Hyaluronic-Acid Based Hydrogels for 3-Dimensional Culture of Patient-Derived Glioblastoma Cells
Summary
Here, we present a protocol for three-dimensional culture of patient-derived glioblastoma cells within orthogonally tunable biomaterials designed to mimic the brain matrix. This approach provides an in vitro, experimental platform that maintains many characteristics of in vivo glioblastoma cells typically lost in culture.
Abstract
Glioblastoma (GBM) is the most common, yet most lethal, central nervous system cancer. In recent years, many studies have focused on how the extracellular matrix (ECM) of the unique brain environment, such as hyaluronic acid (HA), facilitates GBM progression and invasion. However, most in vitro culture models include GBM cells outside of the context of an ECM. Murine xenografts of GBM cells are used commonly as well. However, in vivo models make it difficult to isolate the contributions of individual features of the complex tumor microenvironment to tumor behavior. Here, we describe an HA hydrogel-based, three-dimensional (3D) culture platform that allows researchers to independently alter HA concentration and stiffness. High molecular weight HA and polyethylene glycol (PEG) comprise hydrogels, which are crosslinked via Michael-type addition in the presence of live cells. 3D hydrogel cultures of patient-derived GBM cells exhibit viability and proliferation rates as good as, or better than, when cultured as standard gliomaspheres. The hydrogel system also enables incorporation of ECM-mimetic peptides to isolate effects of specific cell-ECM interactions. Hydrogels are optically transparent so that live cells can be imaged in 3D culture. Finally, HA hydrogel cultures are compatible with standard techniques for molecular and cellular analyses, including PCR, Western blotting and cryosectioning followed by immunofluorescence staining.
Introduction
Three-dimensional (3D) culture systems recapitulate interactions between cells and their surrounding extracellular matrix (ECM) in native tissues better than their two-dimensional (2D) counterparts1,2. Advancements in tissue engineering have yielded sophisticated, 3D culture platforms that enable controlled investigations into 1) how chemical and physical components of the matrix microenvironment affect cell behaviors and 2) efficacy of new therapeutic strategies for a number of diseases, including cancers2. While in vitro models cannot account for systemic factors, such as endocrine and immune signals, and thus cannot completely replace in vivo models, they provide several advantages including reproducibility, experimental control, affordability and speed. Here, we describe the use of brain-mimetic hydrogels in which 3D cultures of patient-derived brain tumor cells capture many aspects of tumor physiology, in particular, the dynamics of acquiring treatment resistance3. Compared to other in vitro methods, these cultures better represent in vivo tumor models and clinical observations3.
Glioblastoma (GBM) is the most frequent and lethal cancer originating in the brain, with a median survival of only 1-2 years4,5. In recent years, many studies have focused on the influence of tumor matrix environment in GBM6,7,8. The unique brain ECM has been reported to affect GBM cell migration, proliferation, and therapeutic resistance6,7,8,9,10,11,12. Hyaluronic acid (HA) is an abundant glycosaminoglycan (GAG) in the brain, where it interacts with other GAGs and proteoglycans to form a hydrogel-like mesh13. Many studies have reported HA overexpression in GBM tumors and its subsequent effects on cancer progression8,9,13,14,15,16,17. Other ECM components also affect GBM tumor growth and invasion6,7,15,18. For example, fibronectin and vitronectin, which are typically overexpressed in GBM, induce heterodimerization of cell surface integrin receptors through binding to the "RGD" sequence and initiate complex signaling cascades that promote tumor survival19,20,21. Besides biochemical influences, physical properties of the tissue matrix also affect GBM progression22,23.
Continual acquisition of resistance to therapies is one of the main drivers of GBM lethality4. Drugs showing promising results in 2D or gliomasphere models have failed in subsequent animal studies and clinical cases3, indicating that the effects of microenvironmental factors significantly contributed to GBM tumor response1. While animal models can provide a 3D, physiologically appropriate microenvironment to xenografted patient cells and generate clinically relevant outcomes24,25, the complexity of the brain microenvironment in vivo makes it challenging to determine which features, including cell-matrix interactions, are key to specific biological outcomes. Identification of new therapeutic targets will benefit from the use of simplified culture platforms in which biochemical and biophysical properties are defined.
Unlike previously reported biomaterial models of the GBM tumor microenvironment26,27 which have not achieved true orthogonal control over individual biochemical and physical features of the ECM, the biomaterial platform reported here enables decoupling of the contributions of multiple independent features to GBM cell phenotype. Here, we present an HA-based, orthogonally tunable, hydrogel system for 3D culture of patient-derived GBM cells. Hydrogels are formed from two polymer components: 1) biologically active HA and 2) biologically inert polyethylene glycol (PEG). PEG is a widely used biocompatible and hydrophilic material with low protein adsorption and minimal immunogenicity28. Here, approximately 5% of glucuronic acid moieties on HA chains are functionalized with thiol groups to enable crosslinking to a commercially available 4-arm-PEG terminated with maleimides via Michael-type addition. In its most common form in the body, HA exists in high molecular weight (HMW) chains. Here, a low degree of modification of HMW HA (500-750 kDa) helps to preserve native interactions of HA and its cell receptors, including CD4429. By substituting PEG-thiol for HA-thiol while maintaining a constant molar ratio of total thiols to maleimides, HA concentration can be decoupled from mechanical properties of the resulting hydrogels. Furthermore, stoichiometric controls can be used to conjugate cysteine-terminated peptides to a defined average number of maleimide-terminated arms on each 4-arm-PEG. Incorporation of ECM-derived, adhesive peptides enables interactions with integrins on cultured cells, through which biochemical and chemical signals are transduced1. Maleimide-thiol addition occurs very quickly under physiological conditions, minimizing the time required for cell encapsulation and maximizing survival of patient-derived cells. Moreover, hydrogel cultures can be treated like typical tissue specimen and are compatible with standard characterization techniques including Western blotting, flow cytometry, and immunofluorescence staining. The following protocol describes the procedures for fabricating hydrogels, establishing 3D cultures of patient-derived GBM cells and techniques for biochemical analysis.
Protocol
All human tissue collection steps were carried out under institutionally approved protocols.
1. Thiolation of Hyaluronic Acid
Note: Molar ratios are stated with respect to total number of carboxylate groups unless otherwise specified.
- Dissolve 500 mg of sodium hyaluronate (HA, 500-750 kDa) at 10 mg/mL in deionized, distilled water (DiH2O) in an autoclave sterilized, 250 mL Erlenmeyer flask. Stir the solution (~200 rpm) at room temperature for 2 hours to fully dissolve HA. Use a stir bar and magnetic stir plate to keep reaction stirring during thiolation procedure.
- Using 0.1 M hydrochloric acid (HCl), adjust the pH of the HA solution to 5.5. Weigh out 69.6 mg (0.25x molar ratio) of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC).
NOTE: Although undissolved EDC can be added directly to the HA solution, it is typically easier to dissolve EDC first in 1 mL of DiH2O and then quickly add the EDC solution to the HA solution. Pre-dissolution in 1 mL of DiH2O can also be done with N-hydroxysuccinimide (NHS) and cystamine dihydrocholoride before adding to the reaction in steps 1.3 and 1.4. - Add 17.9 mg (0.125x molar ratio) of NHS to the reaction. Adjust the pH of the reaction solution to 5.5 using 0.1 M HCl and incubate the reaction at room temperature for 45 minutes.
- Add 70.0 mg (0.25x molar ratio) of cystamine dihydrochloride into the reaction solution. Use 0.1 M sodium hydroxide (NaOH) to adjust pH to 6.25. Incubate the reaction at room temperature overnight while stirring.
- On the following day, use 1 M NaOH to adjust the pH of the reaction solution to around 8. Add 192 mg (4x molar ratio) of dithiothreitol (DTT) to the reaction. Adjust the pH back to 8 after addition of DTT and leave stirring for 1-2 hours at room temperature.
- Use 1 M HCl to adjust pH to 4. Transfer the entire reaction mixture into pre-soaked dialysis membrane (molecular weight cut-off around 13 kDa) and dialyze against DiH2O pre-adjusted to pH 4.
NOTE: The volume of DiH2O for dialysis should be at least 40 times the volume of the reacted HA solution (e.g., 50 mL sample in 2 L of DiH2O, pH 4). - Replace the dialysis solution (DiH2O, pH 4) twice daily for 3 days at room temperature.
- Filter the dialyzed solution through a 0.22 µm membrane, vacuum-driven filter. Flash freeze solution of thiolated HA in liquid nitrogen. Use a lyophilizer to freeze dry samples over 2 days. Thiolated HA in dry form can be stored in a vacuum-sealed desiccator at -20 °C for at least 6 months.
- To determine the degree of thiolation, use Ellman's test and/or proton NMR spectroscopy30,31.
2. Preparation of Crosslinking Materials
- Dissolve thiolated HA at the desired concentration (in this example, 13.3 mg/mL) in HEPES-buffered saline (20 mM HEPES in Hank's balanced salt saline (HBSS) buffer, adjusted to fall within a pH of 8-9) in an amber vial to minimize exposure to ambient light. Keep the solution stirring constantly.
NOTE: Formation of disulfide bonds between thiols conjugated to HA may occur if the pH is above 8, solution concentration is too high, or the solution is left stirring for too long before gelation. To avoid this issue, use below 15 mg/mL of thiolated HA and dissolve thiolated HA within 2 hours of forming hydrogels. - Dissolve 4-arm-PEG-maleimide (PEG-Mal, 20 kDa) and 4-arm-PEG-thiol (PEG-thiol, 20 kDa) in phosphate buffered saline (PBS), pH 7.4, to prepare 50 mg/mL PEG-Mal and PEG-thiol stock solutions.
NOTE: It takes at least 1 hour to fully dissolve PEG reagents. - If adding ECM-derived peptides, dissolve cysteine-containing peptides in PBS to prepare a stock solution.
NOTE: Use a stock concentration of 2-4 mM. - Soak silicone rubber molds in ethanol for at least 20 min to clean them and then autoclave them for sterilization.
3. Hydrogel Crosslinking and Cell Encapsulation
Note: As an example here, the encapsulation of four individual, 80 µL hydrogels with 0.5% (w/v) HA and compressive modulus of 1 kPa is described3. Please see Table 1 for example recipes that yield hydrogels with varying properties: two hydrogels incorporating the integrin-binding peptide RGD and two hydrogels incorporating cysteine caps as a negative control for peptide activity. Seeding concentration of patient-derived GBM cells is 500,000 cells/mL.
- Dilute PEG-Mal stock solution (50 mg/mL, from step 2.2) to 12.5 mg/mL by adding 40 µL of the stock solution to 120 µL PBS. Split the diluted solution into two 1.5 mL microcentrifuge tubes so that each tube has 80 µL.
- Add 16 µL of stock RGD solution (2.8 mM, from step 2.3) to one tube and 16 µL of stock cysteine solution (2.8 mM, from step 2.3) to the second tube. Vortex to mix. Place the PEG-Mal-RGD or PEG-Mal-CYS solutions on ice until use in step 3.9.
NOTE: The procedure as described here yields hydrogels with ~140 µM of peptide. This is equivalent to approximately 1 out of every 8 available PEG arms being occupied with a peptide. This can be varied by altering the molar ratio of cysteine-terminated peptides to available maleimide groups. In general, a maximum concentration of 280 µM of peptide can be achieved while still leaving sufficient numbers of maleimide groups available for hydrogel crosslinking. - Dilute the PEG-thiol stock solution (50 mg/mL, from step 2.2) to 5 mg/mL by mixing 4 µL of 50 mg/mL stock solution with 36 µL of PBS. Add 120 µL of dissolved, thiolated HA from step 2.1 to the mixture.
NOTE: Solutions of HMW HA are very viscous. Thus, we recommend using wide-orifice micropipette tip to transfer solutions. Pipette slowly to avoid solution sticking to the walls of micropipette tip and to improve measurement accuracy. - Place the clean, dry molds (prepared in step 2.4) into each well of a non-tissue culture treated 12-well plate. Using the clean, blunt end of a pipette tip, press the gel mold and double check sealing between molds and bottom of well plate.
NOTE: Check the seal again right before encapsulation to prevent leakage. - Passage cultured GBM cells, dissociate to single cells and determine cell concentration as previously described3.
NOTE: Some GBM lines cannot be dissociated to single cells. In this case, whole gliomaspheres can be encapsulated. However, prepare dissociated single cells after precise cell counting to improve reproducibility. The protocol for gliomasphere passaging varies among different labs and many of these are likely compatible with hydrogel encapsulation.- (Recommended) Centrifuge gliomaspheres at 500 x g for 5 min. Remove the supernatant and add 1 mL of cell dissociation enzyme to the cell pellet. Then, incubate for 5 min, gently tapping the tube to agitate.
- Add 4 mL of complete culture medium (50 ng/mL EGF, 20 ng/mL FGF-2, 25 µg/mL heparin, G21 supplement and 1% penicillin/streptomycin in DMEM/F12) to the cells.
- Centrifuge again at 500 x g for 5 min and remove the supernatant. Finally, resuspend the cell pellet in 1 mL of the complete medium and pass suspended cells through a 70 µm cell strainer. (Recommended) wash the strainer with an additional 4 mL of the complete medium to maximize the number of cells recovered.
- Estimate the concentration of cells in the suspension using a hemocytometer. Split the cell suspension into 2 centrifuge tubes, where each tube contains ~80,000 cells. Centrifuge at 500 x g for 5 min; generally, 80,000 cells will make up 2 80 µL hydrogels.
- Remove the supernatant and resuspend one pellet in 80 µL of PEG-MAL-RGD solution and a second pellet in 80 µL of PEG-MAL-CYS solution (prepared in step 3.1).
- Using a 200 µL, wide-orifice micropipette tip, dispense 40 µL of HA solution (from step 3.3 above) into each rubber silicone mold (as prepared in step 3.4).
- Using a 200 µL, wide-orifice micropipette tip, mix the 40 µL of PEG-MAL-CYS or PEG-MAL-RGD cell solution with the HA solution in the mold. Pipette up and down quickly no more than 10 times. Repeat for each gel culture being prepared.
NOTE: This step takes practice, since initial gelation occurs quickly (within 30 s). Pipette up and down while moving the tip to different locations in molds to ensure even mixing. Always keep the tip below liquid level to avoid formation of air bubbles. Do not mix the solution too many times as gel may get formed inside the micropipette tip. PEG-MAL-CYS and PEG-MAL-RGD can be combined at varying ratios to achieve the desired RGD peptide concentration. - Place the well-plate containing gel-encapsulated cells into a 37 °C cell culture incubator for 5-10 minutes to ensure completion of the reaction.
- Add 2-2.5 mL of culture medium to each well with formed gels. Use a sterile, 2 µL pipette tip or micro-spatula, to gently separate the mold and gel. Use pre-sterilized forceps to remove the mold from the well plate. Place the well plate back to cell incubator (37 °C and 5%CO2) for culture and future experiments.
NOTE: Pull the mold vertically upward to avoid harming the gel cultures. In general, medium in gel cultures should be replaced every 3-4 days. Take care not to aspirate hydrogels when removing medium. If this is an issue, we recommend using a plastic transfer pipette. If bioluminescence imaging to track cell number is planned, GBM cells must be transduced with lentivirus encoding constitutively luciferase expression before encapsulation, as previously described3. For bioluminescence imaging, add 1 mM of D-luciferin to culture medium 1 h prior to luminescence imaging.
4. Lysate Preparation for Western Blotting
- Chill 1.5 mL microcentrifuge tubes on ice (1 per gel samples). Cool centrifuge to 4 °C.
- Remove medium from well plates. Transfer gels into prechilled microcentrifuge tubes.
- Add 100 µL of RIPA buffer with 1x protease/phosphatase inhibitors.
- Using a 1 mL syringe with a 20 G needle, break up the gel by pushing the whole mixture through needle at least 20 times.
- Flash spin the sample using bench top microcentrifuge and place the sample back on ice.
- Vortex samples briefly every 5 min for a total of 20 min.
- Centrifuge samples at 14000 x g for 15 min at 4 °C.
- Transfer supernatant to new, pre-chilled microcentrifuge tubes and store at -20 °C (short-term) or and -80 °C (long-term). Perform gel electrophoresis using standard procedures, as previously described3.
5. Extracting Single Cells from Hydrogel Cultures for Flow Cytometry
- Remove medium and transfer gel culture (from step 3.11) into a 1.5 mL microcentrifuge tube.
- Incubate gel with 500 µL of cell dissociation enzyme (e.g., TrypLE Express) at 37 °C for 5 min, occasionally flicking the tube to agitate.
- Transfer the mixture into a 50 mL centrifuge tube with 5 mL of complete medium. Place the tube on ice.
- Attach a 20G needle onto a 10 mL syringe. Gently pull the cell suspension up and down through the needle 8 times.
- Flow the mixture through 70 µm cell strainer into a new 50 mL centrifuge tube. Apply an additional 5 mL of complete medium through the strainer to collect any remaining cells.
- Centrifuge the sample at 400 x g for 5 min. Remove the supernatant and resuspend the pellet in desired buffer for flow cytometry (using standard protocols3).
6. Cryopreservation of Hydrogels for Sectioning
- Remove medium from gel cultures (from step 3.11).
- Incubate gel cultures with 2 mL of 4% paraformaldehyde (PFA) at 4 °C overnight.
CAUTION: PFA is toxic and must be handled carefully. - On the next day, remove PFA. Add 2 mL of 5% sucrose in PBS to gels and incubate for 1 h at room temperature.
- Replace 5% sucrose solution with 2 mL of 20% sucrose in PBS and incubate for 30 min.
- Replace sucrose solution with 2 mL of fresh 20% sucrose in PBS and incubate for an additional 30 min.
- For a third time, replace the sucrose solution with 2 mL of fresh 20% sucrose in PBS. Incubate at 4 °C overnight.
- Prepare 20% sucrose in Optimal Cutting Temperature (OCT) compound.
NOTE: Due to the viscosity of OCT, dissolution of sucrose can take some time (up to overnight). We recommend putting the mixture onto a shaker during dissolution to maintain agitation. - The next day, remove the 20% sucrose solution and gently pour the 20% sucrose in OCT solution over the gel. Make sure to cover the entire gel. Incubate at 4 °C for 3 h.
- Transfer the gel into the center of an embedding mold using a large, flat spatula; this must be done carefully to avoid damaging the gel.
- Cool 2-methylbutane inside a cryochamber with an excess of dry ice.
- Fill the embedding mold with pure OCT (no sucrose). Fill to just below top edge of mold.
- Freeze the hydrogel sample by immersion in cooled 2-methylbutane and then proceed to sectioning.
- Section the sample using a cryostat; section thickness of 10-18 µm and sectioning temperature of around -26 °C are recommended.
- Perform standard immunostaining procedures on sectioned gel culture, as previously described3.
NOTE: PFA is known irritant and carcinogen. Please handle and dispose PFA according to locally applicable standards and regulations.
7. Total RNA Extraction from Samples in Hydrogel
Note: Here, we describe a protocol using a commercial kit (see the Table of Materials) to extract total RNA from hydrogel cultured cells. The buffers and all material are available within the kit used.
- Remove culture medium. Use a P1000 transfer pipette with wide-bore tip to transfer the hydrogel culture into 1.5 mL microcentrifuge tube.
- Add 350 µL of buffer RLT to the hydrogel culture.
- Using a 20 G needle attached to a 1 mL syringe, shred the gel by pushing the whole mixture through needle at least 20 times.
- Transfer the whole mixture into a homogenizer column placed in a 2 mL collection tube. Centrifuge at 13,000 x g for 2 min.
- Transfer the supernatant into a clean 1.5 mL microcentrifuge tube. Be careful not to disturb the gel precipitate at the bottom.
- Mix the lysate sample with 350 µL of 100% ethanol. Transfer all the sample to a spin column (from the kit) place in a 2 mL collection tube. Centrifuge for 1 min at 13,000 x g.
- For subsequent steps for RNA purification for PCR, follow the general protocol provided by the kit manufacturer.
NOTE: Once RNA is extracted, standard protocols for PCR can be used.
Representative Results
For each batch of thiolated HA, the degree of thiolation should be verified using H1-NMR or an Ellman's test. HA modification using the procedure described here consistently generates ~5% thiolation (defined as the molar ratio of thiols to HA disaccharides) (Figure 1).
Setting up this new culture platform will require each laboratory to perform rigorous testing to ensure good culture viability prior to implementing large-scale experiments. Our 80 µL hydrogels with a seeding density 500,000 cells/mL (40,000 cells/gel) consistently result in proliferation rates that are comparable to, or better than, gliomasphere cultures (Figure 2A-2C). As HA/PEG hydrogels are optically transparent, cell behaviors can be observed directly in live, 3D cultures using phase contrast or fluorescence microscopy. Figure 3A shows that, 4 days after encapsulation, GBM cells in RGD-containing hydrogels exhibit an invasive phenotype, while cells cultured in hydrogel using PEG-MAL-CYS controls have a spherical morphology.
As is typical with xenograft models where researchers must wait for days to months until implanted tumors reach progressive growth24,32, patient-derived cells also take time to adjust to a new culture environment. Thus, we recommend culturing 4-8 days before commencing experiments, such as drug treatment, to ensure most cells have entered the exponential growth phase. Beyond drug treatments, reagents like soluble cyclo-RGD, which competitively disrupt cell interactions with RGD in hydrogels, can be added to culture medium (Figure 3A).
Our hydrogel system is compatible with many common methods for investigating glioma cell biology. Using bioluminescence imaging, which is commonly used to monitor rodent xenograft tumors, relative numbers of viable cells can be observed in hydrogel cultures during the course of treatment. Figure 3B provides an example of this method, where effects of erlotinib treatment on hydrogel-cultured GBM cells were evaluated over 6 days. In general, hydrogel cultures can be treated as tissue samples when preparing lysates for Western blot or PCR. Western blot analysis of cleaved poly ADP polymerase indicates that relative degree of apoptosis in treated CD44 knockdown cells is higher than wildtype GBM cells cultured in 0.5% HA hydrogels (Figure 3C). Similarly, single cell suspensions can be prepared from hydrogel cultures for analysis via flow cytometry using standard protocols for liberating single cells from intact tissues (Figure 2). In addition to cell features, cryosections of hydrogel-based, 3D cultures preserve ECM deposited by cultured cells. For example, deposition patterns of type IV collagen shift upon erlotinib treatment of hydrogel cultures (Figure 3D).
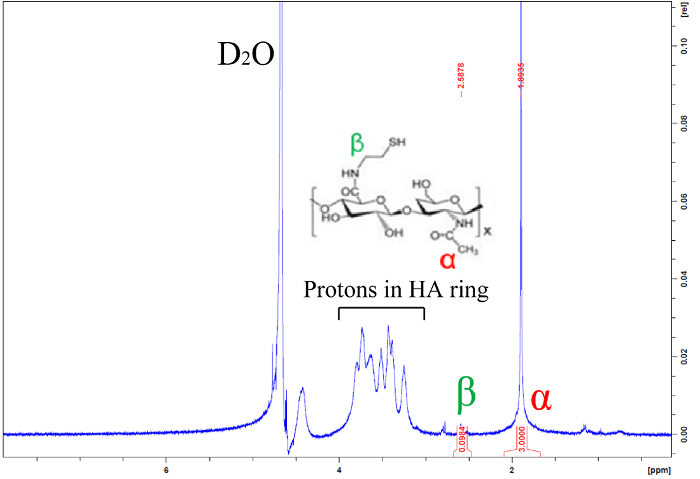
Figure 1: Representative H1-NMR spectrum of thiolated hyaluronic acid. Integrated peaks indicate that approximately 5% of HA glucuronic acid groups have been modified with a thiol. Please click here to view a larger version of this figure.
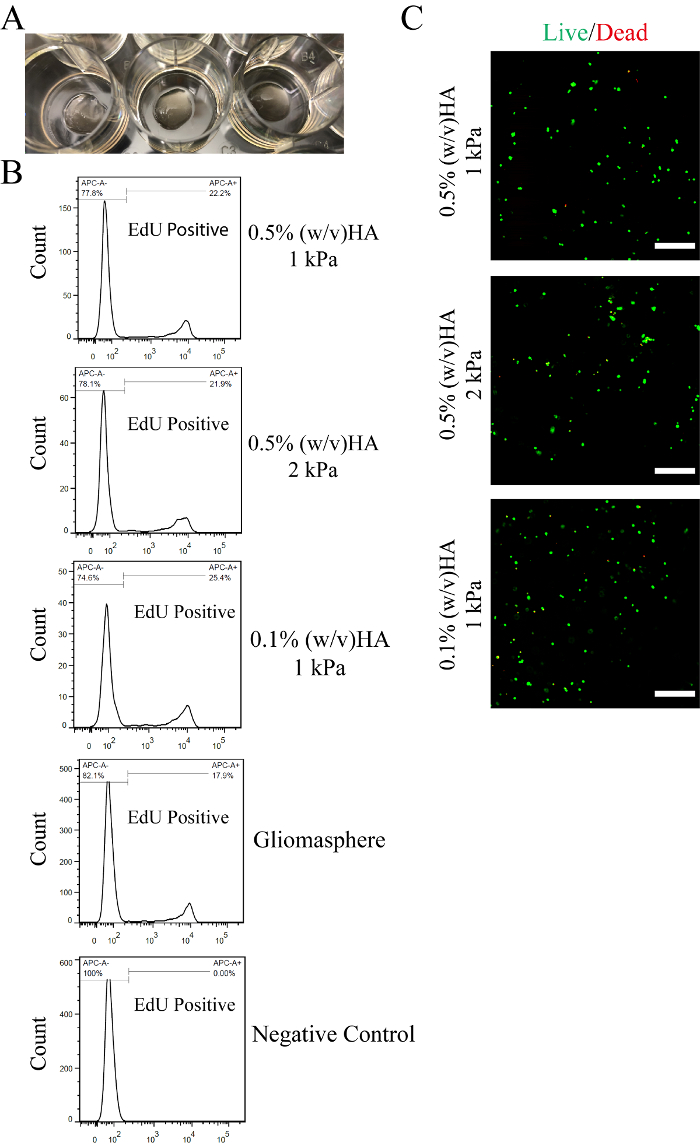
Figure 2: Proliferation of hydrogel-encapsulated cells. A) Example images of fabricated 80 µL gels in 12-well plates. After crosslinking, hydrogels were swollen in cell culture medium. B) Representative results of proliferation rate measured using flow cytometry. GBM cells (HK301) were incubated with 1 µM EdU (5-ethynyl-2′-deoxyuridine) for 2.5 h on the fourth day after encapsulation or passaging. A click-reaction was used to conjugate fluorescent dye to incorporated EdU, as detailed in Xiao et al. 2017.3 Negative controls, where no EdU was added to cultures, were included. C) Representative confocal microscopy images of live (green) and dead (red) cells 24 hours after hydrogel cultures of GBM cells (HK157) were established. Scale bars = 200 µm. Please click here to view a larger version of this figure.
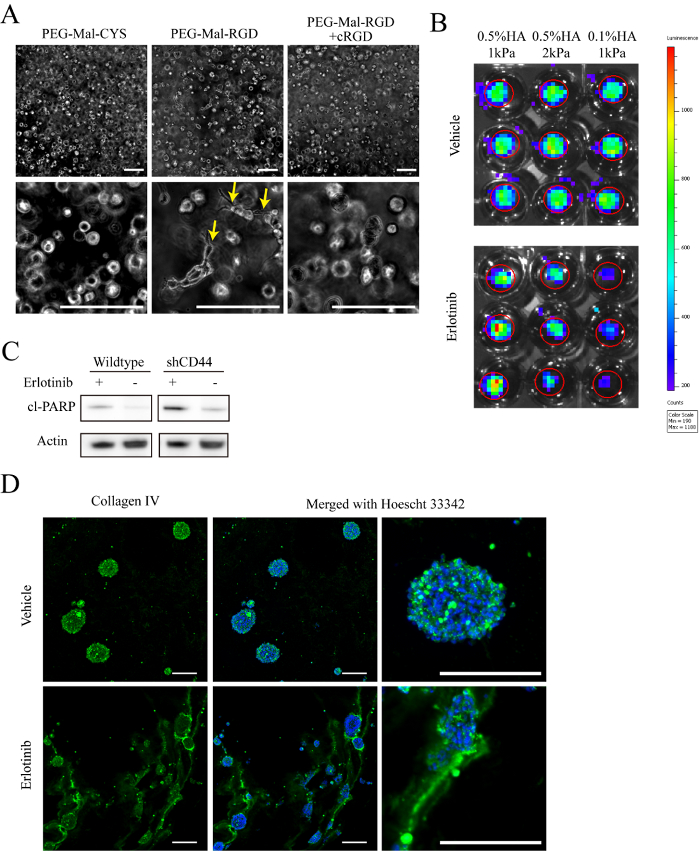
Figure 3: Characterization. A) Representative phase contrast images of 0.5% (w/v) HA hydrogel-cultured cells under varying conditions for 8 days after encapsulation. Arrows indicate cells with an invasive morphology. B) Representative images of bioluminescence signal measured after 15 days of treatment with 1 µM erlotinib or vehicle (DMSO). 1 mM D-luciferin was added to cell culture medium 1 h prior to imaging. Cells were transduced with lentivirus encoding for constitutive expression of firefly luciferase prior to encapsulation. C) Representative immune-blot images analyzing cleaved poly ADP polymerase (cl-PARP) expression in GBM cells (HK301) cultured in hydrogels with 0.5% (w/v) HA and 1 kPa compressive modulus. All cropped images shown were from the same blot. D) Representative staining of collagen IV (green) and Hoechst 33342 (blue) in hydrogel-cultured HK301 cells 12 days after treatment with 1 µm erlotinib or vehicle (DMSO). Scale bar = 200 µm. Please click here to view a larger version of this figure.
| Gel Type | Part A (40 µL each) | Part B (40 µL each) | ||||
| 4Arm-PEG-MAL (50mg/mL) | Cysteine or RGD (2.81mM) | PBS (pH 7.4) | 4Arm-PEG-thiol (50mg/mL) | PBS (pH 7.4) | HA-S (13.3mg/mL) | |
| 0.5% (w/v) HA 1kPa | 10 µL | 4.00 µL | 26 µL | 1.00 µL | 9.00 µL | 30.0 µL |
| 0.5% (w/v) HA 2kPa | 20.0 µL | 4.00 µL | 16.0 µL | 8.00 µL | 2.00 µL | 30.0 µL |
| 0.1% (w/v) HA 1kPa | 10 µL | 4.00 µL | 26 µL | 8.00 µL | 26.0 µL | 6.00 µL |
Table 1. Hydrogel formulations yielding independent control of HA concentration and mechanical properties.
Discussion
Generation of reproducible data using this 3D culture system requires: 1) consistent batch-to-batch thiolation of HA, 2) practice to achieve efficient mixing of hydrogel precursors and handling of hydrogel cultures to prevent damage and 3) optimized seeding density for each cell line used.
When a particular weight percentage of HA is desired in the hydrogel, the degree of thiolation of HA determines the crosslink density. We recommend using a consistent amount of HA for each thiolation reaction to minimize batch-to-batch variation. We suggest that at least 300 mg of HA be used for each thiolation reaction so that the same batch of thiolated HA can be used for multiple experimental repeats. The molar ratio of thiols to maleimide groups on 4-arm PEG should be always be 1.1:1 to ensure maximum crosslinking efficiency. Thus, if the degree of thiolation is varied, then the amount of PEG-Mal added must be adjusted accordingly. When peptides are conjugated to maleimides prior to gel formation, the estimated number of PEG arms with tethered peptides must be subtracted from the total number of available maleimides for this calculation. In addition, we recommend keeping the degree of HA thiolation below 10-15% to avoid disulfide bond formation that will prevent dissolution and consistent gelation.
The Michael-type addition reaction between thiol and maleimide groups ensures that gelation occurs in under a minute. While this fast gelation minimizes the amount of time cells suspended in hydrogel precursors are without complete medium, which may compromise their viability, the encapsulation process requires experience with all pipetting and mixing steps to achieve reproducible results. Typically, we have new trainees practice several rounds of gelation without cells before performing "real" encapsulation experiments.
Two factors can be tweaked to modulate gelation speed: reaction temperature and precursor pH. Lowering the reaction temperature by placing the well plate on ice while mixing slows the reaction and provides for greater time to mix precursor solutions. However, this must be done under sterile conditions using proper aseptic techniques. Similarly, the pH of precursor solutions can be altered to increase or decrease reaction time by using higher or lower pH, respectively. Since thiolated HA is dialyzed against acidic water to prevent disulfide bond formation, reconstituted thiolated HA will yield lower pH than might otherwise be expected. In general, reconstitution in 20 mM HEPES in HBSS buffer, adjusted to pH 9, will yield a pH near 7 when 15 mg/mL of thiolated HA (~5% thiolated) is dissolved. We recommend using pH 6.8-7 thiolated HA solution to allow even mixing of gel solution while maximizing cell health.
Seeding density during encapsulation is critical for cellular viability and experimental reproducibility. We recommend seeding in the range of 100,000 to 2,000,000 cells/mL. We have found that this range is optimal for viability of most cell types while preventing over-confluency within an experimental time frame of at least 3 weeks. The initial seeding density that optimizes viability should be determined experimentally for each cell type used.
Extreme care should be taken when changing the cell medium so as to avoid damaging hydrogel cultures. In particular, take care not to aspirate hydrogel into the pipette. Thus, do not use vacuum aspiration and instead use a sterile transfer pipette to aspirate manually. Changing only half the medium in a culture well at a time may also help.
In general, this technique requires practice to achieve the necessary manual dexterity and consistency. Once a lab-specific protocol is established, the 3D culture system enables the researcher to use many characterization methods as typical for cell culture and explanted tissues. Here, we have described several techniques for analysis, including immunofluorescence, flow cytometry, western blotting, and bioluminescence imaging of live, 3D cultures (Figure 2-3). For more detailed protocols of characterization methods, please see Xiao et al. 20173. Finally, as HA/PEG hydrogels are optically transparent, standard light microscopy techniques, including confocal imaging, can be used to monitor live, 3D cultures.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported with funding from the NIH (R21NS093199) and the UCLA ARC 3R’s Award. Our sincerest thanks go to the lab of Dr. Harley Kornblum for provision of the HK301 and HK157 cell lines. We also thank UCLA Tissue Pathology Core Laboratory (TPCL) for cryosectioning, Advanced Light Microscopy/Spectroscopy core facility (ALMS) in California Nanosystems Institute (CNSI) at UCLA for use of the confocal microscope, UCLA Crump Institute for Molecular Imaging for using IVIS imaging system, UCLA Molecular Instrumentation Center (MIC) for providing magnetic resonance spectroscopy, and Flow Cytometry Core in Jonsson Comprehensive Cancer Center (JCCC) at UCLA for providing instrumentation for flow cytometry.
Materials
| pH meter | Thermo Fisher | N/A | Any pH meter that has pH 2-10 sensitivity |
| Stir plate | Thermo Fisher | N/A | General lab equipment |
| Erlenmeyer flask (125mL) | Thermo Fisher | FB-501-125 | |
| dialysis tubes | Thermo Fisher | 21-152-14 | |
| 2L polypropylene beaker | Thermo Fisher | S01916 | |
| sodium hyaluronan | Lifecore | HA700k-5 | 500-750 kDa range |
| 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) | Thermo Fisher | PI-22980 | |
| N-hydroxysuccinimide (NHS) | sigma aldrich | 130672-5G | |
| Hydrochloric acid (HCl) | Thermo Fisher | SA48-500 | |
| Sodium hydroxide (NaOH) | Thermo Fisher | SS266-1 | |
| Cystamine dihydrochloride | Thermo Fisher | AC111770250 | |
| Dithiolthreitol (DTT) | Thermo Fisher | BP172-25 | |
| Ellman's test reagent (5-(3-Carboxy-4-nitrophenyl)disulfanyl-2-nitrobenzoic acid | Sigma Aldrich | D218200-1G | |
| Deuterated water (deuterium oxide) | Thermo Fisher | AC166301000 | |
| 0.22µm vacuum driven filter | CellTreat | 229706 | |
| Phosphate buffered saline (PBS) | Thermo Fisher | P32080-100T | |
| Hanks' balanced salt saline (HBSS) | Thermo Fisher | MT-21-022-CV | |
| 4-arm-PEG-maleimide | JenKem Technology | A7029-1 | molecular weight around 20kDa |
| 4-arm-PEG-thiol | JenKem Technology | A7039-1 | molecular weight around 20kDa |
| L-Cysteine | sigma aldrich | C7880-100G | |
| RGD ECM mimetic peptide | Genscript Biotech | N/A | Custom peptide with sequence "GCGYGRGDSPG", N-terminal should be acetylated |
| silicone molds | Sigma Aldrich | GBL664201-25EA | Use razor blade to cut into single pieces |
| complete culture medium | Various | Various | DMEM/F12 (Thermofisher) with non-serum supplement (G21 from GeminiBio), epidermal growth factor 50ng/mL (Peprotech), fibroblast growth factor 20ng/mL (Pepro Tech) and heprain 25µg/mL (Sigma Aldrich), culture medium varies in different labs |
| patient derived GBM cell | N/A | N/A | |
| 20G needle | BD medical | 305175 | |
| 1mL syringe | Thermo Fisher | 14-823-434 | |
| 10mL syringe | BD medical | 302995 | |
| RIPA Buffer | Thermo Fisher | PI-89901 | |
| protease/phosphatase inhibitor mini tablet | sigma aldrich | 5892970001 | |
| vortex shaker | Thermo Fisher | 12-814-5Q | |
| TrypLE express | Thermo Fisher | 12604013 | |
| 70µm cell strainer | Thermo Fisher | 22-363-548 | |
| Paraformaldehyde | Thermo Fisher | AC416785000 | Dissolve 4% (w/v) in PBS, keep pH 7.4 |
| D-sucrose | Thermo Fisher | BP220-1 | |
| Optimal Cutting Temperature (O.C.T.) compound | Thermo Fisher | NC9373881 | |
| Cell culture incubator | Thermo Fisher | N/A | Any General One with 5% CO2 and 37C |
| fridge/freezer | Thermo Fisher | N/A | Any General Lab equipment with -20C and -80C capacity |
| Disposable embedding molds | Thermo Fisher | 12-20 | |
| Lyapholizer | Labconco | N/A | Any -105C freeze dryers |
| HEPES | Thermo Fisher | BP310-500 | |
| Amber vial | Kimble Chase | 60912D-2 | |
| Wide orifice pipette tips | Thermo Fisher | 9405120 | |
| 2-methylbutane | Thermo Fisher | 03551-4 | |
| Dry Ice | N/A | N/A |
References
- Xiao, W., Sohrabi, A., Seidlits, S. K. . Integrating the glioblastoma microenvironment into engineered experimental models. , (2017).
- Tibbitt, M. W., Anseth, K. S. Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnology and bioengineering. 103 (4), 655-663 (2009).
- Xiao, W., et al. Brain-mimetic 3d culture platforms allow investigation of cooperative effects of extracellular matrix features on therapeutic resistance in glioblastoma. 암 연구학. 78 (5), 1358-1370 (2018).
- Holland, E. C. Glioblastoma multiforme: the terminator. Proceedings of the National Academy of Sciences. 97 (12), 6242-6244 (2000).
- Ostrom, Q. T., et al. CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2006 – 2010. Journal of Neuro-Oncology. 15 (6), 788-796 (2013).
- Bellail, A. C., Hunter, S. B., Brat, D. J., Tan, C., Van Meir, E. G. Microregional extracellular matrix heterogeneity in brain modulates glioma cell invasion. International Journal of Biochemistry and Cell Biology. 36 (6), 1046-1069 (2004).
- Zamecnik, J. The extracellular space and matrix of gliomas. Acta Neuropathologica. 110 (5), 435-442 (2005).
- Jadin, L., et al. Hyaluronan expression in primary and secondary brain tumors. Annals of translational medicine. 3 (6), (2015).
- Park, J. B., Kwak, H. J., Lee, S. H. Role of hyaluronan in glioma invasion. Cell adhesion & migration. 2 (3), 202-207 (2008).
- Pedron, S., et al. Spatially graded hydrogels for preclinical testing of glioblastoma anticancer therapeutics. MRS communications. 7 (3), 442-449 (2017).
- Jiglaire, C. J., et al. Ex vivo cultures of glioblastoma in three-dimensional hydrogel maintain the original tumor growth behavior and are suitable for preclinical drug and radiation sensitivity screening. Experimental cell research. 321 (2), 99-108 (2014).
- Florczyk, S. J., et al. Porous chitosan-hyaluronic acid scaffolds as a mimic of glioblastoma microenvironment ECM. Biomaterials. 34 (38), 10143-10150 (2013).
- Day, A. J., Prestwich, G. D. Hyaluronan-binding proteins: tying up the giant. Journal of Biological Chemistry. 277 (7), 4585-4588 (2002).
- Wiranowska, M., Tresser, N., Saporta, S. The effect of interferon and anti-CD44 antibody on mouse glioma invasiveness in vitro. Anticancer Research. 18 (5A), 3331-3338 (1998).
- Charles, N. A., Holland, E. C., Gilbertson, R., Glass, R., Kettenmann, H. The brain tumor microenvironment. GLIA. 59 (8), 1169-1180 (2011).
- Gilg, A. G., et al. Targeting hyaluronan interactions in malignant gliomas and their drug-resistant multipotent progenitors. Clinical Cancer Research. 14 (6), 1804-1813 (2008).
- Misra, S., Hascall, V. C., Markwald, R. R., Ghatak, S. Interactions between hyaluronan and its receptors (CD44, RHAMM) regulate the activities of inflammation and cancer. Frontiers in immunology. 6, 201 (2015).
- Varga, I., et al. Expression of invasion-related extracellular matrix molecules in human glioblastoma versus intracerebral lung adenocarcinoma metastasis. Zentralblatt fur Neurochirurgie. 71 (4), 173-180 (2010).
- Guo, W., Giancotti, F. G. Integrin signalling during tumour progression. Nature Reviews Molecular Cell Biology. 5 (10), 816-826 (2004).
- Bello, L., et al. αvβ3 and αvβ5 integrin expression in glioma periphery. Neurosurgery. 49 (2), 380-390 (2001).
- Chamberlain, M. C., Cloughsey, T., Reardon, D. A., Wen, P. Y. A novel treatment for glioblastoma: integrin inhibition. Expert review of neurotherapeutics. 12 (4), 421-435 (2012).
- Chopra, A., et al. Augmentation of integrin-mediated mechanotransduction by hyaluronic acid. Biomaterials. 35 (1), 71-82 (2014).
- Kim, Y., Kumar, S. CD44-mediated adhesion to hyaluronic acid contributes to mechanosensing and invasive motility. Molecular Cancer Research. 12 (10), 1416-1429 (2014).
- Joo, K. M., et al. Patient-specific orthotopic glioblastoma xenograft models recapitulate the histopathology and biology of human glioblastomas in situ. Cell Reports. 3 (1), 260-273 (2013).
- Oh, Y. T., et al. Translational validation of personalized treatment strategy based on genetic characteristics of glioblastoma. PloS one. 9 (8), e103327 (2014).
- Pedron, S., Becka, E., Harley, B. A. C. Regulation of glioma cell phenotype in 3D matrices by hyaluronic acid. Biomaterials. 34 (30), 7408-7417 (2013).
- Wang, C., Tong, X., Yang, F. Bioengineered 3D brain tumor model to elucidate the effects of matrix stiffness on glioblastoma cell behavior using PEG-based hydrogels. Molecular pharmaceutics. 11 (7), 2115-2125 (2014).
- Zhu, J. Bioactive modification of poly(ethylene glycol) hydrogels for tissue engineering. Biomaterials. 31 (17), 4639-4656 (2010).
- Stern, R., Asari, A. A., Sugahara, K. N. Hyaluronan fragments: an information-rich system. European journal of cell biology. 85 (8), 699-715 (2006).
- Riddles, P. W., Blakeley, R. L., Zerner, B. Ellman’s reagent: 5, 5′-dithiobis (2-nitrobenzoic acid)-a reexamination. Analytical biochemistry. 94 (1), 75-81 (1979).
- Jin, R., et al. Synthesis and characterization of hyaluronic acid-poly (ethylene glycol) hydrogels via Michael addition: An injectable biomaterial for cartilage repair. Acta biomaterialia. 6 (6), 1968-1977 (2010).
- Ozawa, T., James, C. D. Establishing intracranial brain tumor xenografts with subsequent analysis of tumor growth and response to therapy using bioluminescence imaging. Journal of visualized experiments. 41, 3-7 (2010).

