Enrichment of mRNA and Bisulfite-mRNA Library Preparation for Next-Generation Sequencing
Summary
This protocol provides an easy-to-follow workflow to conduct poly(A) RNA purification, bisulfite conversion, and library preparation using standardized equipment for a biological sample of interest.
Abstract
RNA post-transcriptional modifications in various types of RNA transcripts are associated with diverse RNA regulation in eukaryotic cells. Aberrant RNA 5-methylcytosine modifications and the dysregulated expression of RNA methyltransferases have been shown to be associated with various diseases, including cancers. Transcriptome-wide bisulfite-sequencing was developed to characterize the positions and the quantitative cytosine methylation levels in the bisulfite-converted RNA at the base-pair resolution. Herein, this protocol presents the procedures of two rounds of poly(A) RNA purification, three cycles of bisulfite reaction, and library preparation in detail to allow the transcriptome-wide mapping of mRNA 5-methylcytosine modification sites. The assessment of RNA quantity and quality after the main reaction is essential to monitor RNA integrity and is a critical step for ensuring high-quality sequencing libraries. Ideally, the procedures can be completed within three days. With this protocol, using high-quality total RNA as the input can practically build up robust bisulfite-mRNA libraries for next-generation sequencing from the sample of interest.
Introduction
Among over 150 types of post-transcriptional modifications1, 5-methylcytosine (m5C) modification has been identified in various types of RNAs, including ribosomal RNA, transfer RNA, messenger RNA, micro RNA, long non-coding RNA, vault RNA, enhancer RNA, and small cajal body-specific RNAs2. The RNA m5C is associated with diverse biological and pathological mechanisms such as regulating plant root development3, viral gene expression4, and cancer progression5. The aim of this protocol is to provide streamlined pipelines to characterize the transcriptome-wide mRNA m5C modification profile of biological samples in different developmental stages or in the disease setting. Transcriptome-wide bisulfite-sequencing was developed to characterize the positions and the quantitative cytosine methylation levels in the bisulfite-converted RNA at base-pair resolution6,7,8,9. This is particularly useful when studying the association of m5C with gene expression and RNA fate that is involved in the biological regulatory mechanisms in cells. In the mammalian cell, there are two known m5C readers: ALYREF can recognize m5C at the nucleus and serves as an mRNA nucleus-to-cytosol transporter10, while YBX1 can recognize m5C in the cytoplasm and increase mRNA stabilization11. Aberrant m5C mRNAs related to immune pathways were reported in Systemic Lupus Erythematosus CD4+ T cells12. Studies have revealed an association between mRNA m5C modification and modulation of cancer immunity and cancer progression13,14. Hence, mapping the m5C modification profile on the mRNA can provide crucial information to elucidate the potential regulatory machinery.
To investigate the functional roles of RNA m5C modification under certain biological conditions, the bisulfite conversion-based (bsRNA-seq) and antibody affinity enrichment-based approaches such as m5C-RIP-seq, miCLIP-seq, and 5-Aza-seq can be combined with the high-throughput sequencing platform to provide efficient detection of targeted regions and sequences with the m5C modifications on a transcriptome-wide scale15,16. The advantage of this protocol provides the comprehensive RNA m5C landscape at a single-base resolution since the antibody affinity enrichment-based approach relies upon the availability of high-quality antibodies and could achieve the single-fragment resolution of m5C methylation landscape17.
All RNA samples will be processed with two rounds of mRNA enrichment using oligo(dT) beads, three cycles of bisulfite reaction, and the sequencing library preparation. To monitor the RNA quality, each RNA sample will be examined by capillary gel electrophoresis before and after the procedures of mRNA purification and bisulfite reaction to assess fragment distribution. The purified libraries will be examined by their PCR amplicon qualities, DNA size distribution fragments by capillary gel electrophoresis, and their overall quantities examined by fluorescence dyes-based quantitative assays before sequencing. The system can also be used to analyze a broad spectrum of biological samples such as agricultural produce, isolated virions, cell lines, model organisms, and pathological specimens.
Protocol
1. Poly(A) RNA purification
NOTE: Use the total RNA treated with DNase I and examine the total RNA quality and integrity by capillary or conventional gel electrophoresis assessment before proceeding to poly(A) RNA purification. Investigators should be able to identify the 28S and 18S rRNA ribosomal bands in the high molecular weight field and the 5.8S rRNA band in the low molecular weight field without any significant smear bands in the electropherogram. The purification steps essentially follow the manufacturer's instructions with minor modifications indicated in the specific steps. See the Table of Materials for details related to all materials and instruments used in this protocol.
- Preparation
- Warm up the reagent, washing buffer, oligo(dT) beads, and lysis buffer at room temperature for 30 min before conducting the following procedures.
- For the isolation of poly(A)-enriched RNA, prepare 10-20 µg of an aliquot of high-quality total RNA as the starting material by transferring an adequate quantity of RNA into a nuclease-free 1.5 mL microcentrifuge tube. Incubate in the heat block at 70 °C for 5 min, then keep on ice for 2 min.
NOTE: The quantities of total RNA as the starting material should be appropriately adjusted depending on the desired quantity of mRNA and the nature of the cells used. Empirically, the poly(A)-enriched RNA may account for 1-5% abundances in the total RNA. An aliquot of 10 µg of high-quality total RNA, should provide high-quality poly(A)-enriched RNA in the 100-500 ng range for the downstream bisulfite conversion. - Meanwhile, fully resuspend the oligo(dT) beads. Transfer the required amounts of beads suspension to a new 1.5 mL microcentrifuge tube, then place on the magnet for 3 min till the magnetic beads form a pellet.
NOTE: Use 50 µL oligo(dT) beads for 10 µg of total RNA input. The volume of beads in this step could be further optimized depending on the cells used. - Remove the supernatant and resuspend the oligo(dT) beads with equal amounts (the volume of the beads suspension used in step 1.1.3.) of lysis buffer and place on the magnet for 3 min before removing the supernatant.
NOTE: Do not overdry the beads; the user should quickly proceed to step 1.2.1.
- The first round of purification
- Add lysis buffer to the RNA sample tube (volume ratio 4:1) and mix thoroughly before transferring to the oligo(dT) beads.
NOTE: If the RNA sample volume is 20 µL, then add 80 µL of lysis buffer. - Resuspend the RNA: lysis buffer mix and oligo(dT) beads fully by pipetting at least 10x.
- Allow the samples to incubate with continuous rotation for 15 min at room temperature.
- Place the sample on the magnet for 5 min or until the supernatant is clear; discard the supernatant.
- Add 600 µL of Wash Buffer 1 to the beads and carefully mix to resuspend the beads.
- Place the tube on the magnet for 5 min or until the supernatant is clear, discard the supernatant, and repeat wash step 1.2.5 once.
- Wash the beads by adding 300 µL of Wash Buffer 2, and gently resuspend the beads.
- Place the tube on the magnet for 5 min or until the supernatant is clear, discard the supernatant, and repeat wash step 1.2.7 once.
- Add 30 µL of cold 10 mM Tris-HCl (Elution buffer) to the sample tube and incubate at 70 °C for 5 min.
- Quickly transfer the tube onto the magnet, and when the suspension is clear after 20 s or more, transfer the supernatant containing eluted poly(A) enriched RNA to the new 1.5 mL microcentrifuge tube.
- Add lysis buffer to the RNA sample tube (volume ratio 4:1) and mix thoroughly before transferring to the oligo(dT) beads.
- The second round of purification
- Add 120 µL (4x the volume of eluted sample) lysis buffer to the eluted RNA sample and mix thoroughly.
- Wash the beads used in the first round of purification with an equal amount of lysis buffer as step 1.1.4, pipet 10x to resuspend the beads, and then place them on the magnet for 5 min before discarding the supernatant.
- Transfer the RNA: lysis buffer mix to the washed beads and mix thoroughly by pipetting at least 10x.
- Incubate the sample with continuous rotation for 15 min at room temperature.
- Repeat steps 1.2.4 to 1.2.8 to remove salt and other RNA subtypes.
NOTE: The volume of washing buffer can be reduced to 300 µL of Wash Buffer 1 and 150 µL of Wash Buffer 2. - Add 25 µL (desired volume) of cold 10 mM Tris-HCl (Elution buffer) to the sample tube and incubate at 70 °C for 5 min.
- Quickly transfer the tube onto the magnet, and when the suspension is clear after 20 s or more, transfer the supernatant containing eluted poly(A) enriched RNA to a new 1.5 mL microcentrifuge tube.
- Transfer 2.2 µL to 8-strip tubes for quality assessment of RNA quantity and quality using RNA high-sensitivity assays.
- Store the sample at -20 °C for short periods and -70 °C for a longer period.
NOTE: This can be a pause point in the protocol.
2. Bisulfite conversion
NOTE: The centrifugation steps were all performed at room temperature. The procedures are essentially performed accordingly to the manufacturer's instructions but with an additional step 2.2 for adding the spike-in control mRNA sequence(s) before the bisulfite reaction step 2.3.
- Transfer 19 µL of purified poly(A) enriched RNA sample to a 0.2 mL 8-strip tube.
- Add 1.0 µL of spike-in control, which is free from any modifications at the predetermined 1:10,000 quantity ratio (i.e., 0.1 ng of luciferase mRNA: 1 µg of purified poly(A) enriched RNA) into the RNA sample of step 2.1.
NOTE: The spike-in control sequence can be Firefly luciferase RNA, Renilla luciferase, or in vitro transcribed RNA sequence with no methylated cytosines. - Add 130 µL of conversion regent to the tube and gently invert several times for thorough mixing.
- Place the tube in the PCR machine and perform three cycles of heating at 70 °C for 10 min followed by incubating at 64 °C for 45 min each and then a final step to cool down to 4 °C.
- Place the column onto the collection tube and transfer 250 µL of RNA binding buffer to the column.
- Transfer the sample from step 2.4 to the column from step 2.5 and pipet thoroughly.
- Add 400 µL of 95-100% of ethanol to the column, quickly close the cap, and invert several times.
- Centrifuge at 10,000 × g for 30 s at 25 °C and discard the flowthrough.
- Add 200 µL of RNA wash buffer to the column.
- Centrifuge at 10,000 × g for 30 s at 25 °C and discard the flowthrough.
- Add 200 µL of desulfonation buffer and incubate at room temperature for 30 min.
- Centrifuge at 10,000 × g for 30 s at 25 °C and discard the flowthrough.
- Add 400 µL of RNA wash buffer and centrifuge at 10,000 × g for 30 s at 25 °C and discard the flowthrough. Repeat this step once.
- Centrifuge again at 10,000 × g for 2 min at 25 °C to completely remove the residual wash buffer.
NOTE: The other centrifugation step 2.14 for 2 min before elution is to completely remove the wash buffer. The wash step is performed twice to wash out the high-salt component from the bisulfite reagent and the desulfonation buffer on the column. - Transfer the column to a new 1.5 mL microcentrifuge tube.
- Add 20-30 µL of Nuclease-free H2O to the column and stand at room temperature for 1 min.
- Centrifuge at 10,000 × g for 30 s at 25 °C.
- Transfer 2.2 µL to 8-strip tubes for the assessment of RNA quantity and quality.
- Store the bisulfite-treated RNA sample at -20 °C for short periods and -70 °C for a longer period.
NOTE: This can be a pause point in the protocol.
3. Bisulfite-treated mRNA library preparation
NOTE: Follow the library preparation instruction protocol section 4 for use with purified mRNA or rRNA-depleted RNA. The first priming step should follow the FFPE RNA protocol since the bisulfite treatment fragmentizes the RNA. Perform every step in the laminar flow hood and add the reaction mixture on an ice-chilled cooling rack..
- Priming the input RNA
- Transfer the desired amount of bisulfite-treated mRNA sample in a total maximum of 5 µL or add Nuclease-free H2O to make a total of 5 µL.
NOTE: A minimum of 10 ng of bisulfite-treated mRNA as input is recommended for this type of RNA library preparation and to generate the final molarity of the 4nM library product. - Add 1 µL of Random primer (lilac) to the sample and mix by pipetting.
- Place the sample tube in the PCR machine preset at 65 °C and lid at 105 °C for 5 min and hold at 4 °C.
- Transfer the desired amount of bisulfite-treated mRNA sample in a total maximum of 5 µL or add Nuclease-free H2O to make a total of 5 µL.
- First-strand cDNA synthesis
- Add 4 µL of synthesis reaction buffer (lilac), 8 µL of strand specificity reagent (brown), and 2 µL of synthesis enzyme mix (brown) to the sample, which makes a total volume of 20 µL. Mix thoroughly by pipetting at least 10x and quickly spin down.
- Place the sample tube in the PCR machine preset at 25 °C for 10 min, 42 °C for 50 min, 70 °C for 15 min, and hold at 4 °C.
NOTE: For inserts over 200 bases, a 42 °C incubation period of 50 min is suggested; for insert size below 200 bases, a 42 °C incubation period of 15 min is suggested.
- Second-strand cDNA synthesis
- Add 8 µL of synthesis reaction buffer with dUTP mix (orange), 4 µL of synthesis enzyme mix (orange), and 48 µL of Nuclease-free H2O. Mix thoroughly by pipetting at least 10x and quickly spin down.
- Place the sample tube in the PCR machine preset at 16 °C for 1 h with the lid set off or ≤40 °C.
- Purification of double-stranded cDNA with DNA purification beads
- Prewarm the DNA purification beads 30 min before conducting the purification and vortex to resuspend the beads.
- Add 144 µL of resuspended DNA purification beads to the cDNA sample from step 3.3.2 and mix thoroughly by pipetting at least 10x.
- Incubate at room temperature for 5 min.
- Place the sample tube on a magnetic rack for 5 min until the solution is clear and then, remove the supernatant and keep the beads that contain DNA.
- Add 200 µL of freshly made 80% ethanol to the tube with the beads while keeping the sample on the magnetic rack and incubate at room temperature for 30 s.
- Remove the supernatant and repeat step 3.4.5 once, but this time carefully remove all the residual supernatant.
- Air dry the beads for 1-5 min while keeping the sample on the magnet.
NOTE: Do not overdry the beads; the color should be dark brown. - Remove the sample from the magnetic rack and elute the DNA by adding 53 µL of 0.1x TE (10 mM Tris-HCl and 1 mM EDTA, pH 8.0) buffer to the beads. Mix thoroughly by pipetting at least 10x and incubate at room temperature for 2 min.
- Place the tube onto the magnetic rack for 5 min or until the solution is clear.
- Transfer 50 µL of supernatant containing double-stranded cDNA to a new 8-strip tube.
NOTE: The protocol can be stopped at this point and the sample can be stored at -30 °C.
- End preparation of cDNA library
- Add 7 µL of end-polishing Reaction Buffer (green) and 3 µL of the end-polishing Enzyme Mix (green) to the eluted double-stranded cDNA sample.
- Mix the mixture by pipetting 10x with a 50 µL volume setting without creating bubbles, then quickly spin down the sample.
- Place the sample in a PCR machine and incubate at 20 °C for 30 min, 65 °C for 30 min, and hold at 4 °C.
- Adaptor ligation
- Dilute the adaptor (red) according to the manufacturer's instructions.
- Add 2.5 µL of the diluted adaptor, 1 µL of ligation enhancer (red), and 30 µL of Ligation Master Mix (red) to the sample.
- Mix the mixture by pipetting 10x with an 80 µL volume setting without creating bubbles, then quickly spin down the sample.
- Place the sample in a PCR machine and incubate at 20 °C for 15 min.
- Add 3 µL of the mixture of uracil DNA glycosylase and DNA glycosylase-lyase endonuclease VIII (blue) to the sample and pipet thoroughly.
- Place back the sample in the PCR machine and incubate at 37 °C for 15 min with the lid set to 75 °C.
- Purification of ligation reaction treated cDNAs
NOTE: The size selection with SPRIselect Beads, or optionally with AMpure XP beads can both be used for this purpose18.- Prewarm the beads 30 min before conducting the purification and vortex to resuspend them.
- Transfer 25 µL of resuspended beads to the cDNA sample from step 3.6.6 and mix thoroughly by pipetting at least 10x.
- Incubate at room temperature for 5 min.
- Place the sample tube on a magnetic rack for 5 min until the solution is clear and then transfer the supernatant to a new 8-strip tube and discard the beads.
- Add 10 µL of resuspended beads to the supernatant and mix thoroughly by pipetting at least 10x.
- Place the sample tube on a magnetic rack for 5 min until the solution is clear and then discard the supernatant and do not disturb the beads.
- Add 200 µL of freshly made 80% ethanol to the tube with the beads while keeping the sample on the magnetic rack and incubate at room temperature for 30 s.
- Remove the supernatant and repeat step 3.7.7 once, but this time carefully remove all the residual supernatant.
- Air-dry the beads for 1-5 min while keeping the sample on the magnet.
NOTE: Do not overdry the beads; the color should be dark brown. - Remove the sample from the magnetic rack and elute the DNA by adding 17 µL of 0.1x TE buffer to the beads. Mix thoroughly by pipetting at least 10x and incubate at room temperature for 2 min.
- Place the tube onto the magnetic rack for 5 min until the solution is clear.
- Transfer 15 µL of supernatant, which contains target end-prepared dsDNA to a new 8-strip tube.
- PCR enrichment of adaptor-ligated DNA
- Add 25 µL of master mix (blue), 5 µL of 5' adaptor primer, and 5 µL of the 3' adaptor primer to the adaptor-ligated DNA sample. Mix thoroughly by pipetting at least 10x.
- Place the sample in the PCR machine and perform PCR amplification using the following steps: (1) 98 °C for 30 s, (2) 98 °C for 10 s, (3) 65 °C for 75 s, (4) 65 °C for 5 min, and (5) hold at 4 °C; in which the steps (2) and (3) should repeat for an N+2 cycle number where N equals to the RNA input quantity in the instruction manual. This means 2 additional cycles are needed due to the double-side size selection of samples performed in step 3.7.
NOTE: For instance, 8 ng of the purified mRNA input would be recommended with 9-10 cycles of PCR; but with double-side, size-selected, ligated cDNA in step 3.7, the recommended PCR cycle would be 11-12 cycles.
- Purification of PCR-amplified libraries
- Prewarm the DNA purification beads for 30 min before conducting the purification and vortex to resuspend the beads.
- Transfer 45 µL (0.9x) of resuspended beads to the PCR product from step 3.8.2 and mix thoroughly by pipetting at least 10x.
- Incubate at room temperature for 5 min.
- Place the sample tube on a magnetic rack for 5 min until the solution is clear and then remove the supernatant and keep the beads that contain DNA.
- Add 200 µL of freshly made 80% ethanol to the tube with the beads, leave the sample on the magnetic rack and allow it to stand at room temperature for 30 s.
- Remove the supernatant and repeat step 3.9.5 once, but this time carefully remove all the residual supernatant.
- Air-dry the beads for 1-5 min while keeping the sample on the magnet.
NOTE: Do not overdry the beads; the color should be dark brown. - Take the sample off the magnetic rack and add 22 µL of 0.1x TE buffer to the beads to elute the DNA. Mix thoroughly by pipetting at least 10x and incubate at room temperature for 2 min.
- Place the tube onto the magnetic rack and allow it to stand for 5 min or until the solution is clear at room temperature.
- Transfer 20 µL of supernatant to a new 8-strip tube.
- Transfer 2.2 µL to 8-strip tubes for quality assessment of library quantity and quality.
NOTE: The quantity of the library can also be detected by qPCR (see the Table of Materials). - Store the bsRNA-seq library sample at -20 °C.
Representative Results
A series of bsRNA-seq libraries from cell lines19 were generated by following the procedures in this report (Figure 1). After total RNA purification accompanied by DNase treatment performed on cell line samples and the quality checked by gel electrophoresis and UV-Vis spectrophotometry (A260/A280), the RNA sample can proceed to poly(A) RNA enrichment. To determine whether the double purification could remove the majority of ribosomal RNA, the purification efficiency of poly(A) RNA was assessed by capillary electrophoresis total RNA assay that can automatically calculate the rRNA contamination percentage (Figure 2). From the RNA fragment peak pattern and rRNA contamination percentage, RNA samples with twice purification indeed showed decreased contamination of ribosomal RNA from 6.5% to 2.2% and 2.6% to 1.1% in AsPC-1 and BxPC-3, respectively. With human pancreatic cancer cell lines, two rounds of beads purification could reach an average of 2.8 ± 1% poly(A)-enriched RNA abundances from the total RNA samples. Therefore, the poly(A) RNA enrichment by oligo(dT) beads with double purification steps minimized ribosomal RNA and represents a feasible mRNA sample enrichment for the downstream experiments.
The bisulfite conversion protocols had been reported in several studies on RNA 5-methylcytosine modification (Table 1). The bisulfite reaction could be performed with a user-reconstituted reaction mixture consisting of 40% sodium bisulfite and 600 mM hydroquinone in the aqueous solution (pH 5.1) at 75 °C for 4 hours10,20,21. Alternatively, a more stringent bisulfite treatment of RNA sample may also be performed with a commercial kit; which in a number of studies from others and ours6,22,23,24 extended the bisulfite reaction time to three incubation cycles. Since the three-cycle incubation step efficiently converts unmethylated cytosine in the in vitro transcribed and structurally more-folded Escherichia coli 16S rRNA segment8, the three-cycle incubation step was also applied in this protocol.
With twice poly(A) enrichment and the three-cycle bisulfite conversion, RNA quality before and after the reaction was assessed by capillary electrophoresis. The RNA size distribution after the bisulfite treatment showed a peak of ~200-500 nucleotides because of fragmentation caused by the bisulfite reaction (Figure 3). Additionally, the fragmented bsRNA is ideal for library preparation without the need to conduct another fragmentation step. A total of 8-10 ng of bsRNA as input was used to construct the library according to this protocol by using 9 to 11 cycles in the final PCR amplification of adapter-ligated DNA. The capillary electrophoresis of the amplicon showed quite a successful library preparation, with only a small portion of primer remaining and no peak of over-amplification (Figure 4). To check the conversion efficiency in each sample library, the unmethylated firefly luciferase RNA as a spike-in control sequence was added to the sample before conducting the bisulfite treatment. After the sequencing reads aligned to the reference sequence using meRanTK tool kits25, the average of 2,440 (0.006%) of total reads were mapped to the spike-in sequence, and the total analyzed C-to-T conversion rate reached an average of 99.81%; the status can be viewed in Integrated Genomics Viewer (Figure 5).
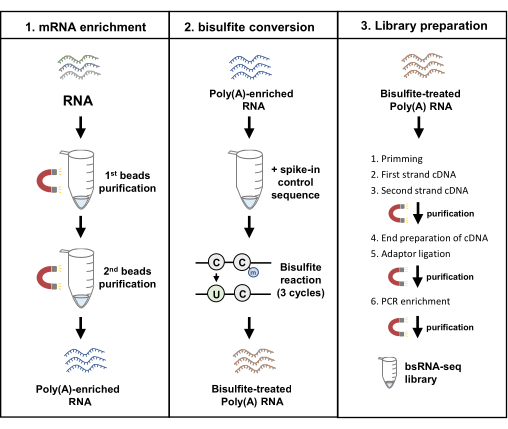
Figure 1: Workflow diagram of bisulfite mRNA sequencing. The pipeline constitutes of three main protocols, including mRNA enrichment, bisulfite conversion, and library preparation. Abbreviation: bsRNA = bisulfite mRNA; QC. = quality control. Please click here to view a larger version of this figure.
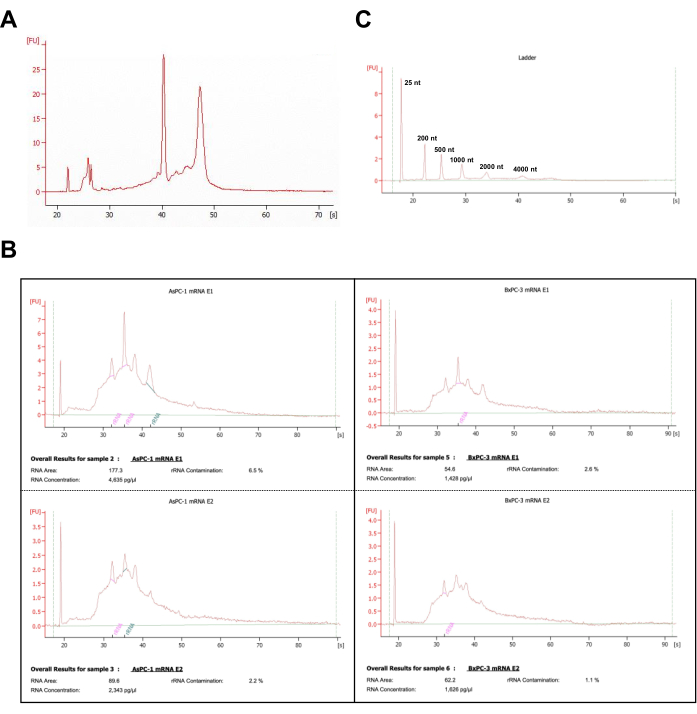
Figure 2: Quality control results of poly(A)-enriched RNA samples with capillary electrophoresis. (A) The representative capillary electrophoresis profile of the total RNA from BxPC-3 cells. (B) The representative capillary electrophoresis profiles of samples processed with oligo(dT) purification from AsPC-1 and BxPC-3 cells. mRNA E1, mRNA elution from one-time purification; mRNA E2, mRNA elution from two-times purification. The rRNA contamination estimates were calculated by the electrophoresis operation system that controls the total RNA analysis assay. (C) The capillary electrophoresis profile of the Ladder markers was obtained in the same run with samples presented in panel B. Please click here to view a larger version of this figure.
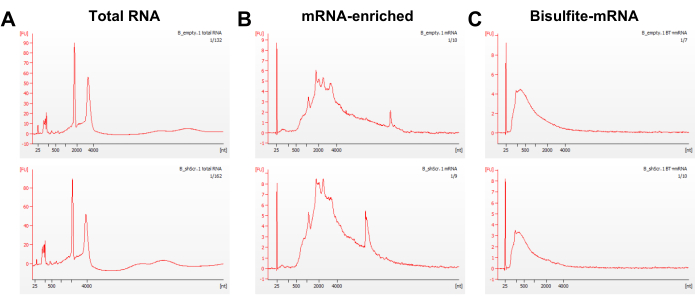
Figure 3: Quality assessment of RNA samples by capillary electrophoresis. The representative capillary electrophoresis profiles of (A) the total RNA, (B) the poly(A)-enriched mRNA from two times purification, and (C) the bisulfite-treated mRNA from the BxPC-3 cells. Abbreviations: B_empty = BxPC-3 cells transduced with empty vector; B_shScr = BxPC-3 cells transduced with the scramble sequences. Please click here to view a larger version of this figure.
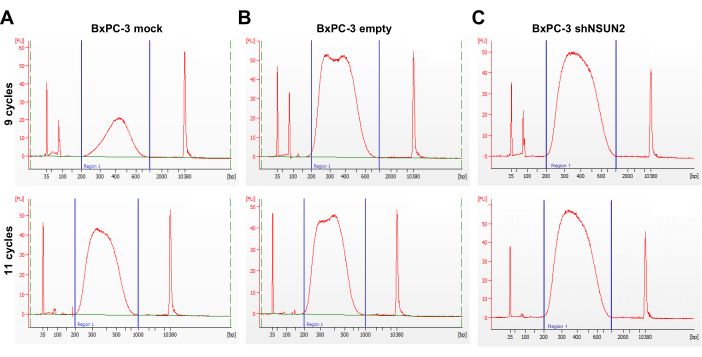
Figure 4: Quality assessment of bsRNA-seq libraries. (A-C) Three sets of representative capillary electrophoresis profiles of constructed bsRNA-seq libraries amplified by PCR at different PCR cycle numbers of 9 and 11 cycles using bsRNA from a set of BxPC-3 cells. The estimated input quantity of the bisulfite-treated RNA was determined by a high-sensitivity RNA quantification assay. A total of 8, 8.4, and 9.6 ng bsRNA were used in the (A) BxPC-3 mock, (B) BxPC-3 empty, and (C) BxPC-3 shN2UN2 samples, respectively. The ladder marker peaks at 35 bp and the 10,380 bp represent the internal standards of the lower- and upper-boundary in the electrophoresis profile of each sample. A prominent peak of around 100 bp and an obscure peak of around 127 bp, respectively, indicate the PCR primers (<100 bp) and the adaptor-dimer (~127 bp), which remained in the eluate after PCR cleanup. Abbreviations: BxPC-3 mock = untransduced BxPC-3 cells; BxPC-3 empty = BxPC-3 cells transduced with the empty vector control; BxPC-3 shN2UN2 = BxPC-3 cells transduced with the shNSUN2 constructs. Please click here to view a larger version of this figure.
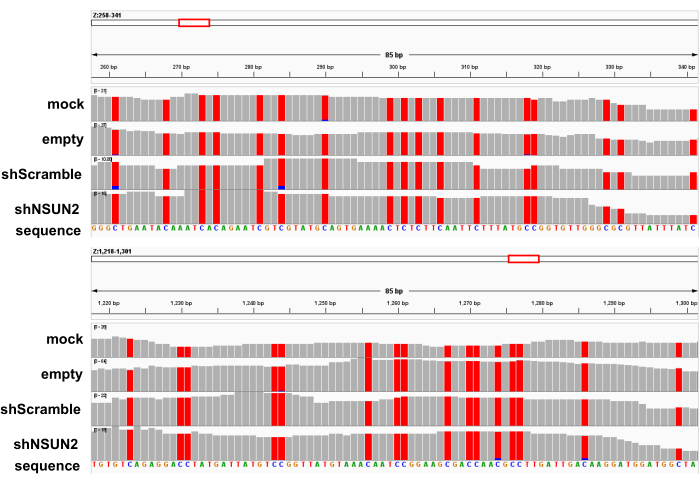
Figure 5: The sequence alignment profiles of each bsRNA-seq library mapped to the spike-in-control reference sequence. The representative alignment profiles of the BxPC-3 mock, empty, scramble, and shNSUN2 bsRNA-seq libraries to the firefly luciferase spike-in control reference sequence (indicated as the "Z" gene); 2 representative regions were shown. Grey bars indicate all reads that present matched consensus nucleotides at the indicated base position to the reference sequence. Red bars highlight the thymidine (T) identity at the base position of the reads; blue bars indicate those reads having the cytosine (C) identity at the position. Please click here to view a larger version of this figure.
Table 1: Comparison of bisulfite reaction protocols and the conversion rate. Please click here to download this Table.
Discussion
In this protocol, a detailed pipeline of poly(A) enrichment, bisulfite conversion, and library preparation was achieved by utilizing standardized components. Further sequencing analysis provided the identification of mRNA 5-methylcytosine in samples of interest.
The critical step is the quality of starting material-total RNA-since the degradation of RNA would impact the recovery rate of poly(A) RNA purification. The sample should be carefully handled and RNase contamination avoided before conducting the poly(A) RNA purification step. Another crucial part of the procedure is the number of PCR cycles in library preparation. The decision of cycle number depends on the quantity of bisulfite-treated mRNA used for library preparation and whether the use of one size selection or double size selection in the step before PCR. The suitable cycle numbers for the sample of interest would thus require a trial run and assess the PCR product size distribution by capillary electrophoresis to determine the optimal cycle number for the same cell line or tissue.
In this protocol, two rounds of oligo(dT) beads purification were used to enrich and purify poly(A) RNA and eliminated a majority of ribosomal RNAs and other RNAs. There are other methods to remove ribosomal RNA and keep other RNA types in the sample, such as oligo-based depletion of rRNA26. Then, the 5-methylcytosine modification present in other RNA types can also be taken into analysis and extend the understanding of 5-methylcytosine features in mRNA and other RNAs.
Bisulfite reaction is known for not being able to distinguish m5C from other modifications, including the 5-hydroxymethylcytosine (hm5C), 3-methylcytidine (m3C), and N4-methylcytidine (m4C)27. However, the experimental design including m5C RNA methyltransferase knockdown or knockout should provide informative evidence of high-confidence regulated m5C sites. Additionally, validation using different methods such as antibody-based enrichment followed by sequencing or PCR or LC-MS/MS-based methods could essentially authenticate the candidate m5C sites16. The emerging technique of direct Nanopore RNA sequencing also provides potential identification of RNA modification via the computational analysis of the current signal or base-called 'error' features28,29,30.
A recent study by Johnson et al. showed that adding the fragmentation step in mitochondria RNA samples after three rounds of bisulfite conversion indeed showed a higher conversion rate and increased the yield of cDNA library product31. This paper specifically analyzed the conversion rate and reads quality mapped to the mitochondria genome, not the mRNA transcriptome. Hence, the use of the fragmentation step has been updated in Table 1 to highlight whether the published reports included an RNA fragmentation step so that the viewers can easily replicate the protocol. Another study by Zhang et al. demonstrated that utilizing in vitro transcribed modification-free RNA library as a negative control is efficient to eliminate the false-positive signal from bisulfite treatment32. Recent studies have compared different protocols used for library construction33. The user can further choose to conduct bsRNA-seq pipeline with or without an additional fragmentation step and in vitro transcribed modification-free RNA library to customize suitable workflow.
The transcriptome-wide identification of m5C by different principles could strengthen the findings of the modification landscapes and provide more understanding of the regulatory mechanisms of RNA modifications in healthy or disease models.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by the National Science and Technology Council of Taiwan. [NSTC 111-2314-B-006-003]
Materials
| Agilent 2100 Electrophoresis Bioanalyzer System | Agilent, Santa Clara, CA | RNA quality detection | |
| AMpure XP beads | Beckman Coulter | A63881 | purify DNA |
| Bioanalyzer DNA high sensitivity kit | Agilent, Santa Clara, CA | 5067-4626 | DNA quality dection |
| Bioanalyzer RNA 6000 Pico kit | Agilent, Santa Clara, CA | 5067-1513 | RNA quality dection |
| DiaMag02 – magnetic rack | Diagenode, Denville, NJ | B04000001 | assist library preparation |
| DiaMag1.5 – magnetic rack | Diagenode, Denville, NJ | B04000003 | assist poly(A) RNA purificaion |
| Dynabeads mRNA DIRECT purification kit | Thermo Fisher Scientific, Waltham, MA | 61011 | poly(A) RNA purificaion; Wash Buffer 1 and Wash Buffer 2 |
| Ethanol | J.T.Baker | 64-17-5 | |
| EZ RNA methylation kit | Zymo, Irvine, CA | R5002 | bisulfite treatment |
| Firefly luciferase mRNA | Promega, Madison, WI, USA | L4561 | spike in control seqeunce |
| KAPA Library Quantification Kits | Roche, Switzerland | KK4824 | library quantification |
| Nanodrop spectrophotometer | Thermo Fisher Scientific, Waltham, MA | Total RNA quantity detection | |
| NEBNext multiplex Oligos for illumina (index Primer set1) | New England Biolabs, Ipswich, MA | E7335S | library preparation |
NEBNext Ultra  Directional RNA Library Prep Kit for Illumina Directional RNA Library Prep Kit for Illumina |
New England Biolabs, Ipswich, MA | E7760S | library preparation |
| Nuclease-free Water | Thermo Fisher Scientific | AM9932 | |
| P2 pipetman | Thermo Fisher Scientific, Waltham, MA | 4641010 | |
| Qubit 2.0 fluorometer | Thermo Fisher Scientific, Waltham, MA | RNA quantity detection | |
| Qubit dsDNA HS Assay Kit | Thermo Fisher Scientific, Waltham, MA | Q32854 | DNA quantity detection |
| Qubit RNA HS Assay Kit | Thermo Fisher Scientific, Waltham, MA | Q32852 | RNA quantity detection |
References
- Boccaletto, P., et al. MODOMICS: a database of RNA modification pathways. 2021 update. Nucleic Acids Research. 50, D231-D235 (2022).
- Bohnsack, K. E., Höbartner, C., Bohnsack, M. T. Eukaryotic 5-methylcytosine (m5C) RNA Methyltransferases: Mechanisms, Cellular Functions, and Links to Disease. Genes. 10 (2), 102 (2019).
- Shinde, H., Dudhate, A., Kadam, U. S., Hong, J. C. RNA methylation in plants: An overview. Frontiers in Plant Science. 14, 1132959 (2023).
- Courtney, D. G., et al. Epitranscriptomic addition of m5C to HIV-1 transcripts regulates viral gene expression. Cell Host & Microbe. 26 (2), 217-227 (2019).
- Jonkhout, N., et al. The RNA modification landscape in human disease. RNA. 23 (12), 1754-1769 (2017).
- Legrand, C., et al. Statistically robust methylation calling for whole-transcriptome bisulfite sequencing reveals distinct methylation patterns for mouse RNAs. Genome Research. 27 (9), 1589-1596 (2017).
- Schaefer, M. RNA 5-methylcytosine analysis by bisulfite sequencing. Methods in Enzymology. 560, 297-329 (2015).
- Amort, T., et al. Distinct 5-methylcytosine profiles in poly(A) RNA from mouse embryonic stem cells and brain. Genome Biology. 18 (1), 1 (2017).
- Schumann, U., et al. Multiple links between 5-methylcytosine content of mRNA and translation. BMC Biology. 18 (1), 40 (2020).
- Yang, X., et al. 5-Methylcytosine promotes mRNA export – NSUN2 as the methyltransferase and ALYREF as an m5C reader. Cell Research. 27, 606 (2017).
- Chen, X., et al. 5-Methylcytosine promotes pathogenesis of bladder cancer through stabilizing mRNAs. Nature Cell Biology. 21 (8), 978-990 (2019).
- Guo, G., et al. Disease activity-associated alteration of mRNA m(5) C methylation in CD4(+) T cells of systemic lupus erythematosus. Frontiers in Cell and Developmental Biology. 8 (5), 430 (2020).
- Song, H., et al. Biological roles of RNA m5C modification and its implications in cancer immunotherapy. Biomarker Research. 10 (1), 15 (2022).
- Xue, C., Zhao, Y., Li, L. Advances in RNA cytosine-5 methylation: detection, regulatory mechanisms, biological functions and links to cancer. Biomarker Research. 8 (1), 43 (2020).
- Helm, M., Motorin, Y. Detecting RNA modifications in the epitranscriptome: predict and validate. Nature Reviews Genetics. 18 (5), 275-291 (2017).
- Guo, G., et al. Advances in mRNA 5-methylcytosine modifications: Detection, effectors, biological functions, and clinical relevance. Molecular Therapy – Nucleic Acids. 26, 575-593 (2021).
- Saplaoura, E., Perrera, V., Colot, V., Kragler, F. Methylated RNA Immunoprecipitation Assay to Study m5C Modification in Arabidopsis. Journal of Visualized Experiments:JoVE. (159), e61231 (2020).
- Quail, M. A., Swerdlow, H., Turner, D. J. Improved protocols for the illumina genome analyzer sequencing system. Current Protocols in Human Genetics. , (2009).
- Chen, S. -. Y., et al. RNA bisulfite sequencing reveals NSUN2-mediated suppression of epithelial differentiation in pancreatic cancer. Oncogene. 41 (22), 3162-3176 (2022).
- Squires, J. E., et al. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Research. 40 (11), 5023-5033 (2012).
- Han, X., et al. Dynamic DNA 5-hydroxylmethylcytosine and RNA 5-methycytosine reprogramming during early human development. Genomics, Proteomics & Bioinformatics. , (2022).
- Amort, T., et al. Long non-coding RNAs as targets for cytosine methylation. RNA biology. 10 (6), 1003-1008 (2013).
- Sun, Z., et al. Effects of NSUN2 deficiency on the mRNA 5-methylcytosine modification and gene expression profile in HEK293 cells. Epigenomics. 11 (4), 439-453 (2019).
- Huang, T., Chen, W., Liu, J., Gu, N., Zhang, R. Genome-wide identification of mRNA 5-methylcytosine in mammals. Nature Structural and Molecular Biology. 26 (5), 380-388 (2019).
- Rieder, D., Amort, T., Kugler, E., Lusser, A., Trajanoski, Z. meRanTK: methylated RNA analysis ToolKit. Bioinformatics. 32 (5), 782-785 (2015).
- Kraus, A. J., Brink, B. G., Siegel, T. N. Efficient and specific oligo-based depletion of rRNA. Scientific Reports. 9 (1), 12281 (2019).
- Edelheit, S., Schwartz, S., Mumbach, M. R., Wurtzel, O., Sorek, R. Transcriptome-wide mapping of 5-methylcytidine RNA modifications in bacteria, archaea, and yeast reveals m(5)C within archaeal mRNAs. PLoS Genetics. 9 (5), e1003602 (2013).
- Furlan, M., et al. Computational methods for RNA modification detection from nanopore direct RNA sequencing data. RNA Biology. 18, 31-40 (2021).
- Leger, A., et al. RNA modifications detection by comparative Nanopore direct RNA sequencing. Nature Communications. 12 (1), 7198 (2021).
- Mateos, P. A., et al. Identification of m6A and m5C RNA modifications at single-molecule resolution from Nanopore sequencing. bioRxiv. , (2022).
- Johnson, Z., Xu, X., Pacholec, C., Xie, H. Systematic evaluation of parameters in RNA bisulfite sequencing data generation and analysis. NAR Genomics and Bioinformatics. 4 (2), 045 (2022).
- Zhang, Z., et al. Systematic calibration of epitranscriptomic maps using a synthetic modification-free RNA library. Nature Methods. 18 (10), 1213-1222 (2021).
- Chao, H. -. P., et al. Systematic evaluation of RNA-Seq preparation protocol performance. BMC Genomics. 20 (1), 571 (2019).

