified in the inner ear11,12, MS is the most commonly used and comprehensive technique in proteomics for protein characterization. Analysis of complex proteomic samples, such as the cochlea, can be challenging. However, the combination of multiple separation techniques with MS enables the identification of a greater number of peptides and proteins, due to an increased dynamic concentration range and peak capacity13. Multidimensional chromatography reduces highly complex protein mixtures by allowing the use of different adsorption mechanisms. There are two commonly used MS proteome analysis approaches, shotgun and bottom-up proteomics. In shotgun proteomics, a mixture of intact proteins is enzymatically digested and separated using multidimensional chromatography with strong cation-exchange chromatography (SCX) followed by reversed-phase liquid chromatography (RPLC)14,15. The separated peptides are subjected to tandem MS and database searching15. A major advantage of this technique is that thousands of proteins can be identified in a single analysis and the technique is better suited to membrane proteins.
In the bottom-up approach, the protein mixture is separated, usually by one- or two-dimensional electrophoresis, and the individual protein bands or spots cut out and digested with an enzyme such as trypsin, usually resulting in multiple peptides. However, another more recently developed electrophoretic approach, used in bottom-up proteomics, is GELFrEE. This technique fractionates protein samples in liquid-phase and makes them less complex prior to analysis. This technique is reproducible, offers high protein recovery, and reduces the distribution of high abundant proteins in complex protein samples16. Peptides, resulting from separated proteins, are analyzed by MS, by using peptide mass fingerprinting or tandem MS (MS/MS), to create sequence tags for database searching17-19. Some of the major advantages of using the bottom-up approach are the ability to obtain high-resolution separations and comprehensive protein coverage. Bottom-up proteomics is the most widely used technique in proteomics20, hence, several bioinformatics tools are available. In addition, proteins can be separated in a complex mixture before digestion, so there is a greater chance of identification.
One of the major challenges in using the inner ear for proteomic analysis is its small size, restricted accessibility, and cell type diversity21. In addition, key proteins that distinguish its functionality, such as ion channels, transporters and receptors, are membrane proteins, which can be difficult to isolate22. Thus, filter-aided sample preparation (FASP) is advantageous for proteomic analyses of tissues that are limited for protein extraction and that require detergents to solubilize membranes23. This filtering allows for the MS analysis of membrane and soluble proteins and for the ability to isolate peptides from low molecular weight contaminants23,24.
The present protocol describes commonly used proteomic approaches that are combined and modified to analyze both soluble and membrane proteins and to maximize the number of protein IDs from the cochlear sensory epithelium. We will describe using shotgun proteomics with FASP multi-digestion, ion exchange chromatography, high resolution MS, and data analysis. In addition, we will describe bottom-up proteomics with GELFrEE, FASP multi-digestion, high resolution MS, and data analysis.
Protocol
Ethics Statement
Experiments using mice tissue were approved by the University of South Florida Institutional Animal Care and Use Committee (Protocols 3931R, 3482R) as set forth under the guidelines of the National Institutes of Health.
1. Protein Extraction
- Isolate cochlear sensory epithelium from 16 30-day-old (P30) CBA/J mice and store at -80 °C.
- On the day of the experiment, wash tissue with 500 µl of 1x phosphate buffered saline (PBS). Centrifuge for 3 min at 1,000 x g, and remove the supernatant. Repeat for a total of three washes.
- Sonicate tissue for 30 sec on ice in 100 μl of lysis buffer containing 50 mM Tris-HCl, pH 8.0, 120 mM NaCl, 50 mM NaF, 5 mM EDTA, 500 μg/ml AEBSF, 10 μg/ml leupeptin, 100 μg/ml pepstatin, 2 μg/ml aprotinin, and 0.5 μg/ml microcystin using a sonic dismembrator (Model 100; Thermo Fisher). Cool lysate on ice for 1 min between each sonication. Sonicate a total of 3x.
- Centrifuge the extract at 750 x g at 4 °C for 2 min and remove the supernatant to a new microtube. Extract the pellet in 50 μl of lysis buffer by sonicating for 30 sec on ice. Centrifuge the extract at 750 x g at 4 °C for 2 min. Combine both lysates and centrifuge at 28,600 x g at 4 °C for 60 min. Remove the supernatant to a new microtube and add 20 μl lysis buffer containing 0.1% ASB-14 to the pellet. Vortex for 1 min and incubate for 60 min at 4 °C.
- Heat the sample at 95 °C for 5 min, then cool at 4 °C for 60 min. Follow with centrifugation at 16,000 x g at 25 °C for 10 min. Collect the supernatant and transfer to a new tube.
- Centrifuge the suspension at 16,000 x g at 4 °C for 5 min and retain the supernatant for digestion.
2. Double Tryptic Protein Digestion of Whole Lysate Using FASP
- Add a 30 μl aliquot (≤400 μg) of cochlear protein extract, containing 4% sodium dodecyl sulfate (SDS), 100 mM Tris-HCl, pH 7.6 and 0.1 M dithiothreitol (DTT) directly to a 30K spin filter and mix with 200 μl of 8 M urea in Tris-HCl. Centrifuge at 14,000 x g for 15 min.
- Dilute the concentrate with 200 μl of 8 M urea solution and centrifuge at 14,000 x g for 15 min.
- Add 10 µl of 10x iodoacetamide (IAA) in 8 M urea solution to the concentrate in the filter and vortex for 1 min. Incubate the spin filter for 20 min at room temperature (RT) in the dark followed by centrifugation at 14,000 x g for 10 min.
- Add 100 µl of 8 M urea solution to the concentrate on the filter unit and centrifuge at 14,000 x g for 15 min. Repeat this step 2x. Add 100 μl of 50 mM ammonium bicarbonate (ABC) solution to the filter unit and centrifuge at 14,000 x g for 10 min. Repeat this step 2x.
- Add 0.4 μg/μl of trypsin in 1:100 (w/w) enzyme-to-protein ratio and incubate overnight (O/N) at 37 °C.
- Following incubation, add 40 μl of 50 mM ABC solution to filter unit and centrifuge at 14,000 x g for 10 min. Repeat this step 1x.
- Add 50 μl of 0.5 M NaCl solution to the spin filter and centrifuge at 14,000 x g for 10 min. Transfer the filtrate containing the tryptic peptides to a fresh microtube and acidify with formic acid (FA) to 1.0% to stop the digestion.
- Wash the filter unit with 40 µl of 8 M urea, and then wash 2x with 40 µl 18 MΩ water.
- Wash the filter unit 3x with 100 μl of 50 mM ABC solution. After the final wash add 0.4 μg/μl of trypsin in 1:100 (w/w) enzyme-to-protein ratio and incubate O/N at 37 °C.
- Elute tryptic peptides from the second digest by adding 40 μl of 50 mM ABC solution to the filter unit and centrifuge at 14,000 x g for 10 min. Repeat this step 1x.
- Add 50 μl of 0.5 M NaCl solution to the filter unit and centrifuge at 14,000 x g for 10 min. Transfer the filtrate containing the tryptic peptides to a fresh microtube and acidify with FA to 1.0%.
- The samples can be quantified using the microplate colorimetric assay using BSA as the standard.
3. Endoproteinase LysC and Tryptic Protein Digestion of Whole Lysate Using FASP
- Add a 30 μl aliquot (≤400 μg) of cochlear protein extract containing 4% SDS, 100 mM Tris-HCl, pH 7.6 and 0.1 M DTT directly to a 30K spin filter and mix with 200 μl of 8 M urea in Tris-HCl. Centrifuge at 14,000 x g for 15 min.
- Dilute the concentrate with 200 μl of 8 M urea solution and centrifuge at 14,000 x g for 15 min.
- Add 10 μl of 10x IAA in 8 M urea solution to the concentrate in the filter and vortex for 1 min. Incubate the spin filter for 20 min at RT in the dark then centrifuge at 14,000 x g for 10 min.
- Add 100 μl of 8 M urea solution to the concentrate on the filter unit and centrifuge at 14,000 x g for 15 min. Repeat this step 2x. Add 100 μl of 100 mM ABC solution to the filter unit and centrifuge at 14,000 x g for 10 min. Repeat this step 2x.
- Add 0.1 μg/μl of endoproteinase LysC in a 1:50 (w/w) enzyme-to-protein ratio and incubate O/N at 30 °C.
- Following incubation, add 40 μl of 100 mM ABC solution to the filter unit and centrifuge at 14,000 x g for 10 min. Repeat this step 1x.
- Add 50 μl of 0.5 M NaCl solution to the spin filter and centrifuge at 14,000 x g for 10 min. Transfer the filtrate containing the LysC peptides to a fresh microtube and acidify with FA to 1.0% to stop the digestion.
- Wash the filter unit with 40 μl of 8 M urea, and then wash 2x with 40 μl 18 MΩ water.
- Wash the filter unit 3x with 100 μl of 50 mM ABC solution. After the final wash add 0.4 μg/μl of trypsin in 1:100 enzyme-to-protein ratio and incubate O/N at 37 °C.
- Elute tryptic peptides by adding 40 μl of 50 mM ABC solution to the filter unit and centrifuge at 14,000 x g for 10 min. Repeat this step 1x.
- Add 50 µl of 0.5 M NaCl solution to the filter unit and centrifuge at 14,000 x g for 10 min. Transfer the filtrate containing the tryptic peptides to a fresh microtube and acidify with 1.0% FA.
- The sample can be quantified using the microplate colorimetric assay with BSA as the standard.
4. Desalting Peptides Using Spin Columns
- Activate a C18 MacroSpin column by adding 500 µl of acetonitrile (ACN) and centrifuge at 1,100 x g for 1 min. Discard the flow-through after centrifugation.
- Equilibrate the column with 500 μl of 0.1% FA and centrifuge at 1,100 x g for 1 min. Discard the flow-through and repeat this step 1x.
- Add up to 500 μl of the peptide digest to the column and centrifuge at 1,100 x g for 1 min. If the sample volume is greater than 500 µl then repeat this step.
- Wash the column with 500 μl of 0.1% FA and centrifuge at 1,100 x g for 1 min. Discard the flow-through. Repeat this step 1x.
- Add 250 µl of a 90:10 ACN-to-water ratio to the column and centrifuge at 1,100 x g for 1 min. Collect the eluent containing the desalted peptides and transfer to a fresh microtube. Repeat this step 1x.
- Dry the desalted peptide sample in a vacuum centrifuge and avoid letting the sample dry completely.
5. Ion Exchange Chromatography
- Inject 50-100 μg of digested protein sample onto a 200 x 2.1 mm, 5 μm SCX column (Polysulfoethyl A) to separate peptides.
- Use a gradient of 2-40% B over 50 min with a flow rate of 250 μl/min. Solvent A is 5 mM ammonium formate, pH 3.0 in 25% acetonitrile and 75% ddH2O. Solvent B is 500 mM ammonium formate, pH 6.0 in 25% ACN and 75% ddH2O.
- Monitor the peptide fractions at 280 nm and collect the fractions in 2 min intervals using a fraction collector.
- Dry fractions in a vacuum concentrator and resuspend in 500 μl of 50% acetonitrile in ddH2O containing 5% FA to assist with salt removal.
- Redry fractions and store at -80 °C until ready to use for nano LC-MS/MS analysis.
6. Acetone Precipitation
Prior to GELFrEE separation the cochlear protein supernatant has to be desalted. Acetone precipitation can be used to desalt and concentrate proteins.
- Add three volumes of ice-cold acetone to the cochlear supernatant. Gently vortex and incubate at -20 °C O/N.
- Centrifuge the sample mixture at 15,000 x g for 15 min at 4 °C and remove the supernatant.
- Wash the protein pellet 3x with chilled acetone and centrifuge at 14,000 x g for 5 min at 4 °C.
- Air-dry the pellet and dissolve in 112 μl of 18 MΩ water.
- Determine the protein concentration using the microplate colorimetric assay.
7. GELFrEE Fractionation of Cochlear Sensory Epithelium
- Add 30 μl of 5x sample buffer (0.25 M Tris-HCl pH 6.8, 10% (w/v) SDS, 50% glycerol, 0.5% (w/v) bromophenol blue) to the desalted protein suspended in 112 μl of 18 MΩ water.
- Add 8 μl of 1M DTT and heat at 95 °C for 5 min to reduce protein disulfide bonds. After heating allow cooling to room temperature.
- Add 8 ml of running buffer to anode reservoir, 150 μl of running buffer to sample collection chamber, and 6 ml of running buffer to cathode reservoir.
- Remove any buffer that may have flowed into the sample-loading chamber using a pipette.
- Load 150 μl (≤1 mg) of the cochlear protein mixture, into the sample-loading chamber using an 8% Tris-acetate cartridge with a mass range of 3.5-150 kDa.
- Lower electrodes and close instrument lid to start the run.
- At the first paused time on the instrument add 2 ml of running buffer to the cathode reservoir and resume run.
- At each paused interval on the instrument, collect the sample from the sample collection chamber using a pipette and add to a freshly labeled microtube. Wash the collection chamber 2x with 150 μl of running buffer and discard.
- Add 150 μl of fresh running buffer to the sample collection chamber and resume run. Collect the protein fractions over a time period of 2.6 hr in a total volume of 150 μl/fraction.
8. 1D Gel Electrophoresis of GELFrEE Fractions
1D gel electrophoresis can be used to visualize the results from GELFrEE fractionation prior to enzymatic digestion and MS analysis. GELFrEE protein fractions can be separated on a 4-15% Tris-HCl gel.
- Mix a 5 μl aliquot of each GELFrEE fraction with 5 µl of sample diluting buffer (350 mM DTT in sample buffer).
- Heat the samples at 95 °C for 3 min.
- Cool samples to RT.
- Load 10 μl of each GELFrEE fraction in individual gel lanes and load 5 μl of molecular weight standard in the last unused lane.
- Run the gel at 125 V for 1.5 hr or until the dye front reaches the bottom of the gel.
- Carefully remove the gel and stain with Silver Stain Plus to visualize the protein separation.
9. Protein Digestion of GELFrEE Fractions Using FASP
A modified FASP procedure is used for detergent removal and digestion of the GELFrEE fractions.
- Add each individual fraction directly to a 30K filter unit; mix with 200 μl of 8 M urea in Tris-HCl and centrifuge at 14,000 x g for 25 min.
- Dilute the concentrate with 200 μl of urea solution and centrifuge at 14,000 x g for 12 min. Repeat this step 1x.
- Add 10 μl of 10x IAA in urea solution to the concentrate in each filter unit, vortex for 1 min, and incubate for 30 min at RT in the dark.
- Add 100 μl of urea solution to the concentrate on the filter units and centrifuge at 14,000 x g for 15 min. Repeat this step 2x. Add 100 μl of 100 mM ABC solution to the filter units and centrifuge at 14,000 x g for 10 min. Repeat this step 2x.
- Add 0.1 μg/μl of LysC for a 1:50 (w/w) enzyme-to-protein ratio to the filter units and incubate O/N at 30 °C.
- Following incubation, add 40 μl of 100 mM ABC solution to the spin filters and centrifuge at 14,000 x g for 10 min. Repeat this step 1x.
- Add 50 μl of 0.5 M NaCl solution to the spin filters and centrifuge at 14,000 x g for 10 min. Transfer the filtrate containing the LysC peptides from each spin filter to a fresh microtube and acidify with trifluoroacetic acid (TFA).
- Wash the spin filters with 40 μl of 8 M urea, then wash 2x with 40 μl 18 MΩ water.
- Wash the spin filters 3x with 100 μl of 50 mM ABC solution. After the final wash add 0.4 μg/μl of trypsin in a 1:100 (w/w) enzyme-to-protein ratio and incubate O/N at 37 °C.
- Elute tryptic peptides from each filter unit by adding 40 μl of 50 mM ABC solution and centrifuge at 14,000 x g for 10 min. Repeat this step 1x.
- Add 50 μl of 0.5 M NaCl solution to the filter units and centrifuge at 14,000 x g for 10 min. Transfer the filtrate containing the tryptic peptides from each filter unit to a fresh microtube and acidify with TFA.
- Dry all digests in a vacuum concentrator.
10. Sample Preparation for LC-MS/MS
- Reconstitute dried LysC and tryptic peptide digests fractions in 20 µl of 0.1% FA and vortex.
- Centrifuge samples at 20,000 x g for 10 min and remove the top 95% of sample to a new sample vial.
- Inject 5 µl of each peptide fraction onto a 100 μm x 25 mm sample trap to remove salts and contaminants and perform chromatographic separation on a 75 μm × 10 cm C18 column.
- Use a gradient of 2-40% B over 100 min with a flow rate of 200 nl/min. Solvent A is 95% ddH2O and 5% acetonitrile containing 0.1% FA. Solvent B is 80% ACN and 20% ddH2O containing 0.1% FA.
- Collect ten tandem mass spectra for each MS scan on a high-resolution mass spectrometer. An LTQ Orbitrap mass spectrometer was used in this experiment.
11. Protein Identification
- Identify proteins using the UniProt mouse database containing both forward and reverse protein sequences and common contaminants. MaxQuant software with MASCOT search engine is used to search the protein database.
- The search parameters that can be used include; fixed modification of carbamidomethyl of cysteine, variable modifications of oxidation of methionine, protein N-terminal acetylation, maximum of 2 missed cleavage. The minimum peptide length to consider for identification is 6 amino acids. Enter a peptide mass concentration error of ±8 ppm and a fragment mass tolerance of 1.2 Da (mass error is dependent on the mass spectrometer being used).
- In the MaxQuant software, use a 1% false discovery rate (FDR) for peptide and protein identification. Protein identification is listed with the number of matched peptides, the percent protein sequence coverage, and the peptide sequences.
Representative Results
To obtain the most comprehensive proteome of the cochlear sensory epithelium, quick tissue dissection is required prior to protein extraction and sample preparation. Two proteomic techniques can be used, shotgun and bottom-up proteomics. To prepare samples for shotgun proteomics, FASP digestion procedure was used as illustrated in Figure 1. The FASP method allows for concentration of proteins, removal of detergents, and digestion of proteins using multiple enzymes. There were two double digestion procedures used, the first was a tryptic digestion followed by a second digestion with trypsin, which were pooled, fractionated on a SCX column into 18 fractions and analyzed by nano LC-MS/MS. A total of 1,485 proteins were identified with a 1% FDR when performing a single-run LC-MS/MS using this experimental approach. Among the identified proteins, 329 and 258 proteins were categorized in mitochondrion and plasma membrane, respectively (Figure 2A). The second double digestion procedure consisted of LysC digestion followed by trypsin digestion. Each digest was individually loaded and separated on the SCX column into 18 fractions and analyzed by nano LC-MS/MS. The results of the LysC and trypsin digestions produced a total of 3,503 proteins with a 1% FDR. Figure 2B shows that 605 and 617 proteins were categorized in mitochondrion and plasma membrane, respectively. This approach provided the largest number of membrane-associated protein IDs. Duplicate analysis of the LysC and trypsin fractions showed more than 65% of the proteins identified were shared between the experiments. However, there were also newly identified proteins in the replicate analysis. The additional peptides and proteins were identified due to small changes in chromatography, therefore leading to different peptides for fragmentation25.
Bottom-up proteomics was applied using GELFrEE fractionation prior to LC-MS/MS as illustrated in Figure 3. There were 12 GELFrEE fractions collected. Prior to digestion and LC-MS/MS analysis, a silver-stained gel was prepared to visualize the results from GELFrEE fractionation as illustrated in Figure 4. The gel showed protein separation by increasing molecular weight for each consecutive fraction. Therefore, the 12 GELFrEE liquid fractions were digested using two different multi-FASP digestion approaches and analyzed by LC-MS/MS. The first digestion approach was performed using a double trypsin digestion, which led to the identification of 2,165 proteins with a 1% FDR when performing a single-run LC-MS/MS. Figure 5A shows that there were 516 and 399 proteins categorized in mitochondrion and plasma membrane, respectively. The second digestion approach was performed using endoproteinase LysC followed by trypsin digestion. Single-run LC-MS/MS analysis identified 2,211 proteins with a 1% FDR. This approach showed a similar number of membrane-associated proteins as when using the trypsin/trypsin approach (Figure 5B). The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium26 with the data set identifier PXD00023125. Combining the results from the different techniques resulted in a large number of membrane and soluble proteins from the mouse cochlea sensory epithelium (Figure 6).
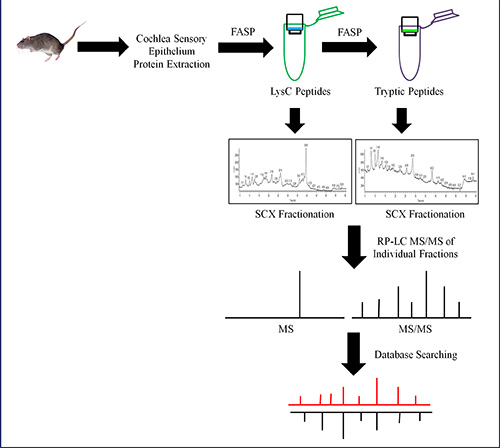
Figure 1. Schematic of a shotgun proteomic experiment using FASP, ion exchange chromatography, and high-resolution MS. Proteins are extracted, solubilized, and digested using FASP with LysC and trypsin endoproteinases. The LysC (green tube) and tryptic peptides (purple tube) are separated into less complex fractions using SCX chromatography and analyzed using nano LC-MS/MS. The MASCOT search engine was used to process the MS data for protein identification. Click here to view larger image.
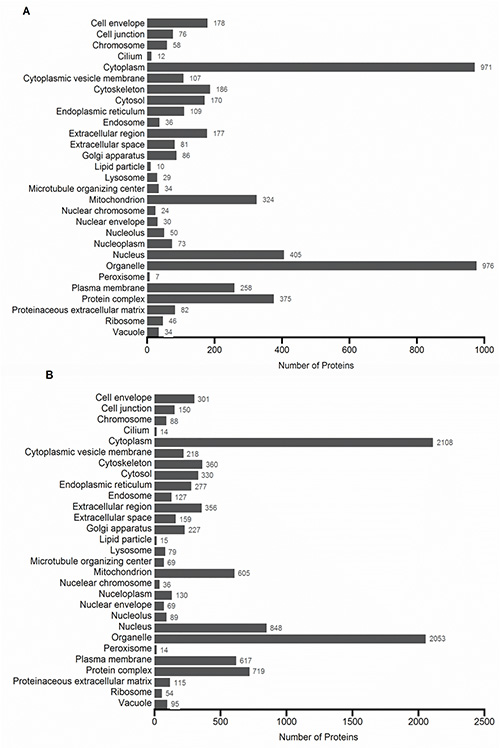
Figure 2. GO cellular components profile of protein IDs when performing a (A) first and second digestion with trypsin followed by SCX separation and (B) first digestion with LysC followed by a second digestion with trypsin followed by SCX separation. All categories are counted nonexclusively, when a protein has more than one category for cellular components. Click here to view larger image.
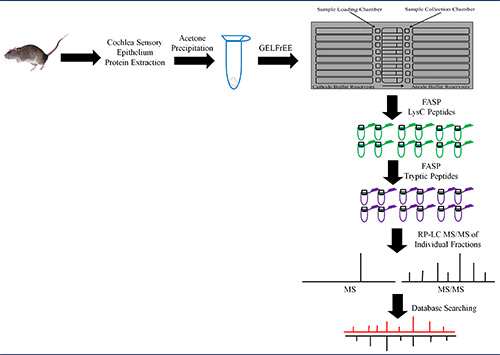
Figure 3. Schematic of a bottom-up proteomic experiment using GELFrEE, FASP, and high-resolution MS. Extracted proteins are solubilized and proteins are precipitated using acetone (blue tube) to remove salts and unwanted contaminants from the lysate. The protein pellet is solubilized and proteins fractionated using GELFrEE. Each fraction was digested using FASP with LysC and trypsin endoproteinases. The LysC (green tubes) and tryptic peptides (purple tubes) from each fraction are analyzed using nano LC-MS/MS and proteins identified by searching MS data using MASCOT. Click here to view larger image.
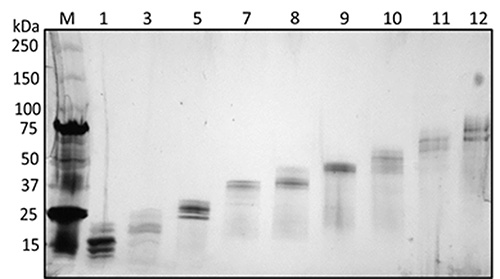
Figure 4. Silver-stained gel of cochlear sensory epithelium GELFrEE fractions, to visualize protein separation in each fraction prior to MS analysis. (M) Protein marker, (1) Fraction 1, (3) Fraction 2, (5) Fraction 5, (7) Fraction 7, (8) Fraction 8, (9) Fraction 9, (10) Fraction 10, (11) Fraction 11, (12) Fraction 12. Reprinted (adapted) with permission from Darville and Sokolowski24. Copyright 2013 by the American Chemical Society. Click here to view larger image.
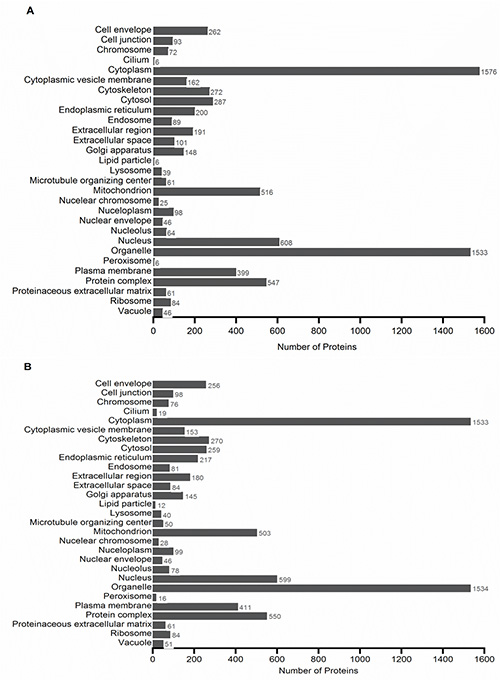
Figure 5. GO cellular components profile of protein IDs when performing a (A) first and second digestion with trypsin after GELFrEE separation and (B) first digestion with LysC followed by a second digestion with trypsin after GELFrEE separation. All categories are counted nonexclusively, when a protein has more than one category for cellular components. Click here to view larger image.
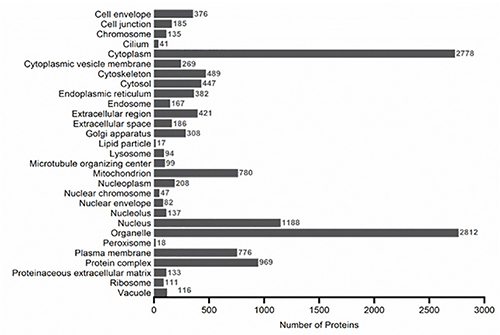
Figure 6. GO cellular components profile for all the proteins identified using SCX, WAX, or GELFrEE separation. When a protein had more than one category for cellular components, all of its categories were counted nonexclusively. Click here to view larger image.
Discussion
The key steps to maximizing protein identification from the cochlear sensory epithelium are: 1) use of multiple endoproteinases for digestion, 2) use of multiple separation techniques, and 3) utilization of a high-resolution mass spectrometer. The application of multiple enzymes increases the number of peptides and improves protein sequence coverage, hence improving the number of identified proteins from the cochlear tissue. Trypsin, the most commonly used protease provides efficient and specific cleavage of proteins, generating peptides that are good for MS ionization and fragmentation. However, using another enzyme prior to trypsin, such as LysC, which also cleaves at lysine residue, provides more efficient peptide cleavage. It was observed that the sequence coverage of the proteins generated from the LysC/trypsin digestion showed an increase in protein sequence coverage relative to the protein sequence coverage from the trypsin/trypsin digest.
Implementation of multidimensional separation prior to MS reduced the high cochlear sample complexity in the shotgun proteomic approach. This allowed for identification of more proteins, including membrane proteins, which are typically difficult to identify. A larger number of membrane proteins were identified using the shotgun approach as opposed to the bottom-up approach. However, the bottom-up approach using GELFrEE allowed for identification of more low abundant proteins. This is due to the isolation of higher abundant proteins, such as cochlin from the cochlea, into individual fractions.
The high quality data achieved in the present protocol by a high-resolution mass spectrometer, such as an LTQ-Orbitrap, improves and increases the number of proteins that can be identified in the cochlea. The LTQ-Orbitrap offers high resolution, high mass accuracy, and high sensitivity for the analysis of peptides. Hence, application of this instrument enables identification of proteins from complex biological samples, such as the cochlear sensory epithelium and enhances identification of low abundant peptides. Utilization of these combined experimental approaches with the powerful tool of MS helps to significantly increase the cochlear protein dataset, therefore improving the opportunity to identify novel protein biomarkers involved in hearing and deafness.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors thank Dr. Kent Seeley, Director of The Center for Drug Discovery and Innovation (CDDI) Proteomics Core Facility at University of South Florida for the use of this facility. This work was supported by NIH/NIDCD grant R01 DC004295 to B.H.A.S.
Materials
| 8% Tris-acetate cartridge | Protein Discovery | 42103 | |
| Acetone | Sigma-Aldrich | 179124 | |
| Acetonitrile | Honeywell | 015-1L | |
| AEBSF | Calbiochem | 101500 | |
| Ammonium formate | Fisher Scientific | AC16861 | |
| Aprotinin | Calbiochem | 616370 | |
| ASB-14 | Calbiochem | 182750-5GM | |
| Bovine serum albumin | BioRad | 500-0112 | |
| C18 column | New Objective | A25112 | 75 μm × 10 cm |
| DC Protein Assay | BioRad | 500-0116 | Microplate Assay Protocol |
| EDTA | Sigma-Aldrich | E9884 | |
| Endoproteinase Lys-C | Sigma-Aldrich | P3428 | |
| FASP Protein Digestion Kit | Protein Discovery | 44250 | |
| Formic acid | Fluka | 94318 | |
| GELFrEE Fractionation System | Protein Discovery | 42001 | GELFrEE 8100 |
| Leupeptin | Calbiochem | 108975 | |
| MacroSpin Column | The Nest Group | SMM SS18V | Silica C18 |
| Microcystin | Calbiochem | 475815 | |
| Pepstatin | Sigma-Aldrich | P5318 | |
| Polysulfoethyl A Column | The Nest Group | 202SE0503 | |
| Sodium dodecyl sulfate (SDS) | Sigma-Aldrich | L3771 | |
| Sonic Dismembrator | Thermo Fisher | 15-338-53 | Model 100 |
| Trypsin | Sigma-Aldrich | T6567 | Proteomics Grade |
References
- Domon, B., Aebersold, R. Review – Mass spectrometry and protein analysis. Science. 312, 212-217 (2006).
- Jamesdaniel, S., Hu, B., Kermany, M. H., Jiang, H., Ding, D., Coling, D., Salvi, R. Noise induced changes in the expression of p38/MAPK signaling proteins in the sensory epithelium of the inner ear. J. Proteomics. 75, 410-424 (2011).
- Thalmann, I., Hughes, I., Tong, B. D., Ornitz, D. M., Thalmann, R. Microscale analysis of proteins in inner ear tissues and fluids with emphasis on endolymphatic sac, otoconia, and organ of Corti. Electrophoresis. 27, 1598-1608 (2006).
- Thalmann, I., Rosenthal, H. L., Moore, B. W., Thalmann, R. Organ of Corti-specific polypeptides: OCP-I and OCP-II. J. Acoust. Soc. Am. 67, (1980).
- Kathiresan, T., Harvey, M., Sokolowski, B. . The use of two-dimensional gels to identify novel protein-protein interactions in the cochlea. In Auditory and vestibular research : methods and protocols. , (2009).
- Zheng, Q. Y., Rozanas, C. R., Thalmann, I., Chance, M. R., Alagramam, K. N. Inner ear proteomics of mouse models for deafness, a discovery strategy. Brain Res. 1091, 113-121 (2006).
- Peng, H., Liu, M., Pecka, J., Beisel, K. W., Ding, S. J. Proteomic Analysis of the Organ of Corti Using Nanoscale Liquid Chromatography Coupled with Tandem Mass Spectrometry. Int. J. Mol. Sci. 13, 8171-8188 (2012).
- Elkan-Miller, T., Ulitsky, I., Hertzano, R., Rudnicki, A., Dror, A. A., Lenz, D. R., Elkon, R., Irmler, M., Beckers, J., Shamir, R., Avraham, K. B. Integration of Transcriptomics, Proteomics, and MicroRNA Analyses Reveals Novel MicroRNA Regulation of Targets in the Mammalian Inner Ear. Plos One. 6, (2011).
- Yang, Y., Dai, M., Wilson, T. M., Omelchenko, I., Klimek, J. E., Wilmarth, P. A., David, L. L., Nuttall, A. L., Gillespie, P. G., Shi, X. Na+/K+-ATPase α1Identified as an Abundant Protein in the Blood-Labyrinth Barrier That Plays an Essential Role in the Barrier Integrity. . Plos One. 6, (2011).
- Shin, J. B., Streijger, F., Beynon, A., Peters, T., Gadzala, L., McMillen, D., Bystrom, C., Vander Zee, C. E., Wallimann, T., Gillespie, P. G. Hair Bundles Are Specialized for ATP Delivery via Creatine Kinase. Neuron. 53, 371-386 (2007).
- Chen, Z. Y., Corey, D. P. An inner ear gene expression database. J. Assoc. Res. Otolaryngol. 3, 140-148 (2002).
- Gabashvili, I. S., Sokolowski, B. H., Morton, C. C., Giersch, A. B. Ion channel gene expression in the inner ear. J. Assoc. Res. Otolaryngol. 8, 305-328 (2007).
- Horvatovich, P., Hoekman, B., Govorukhina, N., Bischoff, R. Multidimensional chromatography coupled to mass spectrometry in analysing complex proteomics samples. J. Sep. Sci. 33, 1421-1437 (2010).
- Ye, M. L., Jiang, X. G., Feng, S., Tian, R. J., Zou, H. F. Advances in chromatographic techniques and methods in shotgun proteome analysis. Trends Anal. Chem. 26, 80-84 (2007).
- Wu, C. C., MacCoss, M. J. Shotgun proteomics: Tools for the analysis of complex biological systems. Curr. Opin. Mol. Ther. 4, 242-250 (2002).
- Bora, A., Anderson, C., Bachani, M., Nath, A., Cotter, R. J. Robust Two-Dimensional Separation of Intact Proteins for Bottom-Up Tandem Mass Spectrometry of the Human CSF Proteome. J. Proteome Res. 11, 3143-3149 (2012).
- Chait, B. T. Mass spectrometry: Bottom-up or top-down. Science. 314, 65-66 (2006).
- Aebersold, R., Mann, M. Mass spectrometry-based proteomics. Nature. 422, 198-207 (2003).
- Reid, G. E., McLuckey, S. A. Top down’ protein characterization via tandem mass spectrometry. J. Mass. Spectrom. 37, 663-675 (2002).
- Han, X., Aslanian, A., Yates, J. R. Mass spectrometry for proteomics. Curr. Opin. Chem. Biol. 12, 483-490 (2008).
- Thalmann, I. Inner ear proteomics: A fad or hear to stay. Brain Res. 1091, 103-112 (2006).
- Lang, F., Vallon, V., Knipper, M., Wangemann, P. Functional significance of channels and transporters expressed in the inner ear and kidney. Am. J. Physiol. Cell Physiol. 293, (2007).
- Wisniewski, J. R., Zougman, A., Nagaraj, N., Mann, M. Universal sample preparation method for proteome analysis. Nat. Meth. 6, 359-362 (2009).
- Wisniewski, J. R., Zielinska, D. F., Mann, M. Comparison of ultrafiltration units for proteomic and N-glycoproteomic analysis by the filter-aided sample preparation method. Anal. Biochem. 410, 307-309 (2011).
- Darville, L. N., Sokolowski, B. H. In-depth Proteomic Analysis of Mouse Cochlear Sensory Epithelium by Mass Spectrometry. J Proteome Res. 12 (8), 3620-3630 (2013).
- Vizcaino, J. A., Cote, R. G., Csordas, A., Dianes, J. A., Fabregat, A., Foster, J. M., Griss, J., Alpi, E., Birim, M., Contell, J., O’Kelly, G., Schoenegger, A., Ovelleiro, D., Perez-Riverol, Y., Reisinger, F., Rios, D., Wang, R., Hermjakob, H. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 41, 1063-1069 (2013).

