From Cells to Cell-Free Protein Synthesis within 24 Hours Using Cell-Free Autoinduction Workflow
Summary
This work describes the preparation of cell extract from Escherichia coli (E. coli) followed by cell-free protein synthesis (CFPS) reactions in under 24 hours. Explanation of the cell-free autoinduction (CFAI) protocol details improvements made to reduce researcher oversight and increase quantities of cell extract obtained.
Abstract
Cell-free protein synthesis (CFPS) has grown as a biotechnology platform that captures transcription and translation machinery in vitro. Numerous developments have made the CFPS platform more accessible to new users and have expanded the range of applications. For lysate based CFPS systems, cell extracts can be generated from a variety of organisms, harnessing the unique biochemistry of that host to augment protein synthesis. Within the last 20 years, Escherichia coli (E. coli) has become one of the most widely used organisms for supporting CFPS due to its affordability and versatility. Despite numerous key advances, the workflow for E. coli cell extract preparation has remained a key bottleneck for new users to implement CFPS for their applications. The extract preparation workflow is time-intensive and requires technical expertise to achieve reproducible results. To overcome these barriers, we previously reported the development of a 24 hour cell-free autoinduction (CFAI) workflow that reduces user input and technical expertise required. The CFAI workflow minimizes the labor and technical skill required to generate cell extracts while also increasing the total quantities of cell extracts obtained. Here we describe that workflow in a step-by-step manner to improve access and support the broad implementation of E. coli based CFPS.
Introduction
The use of cell-free protein synthesis (CFPS) for biotechnology applications has grown substantially over the past few years1,2,3. This development can be attributed in part to increased efforts in understanding the processes that occur in CFPS and the role of each component4,5. Additionally, reduced costs attributed to optimized set-ups and alternative energy sources have made cell-free technology easier to implement for new users6,7,8,9. In order to implement the necessary transcription and translation factors for protein synthesis, cell extract is often used to drive cell-free reactions10. Recently published user guides have provided simple protocols for producing functional extract, making it easier to implement for new and experienced users alike1,11,12,13,14. Cell extract is usually obtained through the lysis of a cell culture, which can be grown using different organisms depending on the specific use desired1,15,16.
Escherichia coli (E. coli) has rapidly become one of the most commonly used host organisms for producing functional extracts17. The BL21 Star (DE3) strain is preferred because it removes the proteases from the outer membrane (OmpT protease) and the cytoplasm (Lon protease), providing an optimal environment for the recombinant protein expression. Additionally, the DE3 contains the λDE3 that carries the gene for T7 RNA polymerase (T7 RNAP) under the control of the lacUV5 promoter; the star component contains a mutated RNaseE gene which prevents cleavage of mRNA4,14,18,19. Under the lacUV5 promoter, isopropyl-thiogalactopyranoside (IPTG) induction allows the expression of T7 RNAP20,21. These strains are used to grow and harvest cells, which give raw material for extract preparation. Cell lysis can be performed using a variety of methods, including bead beating, French press, homogenization, sonication, and nitrogen cavitation1,11,12,22.
The process of bacterial culture and harvesting is consistent across most platforms when using E. coli, but requires multiple days and intense researcher oversight1,11,13. This process generally starts with an overnight seed culture in LB broth, which upon overnight growth is then inoculated into a larger culture of 2xYTPG (yeast, tryptone, phosphate buffer, glucose) the next day. The growth of this larger culture is monitored until it reaches the early-to-mid log phase, at an optical density (OD) of 2.514,20. Constant measurement is required as the components of transcription and translation have been previously demonstrated to be highly active in the early-to-mid log phase23,24. While this process can create reproducible extract, our lab has recently developed a new method using Cell-Free Autoinduction (CFAI) Media, which reduces researcher oversight, increases the overall yield of extract for a given liter of cell culture, and improves access to E. coli-based extract preparation for both experienced and new users (Figure 1). Here we provide the step-by-step guide for implementing the CFAI workflow, to go from a streaked plate of cells to a completed CFPS reaction within 24 hours.
Protocol
1. Media growth
- Prepare 960 mL of CFAI media as described in Table 1 and adjust the pH to 7.2 using KOH.
- Transfer culture media to a 2.5 L baffled flask and autoclave for 30 min at 121 °C.
- Prepare a 40 mL sugar solution as described in Table 1. Filter-sterilize the solution into a separate autoclaved glass container.
NOTE: The sugar solution can be stored in a 30 °C incubator until further use. - Allow the media to completely cool to below 40 °C after autoclaving.
- Prior to inoculation of the CFAI media, add the sugar solution directly to the CFAI media.
- To inoculate the media, swipe a loopful of colonies from a previously streaked E. coli BL21 Star (DE3) plate and insert directly into the media. Swirl the loop into the media but avoid touching the sides of the container. Ensure that the streak plate is fresh, with viable cells.
- Place the flask with the inoculated media in a 30 °C incubator while shaking at 200 rpm. Allow the culture to grow overnight. If cells are inoculated in the evening, the culture will reach an approximate optical density, measured at 600 nm (OD600), of 10 the subsequent morning. Inoculation and harvest times can be adjusted as needed.
2. Cell harvest
- Prepare S30 buffer in advance and keep it cold. Prepare the S30 buffer according to Table 2, to a final pH of 8.2.
NOTE: The S30 buffer can be prepared days prior to the cell harvest. If this is done, prepare without dithiothreitol and store at 4 °C. Add dithiothreitol just before use. - From this point forward, keep all solutions and materials on ice. Transfer 1 L of media into a 1 L centrifuge bottle and centrifuge at 5,000 x g between 4-10 °C for 10 min. Decant and dispose of the supernatant. Using a sterile spatula, transfer the pellet to a pre-chilled and previously weighed 50 mL conical tube.
- Wash once with 30-40 mL of cold S30 buffer by resuspending the pellet via vortexing in 30 s bursts with rest periods on ice.
NOTE: Due to a higher volume of pellet, splitting the cell pellet into two 50 mL conical tubes can be helpful for the wash step. Storing cells in smaller aliquots also provides flexibility for the downstream processing. - Centrifuge the cell resuspension at 5000 x g between 4-10 °C for 10 min.
- Dispose of the supernatant and wipe any excess from the inside walls of the 50 mL conical tube using a clean tissue, avoid touching the pellet itself. Weigh and flash freeze the pellet in liquid nitrogen. Store at -80 °C until further use.
NOTE: Pellets may not require flash-freezing if the user plans to continue with the extract preparation protocol.
3. Extract preparation
- Combine the frozen pellet with 1 mL of S30 buffer for every 1 g of the cell pellet and allow to thaw on ice for approximately 30-60 min. Resuspend thawed pellet via vortexing in bursts of 30 s with rest periods on ice. Vortex until there are no visible clumps of cells remaining.
NOTE: Smaller clumps can be resuspended by mixing using a pipette. - Transfer aliquots of 1.4 mL of cell resuspension into 1.5 mL microfuge tubes for cell lysis. Sonicate each tube with a frequency of 20 kHz and 50% amplitude for three bursts of 45 s with 59 s of rest per cycle surrounded by an ice bath. Invert the tube between cycles and immediately add 4.5 µL of 1 M dithiothreitol after the last sonication cycle.
NOTE: Due to the heat released from sonication, it is extremely important to make sure all aliquots are kept on ice when not being sonicated. The ice bath should be constantly replenished or large enough to stay cool throughout the entire sonication process. - Centrifuge each tube at 18,000 x g and 4 °C for 10 min. Retrieve the supernatant and aliquot into 1.5 mL microfuge tubes in 600 µL aliquots. Flash freeze aliquots and store at -80 °C until further use. Care should be taken to pipette only the supernatant.
4. Cell-free protein synthesis
- Thaw one aliquot of extract from the previous step to perform 15 µL of cell-free protein synthesis reactions in 1.5 mL microfuge tubes in quadruplicate.
- Prepare each reaction by combining 240 ng of DNA (16 µg/mL final concentration), 2.20 µL of Solution A, 2.10 µL of Solution B, 5.0 µL of extract, and a varying volume of molecular-grade water to fill the reaction to 15 µL. This reaction can be scaled to higher volumes. See Table 3 for ratios.
- Prepare Solution A and B according to Table 4. Each solution can be prepared in batches of 100 µL to 1 mL, aliquoted, and stored at -80 °C until further use.
NOTE: The DNA amount can vary depending on the protein of interest. In this case, the plasmid used, pJL1-sfGFP, has been optimized to perform at 16 µg/mL, or 597 µM.
- Prepare Solution A and B according to Table 4. Each solution can be prepared in batches of 100 µL to 1 mL, aliquoted, and stored at -80 °C until further use.
- Let the reactions run for at least 4 h at 37 °C.
5. Quantification of reporter protein, super folder green fluorescent protein (sfGFP)
- Using a half area 96-well black polystyrene plate, combine 2 µL of each cell-free protein synthesis reaction product with 48 µL of 0.05 M HEPES buffer at pH 7.2. Three to four replicates of each reaction tube are recommended.
- Quantify the fluorescence intensity of the sfGFP with an excitation wavelength of 485 nm and an emission wavelength of 528 nm.
- For conversion of relative fluorescence units to volumetric yield (µg/mL) of sfGFP, establish a standard curve using purified pJL1-sfGFP.
Representative Results
When preparing CFAI media, glucose was exchanged for an increase in lactose and glycerol as the main energy substrate in the media. Additionally, the buffering capacity of the CFAI media was increased as well. These specific components are given in Table 1.
The cells were then grown to both an OD600 of 10 and the standard 2.5 in CFAI media to show consistency with extract quality despite varying extract quantities. The 2.5 OD600 CFAI media was grown after inoculating from a seed culture in LB broth at 37 °C, 200 rpm, while the OD600 10 culture was inoculated directly from a plate. Each batch of CFAI media was then monitored and harvested at their respective OD600. The growth to an OD600 of 10 led to an increase in higher amount of cell pellet and overall extract obtained, as it produced 9.60 mL of extract versus the 2.10 mL of extract obtained from the growth to 2.5 OD600 (Figure 2). Further analysis of total protein concentration demonstrated no significant difference in overall protein in each extract (Table 5). Even though they were grown to different levels of optical density, both batches of extract demonstrated similar results in cell-free reactions using sfGFP (Figure 3). This suggests that the combination of the increased buffering capacity, the use of lactose and glycerol as the main carbon source and implementing lactose instead of IPTG for T7RNAP induction help stabilize extract growths to any OD600 below 10.
| Autoclaved CFAI Media: | |
| Components | Amount |
| Sodium Chloride | 5.0 g |
| Tryptone | 20.0 g |
| Yeast Extract | 5.0 g |
| Potassium Phosphate, monobasic | 6.0 g |
| Potassium Phosphate, dibasic | 14.0 g |
| NanopureTM Water | Fill to a total of 960 mL |
| Filter Sterilized Sugar Solution: | |
| Components | Amount |
| D-Glucose | 0.50 g |
| D-Lactose | 4.0 g |
| 80% v/v Glycerol | 7.5 mL |
| NanopureTM Water | 28.0 mL |
Table 1: CFAI components. Components for CFAI media and sugar solutions with their respective amounts. The media should be stirred throughout the addition of each component and the sugar solution filter sterilized. Each solution should be added to a separate sterile container prior to inoculation.
| S30 Buffer | |
| Components | Concentration |
| Tris Acetate pH 8.2 at room temperature | 10 mM |
| Magnesium Acetate | 14 mM |
| Potassium Acetate | 60 mM |
| Dithiothreitol | 2 mM |
Table 2: S30 buffer components: Components for S30 buffer were added with their respective amounts into a sterile 50 mL conical tube.
| Component | Amount |
| Solution A | 2.20 μL |
| Solution B | 2.1 μL |
| Extract | 5 μL |
| DNA Template | Volume for 16 μg/mL final |
| Water | Fill to a total of 15 μL |
Table 3: CFPS reaction ratios: Relative volume percentages for Solution A, Solution B, and extract. The DNA volume can vary depending on the specific plasmid's concentration and may need to be optimized for the user's specific plasmid being used.
| Solution A | Solution B | ||
| Components | Concentration | Components | Concentration |
| ATP | 1.2 mM | Magnesium Glutamate | 10 mM |
| GTP | 0.850 mM | Ammonium Glutamate | 10 mM |
| UTP | 0.850 mM | Potassium Glutamate | 130 mM |
| CTP | 0.850 mM | Phosphoenolpyruvate (PEP) | 30 mM |
| Folinic Acid | 31.50 μg/mL | L-Valine | 2 mM |
| tRNA | 170.60 μg/mL | L-Tryptophan | 2 mM |
| Nicotinamide Adenine Dinucleotide (NAD) | 0.40 mM | L-Isoleucine | 2 mM |
| Coenzyme A | 0.27 mM | L-Leucine | 2 mM |
| Oxalic Acid | 4.00 mM | L-Cysteine | 2 mM |
| Putrescine | 1.00 mM | L-Methionine | 2 mM |
| Spermidine | 1.50 mM | L-Alanine | 2 mM |
| HEPES Buffer pH 7.5 | 57.33 mM | L-Arginine | 2 mM |
| L-Asparagine | 2 mM | ||
| L-Aspartic Acid | 2 mM | ||
| L-Glutamic acid | 2 mM | ||
| L-Glycine | 2 mM | ||
| L-Glutamine | 2 mM | ||
| L-Histidine | 2 mM | ||
| L-Lysine | 2 mM | ||
| L-Proline | 2 mM | ||
| L-Serine | 2 mM | ||
| L-Threonine | 2 mM | ||
| L-Phenylalanine | 2 mM | ||
| L-Tyrosine | 2 mM | ||
Table 4: Solution A and B components. Stock concentrations for the components for Solution A and B were added with their respective amounts, each in a 1.5 mL microfuge tube.
| Extract | Total Protein Concentration (μg/mL) | Standard Deviation |
| 2xYTPG 2.5 OD | 30617 | 3745 |
| CFAI 2.5 OD | 30895 | 2254 |
| CFAI 10.0 OD | 27905 | 3582 |
Table 5: Total extract protein yields. Analysis of the total protein of different cell extract growths. Total protein concentration was determined using a Bradford Assay. Each concentration was determined from triplicates using a 1:40 dilution.
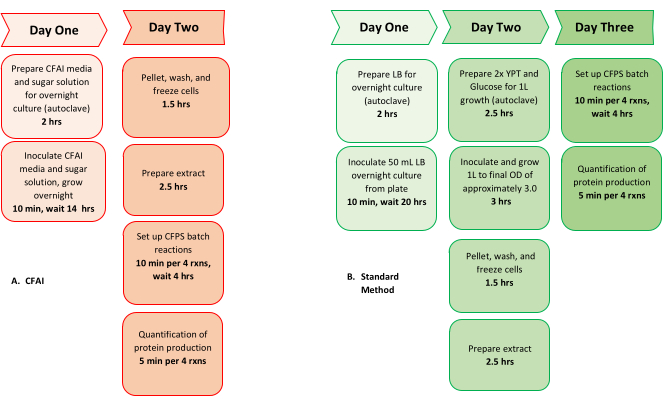
Figure 1: Comparison of CFAI and typical workflow from cells to CFPS: Comparison of the overall timeline from cells to CFPS using the (A) CFAI workflow (left, red) versus the (B) previously established method (green, right). The comparison demonstrates the reduced researcher oversight and timeline when performing CFPS using the CFAI workflow. Please click here to view a larger version of this figure.
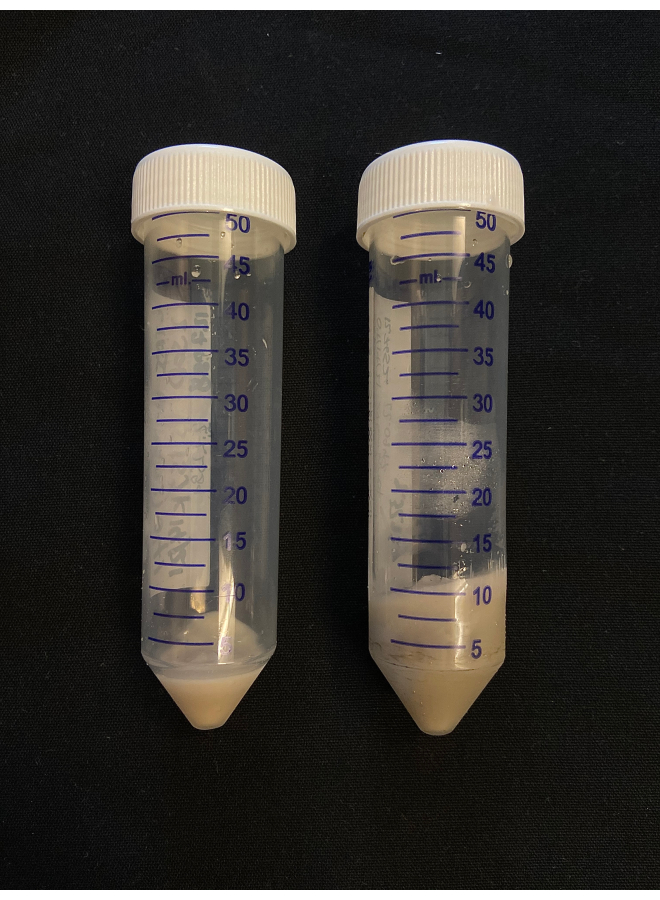

Figure 2: CFAI pellet size comparison. Comparison of CFAI media pellets after cell harvest at different OD600. The media grown to an OD600 of 2.5 produced a 2.23 g cell pellet (left) and the media grown to an OD600 of 10 produced a 9.49 g cell pellet (right). Please click here to view a larger version of this figure.
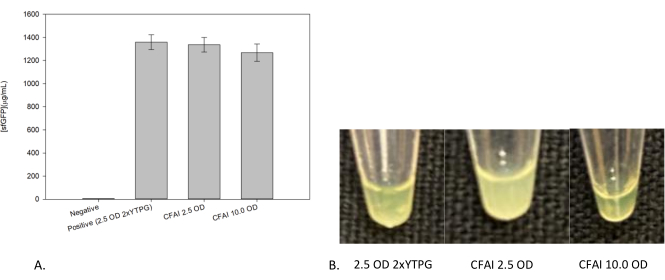
Figure 3: Effects of growth on CFPS reaction yields. (A) Comparison of CFPS reaction yields between growths to 2.5 OD600 and 10 OD600, with (B) images of each CFPS reaction above their respective yield. Cell-free reactions were performed in a 1.5 mL microfuge tube and quantified after 24 h of incubation at 37 °C using a standard curve to correlate fluorescence to sfGFP concentration. The "Negative" corresponds to the set of negative control reactions in which no template DNA was added. The traditional 2xYTPG media (the positive control) and the CFAI extracts are of similar quality as demonstrated through their high CFPS yields. Please click here to view a larger version of this figure.
Discussion
Researcher oversight is traditionally needed for two key actions during cell growth: the induction of T7 RNAP and harvesting cells at a specific OD600. CFAI obviates both of those requirements to decrease the researcher's time and technical training required in order to prepare high quality cell extracts. Auto-induction of T7 RNAP is achieved by replacing glucose with lactose as the primary sugar in the media, obviating the previous need to actively monitor the growth and then induce with IPTG at a precise point during cell growth. The need to actively monitor cell cultures to harvest at a specific OD600 is also obviated, untethering the researcher from the cell culture. This adds to the recent work which have also demonstrated production of quality extracts harvested at non-traditional times13,25,26. The new media formulation improves buffering capacity and carbon sources to support active energy metabolism even as the cell culture approaches stationary phase. The capacity to obtain robust cell extracts from high OD600 cultures allows the researcher to harvest the cultures at their convenience27. The workflow we prefer and recommend is to inoculate the culture in the evening and returning to harvest the next morning.
Harvesting cells at a higher OD600 also results in a significantly larger quantity of cells obtained for extract preparation. For experienced researchers, it is worth noting that the cell pellet is much darker in color compared to cells grown in 2xYTPG media, even when harvested at an OD600 of 2.5. It is also important to note that if the entire cell pellet is being processed at once, the large amount of resuspension obtained per cell pellet when performing lysis via sonication will take some time. Hence, it is important to keep all aliquots cold during this process11,13,14. The increase in extract volume per growth decreases cost proportionally and supports biomanufacturing applications. With the improvements demonstrated, the CFAI workflow provides an easier protocol for new and experienced users of cell-free technology to produce reproducible, functional E. coli extract.
Despite the advantages of the provided CFAI media, there are limitations to this method. The primary challenge is the nascent nature of the workflow. While metabolomics analysis has shed light on the differences in CFAI OD600 10 extracts as well as reaction products compared to 2xYTPG, the implications of these differences on specific applications remain uncharacterized27. Additionally, this workflow has been developed for BL21 E. coli-based lysate. It is unclear whether the media reformulation would support robust extract preparation from other E. coli strains, such as the genomically recoded strains of E. coli28,29. It is possible that the CFAI approach could be utilized for generating extracts from other bacterial organisms, but it is unlikely to support extract preparation for eukaryotic organisms such as Chinese Hamster Ovary or rabbit reticulocyte; however, these have their own established methods30,31. We anticipate that the simplicity of the CFAI workflow will reduce the barriers and incentivize the cell-free community to characterize and evaluate its utility for the broad range of applications that CFPS supports.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
Authors would like to acknowledge Dr. Jennifer VanderKelen and Andrea Laubscher for technical support. Authors would also like to thank Nicole Gregorio, Max Levine, Alissa Mullin, Byungcheol So, August Brookwell, Elizabeth (Lizzy) Vojvoda, Logan Burrington and Jillian Kasman for helpful discussions. Authors also acknowledge funding support from the Bill and Linda Frost Fund, Center for Applications in Biotechnology's Chevron Biotechnology Applied Research Endowment Grant, Cal Poly Research, Scholarly, and the National Science Foundation (NSF-1708919).
Materials
| 1.5 mL Microfuge Tubes | Phenix | MPC-425Q | |
| 1L Centrifuge Tube | Beckman Coulter | A99028 | |
| Avanti J-E Centrifuge | Beckman Coulter | 369001 | |
| CoA | Sigma-Aldrich | C3144-25MG | |
| Cytation 5 Cell Imaging Multi-Mode Reader | Biotek | BTCYT5F | |
| D-Glucose | Fisher | D16-3 | |
| D-Lactose | Alfa Aesar | J66376 | |
| DTT | ThermoFisher | 15508013 | |
| Folinic Acid | Sigma-Aldrich | F7878-100MG | |
| Glycerol | Fisher | BP229-1 | |
| Glycine | Sigma-Aldrich | G7126-100G | |
| HEPES | ThermoFisher | 11344041 | |
| IPTG | Sigma-Aldrich | I6758-1G | |
| JLA-8.1000 Rotor | Beckman Coulter | 366754 | |
| K(Glu) | Sigma-Aldrich | G1501-500G | |
| K(OAc) | Sigma-Aldrich | P1190-1KG | |
| KOH | Sigma-Aldrich | P5958-500G | |
| L-Alanine | Sigma-Aldrich | A7627-100G | |
| L-Arginine | Sigma-Aldrich | A8094-25G | |
| L-Asparagine | Sigma-Aldrich | A0884-25G | |
| L-Aspartic Acid | Sigma-Aldrich | A7219-100G | |
| L-Cysteine | Sigma-Aldrich | C7352-25G | |
| L-Glutamic Acid | Sigma-Aldrich | G1501-500G | |
| L-Glutamine | Sigma-Aldrich | G3126-250G | |
| L-Histadine | Sigma-Aldrich | H8000-25G | |
| L-Isoleucine | Sigma-Aldrich | I2752-25G | |
| L-Leucine | Sigma-Aldrich | L8000-25G | |
| L-Lysine | Sigma-Aldrich | L5501-25G | |
| L-Methionine | Sigma-Aldrich | M9625-25G | |
| L-Phenylalanine | Sigma-Aldrich | P2126-100G | |
| L-Proline | Sigma-Aldrich | P0380-100G | |
| L-Serine | Sigma-Aldrich | S4500-100G | |
| L-Threonine | Sigma-Aldrich | T8625-25G | |
| L-Tryptophan | Sigma-Aldrich | T0254-25G | |
| L-Tyrosine | Sigma-Aldrich | T3754-100G | |
| Luria Broth | ThermoFisher | 12795027 | |
| L-Valine | Sigma-Aldrich | V0500-25G | |
| Mg(Glu)2 | Sigma-Aldrich | 49605-250G | |
| Mg(OAc)2 | Sigma-Aldrich | M5661-250G | |
| Microfuge 20 | Beckman Coulter | B30134 | |
| Molecular Grade Water | Sigma-Aldrich | 7732-18-5 | |
| NaCl | Alfa Aesar | A12313 | |
| NAD | Sigma-Aldrich | N8535-15VL | |
| New Brunswick Innova 42/42R Incubator | Eppendorf | M1335-0000 | |
| NH4(Glu) | Sigma-Aldrich | 09689-250G | |
| NTPs | ThermoFisher | R0481 | |
| Oxalic Acid | Sigma-Aldrich | P0963-100G | |
| PEP | Sigma-Aldrich | 860077-250MG | |
| Potassium Phosphate Dibasic | Acros, Organics | A0382124 | |
| Potassium Phosphate Monobasic | Acros, Organics | A0379904 | |
| PureLink HiPure Plasmid Prep Kit | ThermoFisher | K210007 | |
| Putrescine | Sigma-Aldrich | D13208-25G | |
| Spermidine | Sigma-Aldrich | S0266-5G | |
| Tris(OAc) | Sigma-Aldrich | T6066-500G | |
| tRNA | Sigma-Aldrich | 10109541001 | |
| Tryptone | Fisher Bioreagents | 73049-73-7 | |
| Tunair 2.5L Baffled Shake Flask | Sigma-Aldrich | Z710822 | |
| Ultrasonic Processor | QSonica | Q125-230V/50HZ | |
| Yeast Extract | Fisher Bioreagents | 1/2/8013 |
Riferimenti
- Gregorio, N. E., Levine, M. Z., Oza, J. P. A user’s guide to cell-free protein synthesis. Methods and Protocols. 2 (1), 1-34 (2019).
- Silverman, A. D., Karim, A. S., Jewett, M. C. Cell-free gene expression: an expanded repertoire of applications. Nature Reviews Genetics. 21 (3), (2020).
- Swartz, J. R. Expanding biological applications using cell-free metabolic engineering: An overview. Metabolic Engineering. 50, 156-172 (2018).
- Jared, B., Dopp, L., Tamiev, D. D., Reuel, N. F. Cell-free supplement mixtures: Elucidating the history and biochemical utility of additives used to support in vitro protein synthesis in E. coli extract. Biotechnology Advances. 37 (1), 246-258 (2019).
- Jewett, M. C., Swartz, J. R. Mimicking the Escherichia coli cytoplasmic environment activates long-lived and efficient cell-free protein synthesis. Biotechnology and Bioengineering. 86 (1), 19-26 (2004).
- Calhoun, K. A., Swartz, J. R. Energizing cell-free protein synthesis with glucose metabolism. Biotechnology and Bioengineering. 90 (5), 606-613 (2005).
- Levine, M. Z., Gregorio, N. E., Jewett, M. C., Watts, K. R., Oza, J. P. Escherichia coli-based cell-free protein synthesis: Protocols for a robust, flexible, and accessible platform technology. Journal of Visualized Experiments. (144), e58882 (2019).
- Sun, Z. Z., et al. Protocols for Implementing an Escherichia coli Based TX-TL Cell-Free Expression. System for Synthetic Biology. , 1-14 (2013).
- Pardee, K., et al. Portable, on-demand biomolecular manufacturing. Cell. 167, 248-254 (2016).
- Moore, S. J., Macdonald, J. T., Freemont, P. S. Cell-free synthetic biology for in vitro prototype engineering. Biochemical Society Transactions. 45 (3), 785-791 (2017).
- Cole, S. D., Miklos, A. E., Chiao, A. C., Sun, Z. Z., Lux, M. W. Methodologies for preparation of prokaryotic extracts for cell-free expression systems. Synthetic and Systems Biotechnology. 5 (4), 252-267 (2020).
- Didovyk, A., Tonooka, T., Tsimring, L., Hasty, J. Rapid and scalable preparation of bacterial lysates for cell-free gene expression. ACS Synthetic Biology. 6 (12), 2198-2208 (2017).
- Dopp, J. L., Reuel, N. F. Process optimization for scalable E. coli extract preparation for cell-free protein synthesis. Biochemical Engineering Journal. 138, 21-28 (2018).
- Kwon, Y. C., Jewett, M. C. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. Scientific Reports. 5, 8663 (2015).
- Endo, Y., Sawasaki, T. Cell-free expression systems for eukaryotic protein production. Current Opinion in Biotechnology. 17 (4), 373-380 (2006).
- Zemella, A., Thoring, L., Hoffmeister, C., Kubick, S. Cell-free protein synthesis: Pros and cons of prokaryotic and eukaryotic systems. ChemBioChem. 16 (17), 2420-2431 (2015).
- Laohakunakorn, N., Grasemann, L., Lavickova, B., Michielin, G. Bottom-up construction of complex biomolecular systems with cell-free synthetic biology. Frontiers in Bioengineering and Biotechnology. 8, (2020).
- Ahn, J. H., et al. Cell-free synthesis of recombinant proteins from PCR-amplified genes at a comparable productivity to that of plasmid-based reactions. Biochemical and Biophysical Research Communications. 338 (3), 1346-1352 (2005).
- Kim, T. W., et al. Simple procedures for the construction of a robust and cost-effective cell-free protein synthesis system. Journal of Biotechnology. 126 (4), 554-561 (2006).
- Hunt, J. P., et al. Streamlining the preparation of “endotoxin-free” ClearColi cell extract with autoinduction media for cell-free protein synthesis of the therapeutic protein crisantaspase. Synthetic and Systems Biotechnology. 4 (4), 220-224 (2019).
- Studier, F. W. Protein production by auto-induction in high-density shaking cultures. Protein Expression and Purification. 41, 207-234 (2005).
- Shrestha, P., Holland, T. M., Bundy, B. C. Streamlined extract preparation for Escherichia coli-based cell-free protein synthesis by sonication or bead vortex mixing. BioTechniques. 53 (3), 163-174 (2012).
- Bosdriesz, E., Molenaar, D., Teusink, B., Bruggeman, F. J. How fast-growing bacteria robustly tune their ribosome concentration to approximate growth-rate maximization. FEBS Journal. 282 (10), 2029-2044 (2015).
- Zawada, J., Swartz, J. Maintaining rapid growth in moderate-density Escherichia coli fermentations. Biotechnology and Bioengineering. 89 (4), 407-415 (2005).
- Kim, J., Copeland, C. E., Padumane, S. R., Kwon, Y. C. A crude extract preparation and optimization from a genomically engineered Escherichia coli for the cell-free protein synthesis system: Practical laboratory guideline. Methods and Protocols. 2 (3), 1-15 (2019).
- Failmezger, J., Rauter, M., Nitschel, R., Kraml, M., Siemann-Herzberg, M. Cell-free protein synthesis from non-growing, stressed Escherichia coli. Scientific Reports. 7 (1), 1-10 (2017).
- Levine, M. Z., et al. Activation of energy metabolism through growth media reformulation enables a 24-h workflow for cell-free expression. ACS Synthetic Biology. 9 (10), 2765-2774 (2020).
- Martin, R. W., et al. Cell-free protein synthesis from genomically recoded bacteria enables multisite incorporation of noncanonical amino acids. Nature Communications. , 1-9 (2018).
- Soye, B. J. D., et al. Resource A highly productive, One-pot cell-free protein synthesis platform based on genomically recoded Escherichia coli resource. Cell Chemical Biology. 26 (12), 1743-1754 (2019).
- Ezure, T., Suzuki, T., Ando, E. A cell-free protein synthesis system from insect cells. Methods in Molecular Biology. , 285-296 (2014).
- Heide, C., et al. development and optimization of a functional mammalian cell-free protein synthesis platform. Frontiers in Bioengineering and Biotechnology. 8, 1-10 (2021).

